Abstract
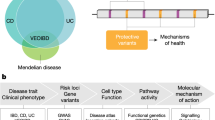
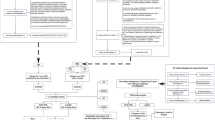
A major challenge in inflammatory bowel disease (IBD) is the integration of diverse IBD data sets to construct predictive models of IBD. We present a predictive model of the immune component of IBD that informs causal relationships among loci previously linked to IBD through genome-wide association studies (GWAS) using functional and regulatory annotations that relate to the cells, tissues, and pathophysiology of IBD. Our model consists of individual networks constructed using molecular data generated from intestinal samples isolated from three populations of patients with IBD at different stages of disease. We performed key driver analysis to identify genes predicted to modulate network regulatory states associated with IBD, prioritizing and prospectively validating 12 of the top key drivers experimentally. This validated key driver set not only introduces new regulators of processes central to IBD but also provides the integrated circuits of genetic, molecular, and clinical traits that can be directly queried to interrogate and refine the regulatory framework defining IBD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Liu, J.Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
Jostins, L. et al. Host–microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Franke, A. et al. Genome-wide meta-analysis increases to 71 the number of confirmed Crohn's disease susceptibility loci. Nat. Genet. 42, 1118–1125 (2010).
Kenny, E.E. et al. A genome-wide scan of Ashkenazi Jewish Crohn's disease suggests novel susceptibility loci. PLoS Genet. 8, e1002559 (2012).
Chen, G.B. et al. Estimation and partitioning of (co)heritability of inflammatory bowel disease from GWAS and Immunochip data. Hum. Mol. Genet. 23, 4710–4720 (2014).
Khor, B., Gardet, A. & Xavier, R.J. Genetics and pathogenesis of inflammatory bowel disease. Nature 474, 307–317 (2011).
Chen, Y. et al. Variations in DNA elucidate molecular networks that cause disease. Nature 452, 429–435 (2008).
Emilsson, V. et al. Genetics of gene expression and its effect on disease. Nature 452, 423–428 (2008).
Wang, I.M. et al. Systems analysis of eleven rodent disease models reveals an inflammatome signature and key drivers. Mol. Syst. Biol. 8, 594 (2012).
Zhang, B. et al. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer's disease. Cell 153, 707–720 (2013).
Haberman, Y. et al. Pediatric Crohn disease patients exhibit specific ileal transcriptome and microbiome signature. J. Clin. Invest. 124, 3617–3633 (2014).
Sandborn, W.J. et al. Ustekinumab induction and maintenance therapy in refractory Crohn's disease. N. Engl. J. Med. 367, 1519–1528 (2012).
Li, Q. et al. Variants in TRIM22 that affect NOD2 signaling are associated with very-early-onset inflammatory bowel disease. Gastroenterology 150, 1196–1207 (2016).
Zenewicz, L.A., Abraham, C., Flavell, R.A. & Cho, J.H. Unraveling the genetics of autoimmunity. Cell 140, 791–797 (2010).
Farh, K.K. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Park, H. et al. A point mutation in the murine Hem1 gene reveals an essential role for Hematopoietic protein 1 in lymphopoiesis and innate immunity. J. Exp. Med. 205, 2899–2913 (2008).
Stevenson, C. et al. Essential role of Elmo1 in Dock2-dependent lymphocyte migration. J. Immunol. 192, 6062–6070 (2014).
Kadoya, M. et al. Allograft inflammatory factor-1 stimulates chemokine production and induces chemotaxis in human peripheral blood mononuclear cells. Biochem. Biophys. Res. Commun. 448, 287–291 (2014).
Dobbs, K. et al. Inherited DOCK2 deficiency in patients with early-onset invasive infections. N. Engl. J. Med. 372, 2409–2422 (2015).
Sakai, Y. et al. The Rac activator DOCK2 regulates natural killer cell–mediated cytotoxicity in mice through the lytic synapse formation. Blood 122, 386–393 (2013).
