Abstract
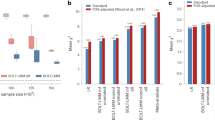
Testing for associations in big data faces the problem of multiple comparisons, wherein true signals are difficult to detect on the background of all associations queried. This difficulty is particularly salient in human genetic association studies, in which phenotypic variation is often driven by numerous variants of small effect. The current strategy to improve power to identify these weak associations consists of applying standard marginal statistical approaches and increasing study sample sizes. Although successful, this approach does not leverage the environmental and genetic factors shared among the multiple phenotypes collected in contemporary cohorts. Here we developed covariates for multiphenotype studies (CMS), an approach that improves power when correlated phenotypes are measured on the same samples. Our analyses of real and simulated data provide direct evidence that correlated phenotypes can be used to achieve increases in power to levels often surpassing the power gained by a twofold increase in sample size.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Stranger, B.E., Stahl, E.A. & Raj, T. Progress and promise of genome-wide association studies for human complex trait genetics. Genetics 187, 367–383 (2011).
Sham, P.C. & Purcell, S.M. Statistical power and significance testing in large-scale genetic studies. Nat. Rev. Genet. 15, 335–346 (2014).
Locke, A.E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Zhou, X. & Stephens, M. Efficient multivariate linear mixed model algorithms for genome-wide association studies. Nat. Methods 11, 407–409 (2014).
O'Reilly, P.F. et al. MultiPhen: joint model of multiple phenotypes can increase discovery in GWAS. PLoS One 7, e34861 (2012).
Aschard, H. et al. Maximizing the power of principal-component analysis of correlated phenotypes in genome-wide association studies. Am. J. Hum. Genet. 94, 662–676 (2014).
Stephens, M. A unified framework for association analysis with multiple related phenotypes. PLoS One 8, e65245 (2013).
Liang, L. et al. A cross-platform analysis of 14,177 expression quantitative trait loci derived from lymphoblastoid cell lines. Genome Res. 23, 716–726 (2013).
Stegle, O., Parts, L., Durbin, R. & Winn, J. A Bayesian framework to account for complex non-genetic factors in gene expression levels greatly increases power in eQTL studies. PLoS Comput. Biol. 6, e1000770 (2010).
Greenland, S., Pearl, J. & Robins, J.M. Causal diagrams for epidemiologic research. Epidemiology 10, 37–48 (1999).
Hernán, M.A., Hernández-Díaz, S., Werler, M.M. & Mitchell, A.A. Causal knowledge as a prerequisite for confounding evaluation: an application to birth defects epidemiology. Am. J. Epidemiol. 155, 176–184 (2002).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Farrar, D.E. & Glauber, R.R. Multicollinearity in regression analysis: the problem revisited. Rev. Econ. Stat. 49, 92–107 (1967).
Aschard, H., Vilhjálmsson, B.J., Joshi, A.D., Price, A.L. & Kraft, P. Adjusting for heritable covariates can bias effect estimates in genome-wide association studies. Am. J. Hum. Genet. 96, 329–339 (2015).
Kettunen, J. et al. Genome-wide association study identifies multiple loci influencing human serum metabolite levels. Nat. Genet. 44, 269–276 (2012).
Shin, S.Y. et al. An atlas of genetic influences on human blood metabolites. Nat. Genet. 46, 543–550 (2014).
Suhre, K. et al. Human metabolic individuality in biomedical and pharmaceutical research. Nature 477, 54–60 (2011).
Rhee, E.P. et al. A genome-wide association study of the human metabolome in a community-based cohort. Cell Metab. 18, 130–143 (2013).
Leek, J.T. & Storey, J.D. Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet. 3, 1724–1735 (2007).
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511 (2013).
Yu, C.H., Pal, L.R. & Moult, J. Consensus genome-wide expression quantitative trait loci and their relationship with human complex trait disease. OMICS 20, 400–414 (2016).