Gotoh, K. et al. Selective control of type I IFN induction by the Rac activator DOCK2 during TLR-mediated plasmacytoid dendritic cell activation. J. Exp. Med. 207, 721–730 (2010).
Watanabe, M. et al. DOCK2 and DOCK5 act additively in neutrophils to regulate chemotaxis, superoxide production, and extracellular trap formation. J. Immunol. 193, 5660–5667 (2014).
Sanui, T. et al. DOCK2 regulates Rac activation and cytoskeletal reorganization through interaction with ELMO1. Blood 102, 2948–2950 (2003).
Zhao, Y.Y., Yan, D.J. & Chen, Z.W. Role of AIF-1 in the regulation of inflammatory activation and diverse disease processes. Cell. Immunol. 284, 75–83 (2013).
Fukui, Y. et al. Haematopoietic cell–specific CDM family protein DOCK2 is essential for lymphocyte migration. Nature 412, 826–831 (2001).
Sirota, M., Schaub, M.A., Batzoglou, S., Robinson, W.H. & Butte, A.J. Autoimmune disease classification by inverse association with SNP alleles. PLoS Genet. 5, e1000792 (2009).
Giguère, P.M. et al. G protein signaling modulator-3 inhibits the inflammasome activity of NLRP3. J. Biol. Chem. 289, 33245–33257 (2014).
Yang, Z.F. et al. Allograft inflammatory factor-1 (AIF-1) is crucial for the survival and pro-inflammatory activity of macrophages. Int. Immunol. 17, 1391–1397 (2005).
Liu, G., Ma, H., Jiang, L. & Zhao, Y. Allograft inflammatory factor-1 and its immune regulation. Autoimmunity 40, 95–102 (2007).
Kim, S. et al. Characterizing the genetic basis of innate immune response in TLR4-activated human monocytes. Nat. Commun. 5, 5236 (2014).
Lösing, M. et al. The Dok-3/Grb2 protein signal module attenuates Lyn kinase-dependent activation of Syk kinase in B cell antigen receptor microclusters. J. Biol. Chem. 288, 2303–2313 (2013).
Ng, C.H., Xu, S. & Lam, K.P. Dok-3 plays a nonredundant role in negative regulation of B-cell activation. Blood 110, 259–266 (2007).
Ou, X., Xu, S., Li, Y.F. & Lam, K.P. Adaptor protein DOK3 promotes plasma cell differentiation by regulating the expression of programmed cell death 1 ligands. Proc. Natl. Acad. Sci. USA 111, 11431–11436 (2014).
Peng, Q., O'Loughlin, J.L. & Humphrey, M.B. DOK3 negatively regulates LPS responses and endotoxin tolerance. PLoS One 7, e39967 (2012).
Steinbach, E.C. & Plevy, S.E. The role of macrophages and dendritic cells in the initiation of inflammation in IBD. Inflamm. Bowel Dis. 20, 166–175 (2014).
Mortha, A. et al. Microbiota-dependent crosstalk between macrophages and ILC3 promotes intestinal homeostasis. Science 343, 1249288 (2014).
Onozawa, Y. et al. Activation of T cell death-associated gene 8 regulates the cytokine production of T cells and macrophages in vitro. Eur. J. Pharmacol. 683, 325–331 (2012).
Shenoy, A.R. et al. GBP5 promotes NLRP3 inflammasome assembly and immunity in mammals. Science 336, 481–485 (2012).
Soucie, E.L. et al. Lineage-specific enhancers activate self-renewal genes in macrophages and embryonic stem cells. Science 351, aad5510 (2016).
Gemperle, C. et al. Regulation of the formyl peptide receptor 1 (FPR1) gene in primary human macrophages. PLoS One 7, e50195 (2012).