Westra, H.J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat. Genet. 45, 1238–1243 (2013).
Gibson, G. Rare and common variants: twenty arguments. Nat. Rev. Genet. 13, 135–145 (2012).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Dahl, A., Guillemot, V., Mefford, J., Aschard, H. & Zaitlen, N. Adjusting for principal components of molecular phenotypes induces replicating false positives. Preprint at https://www.biorxiv.org/content/early/2017/03/26/120899 (2017).
Dahl, A. et al. A multiple-phenotype imputation method for genetic studies. Nat. Genet. 48, 466–472 (2016).
Robinson, L.D. & Jewell, N.P. Some surprising results about covariate adjustment in logistic regression models. Int. Stat. Rev. 59, 227–240 (1991).
Peterson, C.B., Bogomolov, M., Benjamini, Y. & Sabatti, C. Many phenotypes without many false discoveries: error controlling strategies for multitrait association studies. Genet. Epidemiol. 40, 45–56 (2016).
Higham, N.J. Computing the nearest correlation matrix: a problem from finance. IMA J. Numer. Anal. 22, 329–343 (2002).
Devlin, B., Roeder, K. & Wasserman, L. Genomic control, a new approach to genetic-based association studies. Theor. Popul. Biol. 60, 155–166 (2001).
Liu, X., Huang, M., Fan, B., Buckler, E.S. & Zhang, Z. Iterative usage of fixed and random effect models for powerful and efficient genome-wide association studies. PLoS Genet. 12, e1005767 (2016).
Wang, T.J. et al. Metabolite profiles and the risk of developing diabetes. Nat. Med. 17, 448–453 (2011).
Townsend, M.K. et al. Reproducibility of metabolomic profiles among men and women in 2 large cohort studies. Clin. Chem. 59, 1657–1667 (2013).
Mayers, J.R. et al. Elevation of circulating branched-chain amino acids is an early event in human pancreatic adenocarcinoma development. Nat. Med. 20, 1193–1198 (2014).
Wolpin, B.M. et al. Genome-wide association study identifies multiple susceptibility loci for pancreatic cancer. Nat. Genet. 46, 994–1000 (2014).
Li, B. & Dewey, C.N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12, 323 (2011).
Acknowledgements
H.A. and N.Z. were supported by NIH grant R03DE025665. H.A. was also supported by NIH grant R21HG007687, and N.Z. was also supported by NIH career development award K25HL121295 and NIH grant U01HG009080. C.J.P. was supported by NIH grant R00 ES023504.
Author information
Authors and Affiliations
Contributions
H.A. conceived the approach and performed all real-data analyses. H.A., N.Z., B.V., C.J.P., D.S., and P.K. contributed substantially to improving the approach and the study design. C.J.Y. contributed to the quality control and analysis of the gEUVADIS data. B.W. collected the metabolite data and contributed to quality control and analysis of the metabolite data. H.A. and N.Z. conceptualized and performed the simulation study. V.G. contributed to the simulation study. H.A. and N.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–45, Supplementary Tables 1–8 and Supplementary Note (PDF 11510 kb)
Rights and permissions
About this article
Cite this article
Aschard, H., Guillemot, V., Vilhjalmsson, B. et al. Covariate selection for association screening in multiphenotype genetic studies. Nat Genet 49, 1789–1795 (2017). https://doi.org/10.1038/ng.3975
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3975
This article is cited by
-
Phenotype integration improves power and preserves specificity in biobank-based genetic studies of major depressive disorder
Nature Genetics (2023)
-
Wavelet Screening: a novel approach to analyzing GWAS data
BMC Bioinformatics (2021)
-
Mitochondrial DNA variants modulate N-formylmethionine, proteostasis and risk of late-onset human diseases
Nature Medicine (2021)
-
A comprehensive study of metabolite genetics reveals strong pleiotropy and heterogeneity across time and context
Nature Communications (2019)