Ma, C. et al. Receptor signaling lymphocyte-activation molecule family 1 (Slamf1) regulates membrane fusion and NADPH oxidase 2 (NOX2) activity by recruiting a Beclin-1/Vps34/ultraviolet radiation resistance-associated gene (UVRAG) complex. J. Biol. Chem. 287, 18359–18365 (2012).
Duong, B.H. et al. A20 restricts ubiquitination of pro-interleukin-1β protein complexes and suppresses NLRP3 inflammasome activity. Immunity 42, 55–67 (2015).
Glowacka, W.K., Alberts, P., Ouchida, R., Wang, J.Y. & Rotin, D. LAPTM5 protein is a positive regulator of proinflammatory signaling pathways in macrophages. J. Biol. Chem. 287, 27691–27702 (2012).
Present, D.H. et al. Infliximab for the treatment of fistulas in patients with Crohn's disease. N. Engl. J. Med. 340, 1398–1405 (1999).
Ito, H. Treatment of Crohn's disease with anti-IL-6 receptor antibody. J. Gastroenterol. 40 (Suppl. 16), 32–34 (2005).
Daryani, N.E. et al. Significance of IL-1RA polymorphism in Iranian patients with inflammatory bowel disease. Dig. Dis. Sci. 60, 1389–1395 (2015).
Mayer, L. et al. Anti-IP-10 antibody (BMS-936557) for ulcerative colitis: a phase II randomised study. Gut 63, 442–450 (2014).
Globig, A.M. et al. Comprehensive intestinal T helper cell profiling reveals specific accumulation of IFN-γ+IL-17+ coproducing CD4+ T cells in active inflammatory bowel disease. Inflamm. Bowel Dis. 20, 2321–2329 (2014).
Harbour, S.N., Maynard, C.L., Zindl, C.L., Schoeb, T.R. & Weaver, C.T. Th17 cells give rise to Th1 cells that are required for the pathogenesis of colitis. Proc. Natl. Acad. Sci. USA 112, 7061–7066 (2015).
Kinnebrew, M.A. et al. Interleukin 23 production by intestinal CD103+CD11b+ dendritic cells in response to bacterial flagellin enhances mucosal innate immune defense. Immunity 36, 276–287 (2012).
Scott, C.L., Aumeunier, A.M. & Mowat, A.M. Intestinal CD103+ dendritic cells: master regulators of tolerance? Trends Immunol. 32, 412–419 (2011).
Giguère, P.M. et al. G-protein signaling modulator-3, a gene linked to autoimmune diseases, regulates monocyte function and its deficiency protects from inflammatory arthritis. Mol. Immunol. 54, 193–198 (2013).
Park, H., Chan, M.M. & Iritani, B.M. Hem-1: putting the “WAVE” into actin polymerization during an immune response. FEBS Lett. 584, 4923–4932 (2010).
Seinen, M.L., van Nieuw Amerongen, G.P., de Boer, N.K. & van Bodegraven, A.A. Rac attack: modulation of the small GTPase Rac in inflammatory bowel disease and thiopurine therapy. Mol. Diagn. Ther. 20, 551–557 (2016).
Muise, A.M. et al. Single nucleotide polymorphisms that increase expression of the guanosine triphosphatase RAC1 are associated with ulcerative colitis. Gastroenterology 141, 633–641 (2011).
Kim, S.S. et al. DOK3 is required for IFN-β production by enabling TRAF3/TBK1 complex formation and IRF3 activation. J. Immunol. 193, 840–848 (2014).
Sironi, M. et al. An evolutionary analysis of RAC2 identifies haplotypes associated with human autoimmune diseases. Mol. Biol. Evol. 28, 3319–3329 (2011).
Fattouh, R. et al. Rac2-deficiency leads to exacerbated and protracted colitis in response to Citrobacter rodentium infection. PLoS One 8, e61629 (2013).
Muise, A.M. et al. NADPH oxidase complex and IBD candidate gene studies: identification of a rare variant in NCF2 that results in reduced binding to RAC2. Gut 61, 1028–1035 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G.K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
DeLuca, D.S. et al. RNA-SeQC: RNA-seq metrics for quality control and process optimization. Bioinformatics 28, 1530–1532 (2012).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Zhu, J. et al. An integrative genomics approach to the reconstruction of gene networks in segregating populations. Cytogenet. Genome Res. 105, 363–374 (2004).
Zhu, J. et al. Increasing the power to detect causal associations by combining genotypic and expression data in segregating populations. PLOS Comput. Biol. 3, e69 (2007).
Zhu, J. et al. Integrating large-scale functional genomic data to dissect the complexity of yeast regulatory networks. Nat. Genet. 40, 854–861 (2008).
Zhu, J. et al. Stitching together multiple data dimensions reveals interacting metabolomic and transcriptomic networks that modulate cell regulation. PLoS Biol. 10, e1001301 (2012).
Weiner, O.D. et al. Hem-1 complexes are essential for Rac activation, actin polymerization, and myosin regulation during neutrophil chemotaxis. PLoS Biol. 4, e38 (2006).
Weiner, O.D., Marganski, W.A., Wu, L.F., Altschuler, S.J. & Kirschner, M.W. An actin-based wave generator organizes cell motility. PLoS Biol. 5, e221 (2007).
Mathelier, A. et al. JASPAR 2016: a major expansion and update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 44, D110–D115 (2016).
Casimiro, I., Chinnasamy, P. & Sibinga, N.E. Genetic inactivation of the allograft inflammatory factor-1 locus. Genesis 51, 734–740 (2013).
Sanos, S.L. et al. RORγt and commensal microflora are required for the differentiation of mucosal interleukin 22–producing NKp46+ cells. Nat. Immunol. 10, 83–91 (2009).
Ivanov, I.I. et al. The orphan nuclear receptor RORγt directs the differentiation program of proinflammatory IL-17+ T helper cells. Cell 126, 1121–1133 (2006).
Wirtz, S., Neufert, C., Weigmann, B. & Neurath, M.F. Chemically induced mouse models of intestinal inflammation. Nat. Protoc. 2, 541–546 (2007).
Powrie, F., Leach, M.W., Mauze, S., Caddle, L.B. & Coffman, R.L. Phenotypically distinct subsets of CD4+ T cells induce or protect from chronic intestinal inflammation in C. B-17 scid mice. Int. Immunol. 5, 1461–1471 (1993).
Shumway, R.H. & Stoffer, D.S. Time Series Analysis and Its Applications (Springer, 2000).
Acknowledgements
We acknowledge P. Chinnasamy for backcrossing of Aif1−/− mice; E. Esplugues; K. Saulnier from Charles River Laboratories; S. Graham, J. Mena, and G. Lyng from Biomodels; K. Amin from Qiagen; E. Venturini and the New York Genome Center; M. Mahajan, Y. Kasai, and the Genome Core at Mount Sinai; H. Thomas; R. Ng; the Pathology Department and the Histology core at MSH; and Sinai Innovations. This work was supported in part through the computational resources and staff expertise provided by the Department of Scientific Computing at the Icahn School of Medicine at Mount Sinai. This work was funded by the Schadt laboratory at the Icahn Institute for Genomics and Multi-scale Biology, Icahn School of Medicine at Mount Sinai (NewYork). This work was partially funded by NIH/NIA grant R01AG046170 (to E.E.S. and B.Z.), a component of the AMP-AD Target Discovery and Preclinical Validation Project, the Rheumatology Research Foundation (to T.K.T.), the Leading Advanced Projects for Medical Innovation (LEAP; to Y.F.) from the Japan Agency for Medical Research and Development (AMED), U01HG008451 (to J.Z.), NIH R01HL128066 (to N.S.), and RO1 AI092093 and R21 AI109020 (to B.M.I.).
Author information
Authors and Affiliations
Contributions
L.A.P. and E.E.S. conceived of, designed, managed, and performed all analysis in the study. J.P. and E.M.N. performed macrophage knockdown experiments. W.-m.S. performed network enrichments. A.M. performed flow cytometry. A.I. performed histological scoring of mouse tissue, and S.R.L. and L.A.P. performed adoptive T cell transfer colitis experiments. L.A.P. designed and managed DSS experiments, A.D.N. and K.H. performed polygenic risk score and variant calling, A.D.N., K.H., and P.R. performed CRE SNP enrichments, B.A.K. generated differential expression signatures, Y.Z. generated clinical correlations, A.S. performed cell enrichments, J.S., S.E.T., W.-m.S., and Y.Z. performed statistical analysis on macrophage experiments, K.S., R.M., P.R., and L.A.P. constructed the eQTL database, B.L. and H.S. performed RNA-seq analysis, E.L. performed transcription factor analysis, M.W. and C.A. provided visualization tools, C.B., M.C., A.D., J.R.F., J.P., and L.F.M. provided patient population guidance, Y.F., M.B.H., B.M.I., N.S., and T.K.T. provided reagents and guidance, A.K., C.A., and J.J.F. provided project support, J.Z. constructed Bayesian networks, B.Z. constructed coexpression networks, and L.A.P. and E.E.S. wrote the manuscript. C.A., B.Z., J.P., J.R.F., and R.D. provided critical review of the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.P., E.M.N., A.S., J.S., S.E.T., C.B., M.C., A.D., J.R.F., and R.D. were employees of Janssen during the time this work was completed.
Integrated supplementary information
Supplementary Figure 1 Distribution of the IBD polygenic score based on 86 SNPs represented in the three IBD populations (from top to bottom): RISK, CERTIFI, and MSH.
To assess whether the mean polygenic scores for each pair of populations were significantly different, we employed a pairwise t test. None of the tests were significant at a nominal 0.05 P-value threshold: the P value for CERTIFI versus MSH was 0.71; for RISK versus MSH the P value was 0.29; and for RISK versus CERTIFI the P value was 0.40.
Supplementary Figure 2 Heat map of the odds ratios of overlaps between gene lists by cell type (rows) and inflamed anatomical region (columns) in CD (top) and UC (bottom).
The full list of genes reported in any cell type and/or region was used as the background. Overlaps with Fisher’s test P < 0.001 are marked with a black circle. Log-odds ratios quantify the overlap between different anatomical regions (in columns) and cell types (in rows) within disease conditions (horizontal panels). We observed, for example, a significant overlap between the myeloid and rectum gene lists within patients with CD but not UC. Included in the heat map is ileum, plus any cell type and anatomical region with at least one overlap with significance, P < 0.001.
Supplementary Figure 3 Extent of KDG mRNA knockdown.
(a–k) Three siRNA, mock, and non-target control experiments were performed per KDG target, including DOK3 (a), FPR1 (b), TNFAIP3 (c), LAPTM5 (d), SLAMF1 (e), AIF1 (f), NCKAP1L (g), GPR65 (h), GBP5 (i), MAFB (j), and GPSM3 (k), in human primary monocyte-derived macrophages with and without LPS stimulation. Data are representative of duplicate or triplicate samples per experiment.
Supplementary Figure 4 Effect of KDG knockdown in macrophages on cytokine expression.
(a–h) Differential production of cytokines following siRNA-mediated knockdown of KDGs: IL-6 (a), IL-10 (b), TNF-α (c), IL-12p40 (d), IL-1RA (ILRN) (e), MCP-3 (CCL7) (f), MIP-1β (CCL4) (g), and IP-10 (CXCL10) (h). Comparisons were between LPS-treated siRNA versus LPS-treated non-targeting control siRNA cells. Three donors were tested for each siRNA in three separate experiments with two or three replicates per experiment.
Supplementary Figure 5 The KDG module and network structure are conserved across species.
(a–c) Overlap of the mouse brown coexpression module with the CIC subnetwork of the MSH IBD network (184 genes) (a), the CERTIFI IBD network (191 genes) (b), and the RISK IBD network (308 genes) (c). Intestine KDGs are shown as red diamonds. Macrophage KDGs are shown as green triangles.
Supplementary Figure 6 Network validation: significant enrichment of KDG perturbation signatures in various IBD Bayesian networks.
Mouse intestine KDG knockout versus wild-type control interaction with DSS differential expression signature –log10 Fisher’s exact test P-value enrichment within a two-path-length neighborhood of each mouse experimental KDG in the CERTIFI, MSH, and RISK IBD networks.
Supplementary Figure 7 Enrichment of mouse and human transcription factors in KDG subnetworks on each IBD network.
These are transcription factors for regulation of genes in each subnetwork. (a) In monocyte results based on DNase I hypersenstivity regions at a significance threshold of P = 10−4. (b) In T cell ENCODE data at a significance threshold of P = 10−5.
Supplementary Figure 8 Colon weight/length ratio of KDG-knockout mice following treatment.
(a) Gpsm3 DSS: 9 Gpsm3−/− and 8 WT male mice. Pooled data representative of two independent experiments are shown. (b) Dock2 TNBS: 3 Dock2−/− and 6 WT and female mice. Data are representative of one of two independent experiments. An unpaired t test was performed. Data are expressed as ±s.e.m. Statistical significance is indicated: *P < 0.05, **P < 0.01, ***P < 0.001.
Supplementary Figure 9 Stool scores from KDG-knockout mice from DSS (day 7) or TNBS.
(a) Dock2 DSS: 6 Dock2−/− and 5 WT mice. (b) Dock2 TNBS: 2 Dock2−/− and 6 WT mice. The data in a and b are representative of a single experiment. (c) Dok3 DSS: 20 Dok3−/− and 20 WT mice per group. Pooled data representative of two independent experiments are shown. (d) Gpsm3 DSS: 3 Gpsm3−/− and 3 WT mice per group. Data are representative of one of two independent experiments. An unpaired two-tailed Student's t test was performed. Data are expressed as ±s.e.m. Statistical significance is indicated: *P < 0.05, **P < 0.01, ***P < 0.001.
Supplementary Figure 10 Differential weight loss from the T cell adoptive transfer colitis model.
(a–d) C57BL/6 Rag−/− mice were transferred with cells from Aif1−/− (a), Gpsm3−/− (b), Dock2−/− (c), Nckap1l−/− (d), or respective wild-type littermate control mice. Data are expressed as ±s.e.m. Comparisons were performed using an autoregressive model to maximize use of the time series data. Data are expressed as mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Note (PDF 3122 kb)
Supplementary Tables
Supplementary Tables 1–38 (XLSX 5006 kb)
Rights and permissions
About this article
Cite this article
Peters, L., Perrigoue, J., Mortha, A. et al. A functional genomics predictive network model identifies regulators of inflammatory bowel disease. Nat Genet 49, 1437–1449 (2017). https://doi.org/10.1038/ng.3947
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3947
This article is cited by
-
Bone marrow-derived mesenchymal stem cells mitigate chronic colitis and enteric neuropathy via anti-inflammatory and anti-oxidative mechanisms
Scientific Reports (2024)
-
A precise molecular subtyping of ulcerative colitis reveals the immune heterogeneity and predicts clinical drug responses
Journal of Translational Medicine (2023)
-
A temporal classifier predicts histopathology state and parses acute-chronic phasing in inflammatory bowel disease patients
Communications Biology (2023)
-
Predictive network analysis identifies JMJD6 and other potential key drivers in Alzheimer’s disease
Communications Biology (2023)
-
Inflammatory Bowel Disease: Pathophysiology, Treatment, and Disease Modeling
BioChip Journal (2023)