Abstract
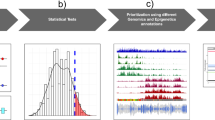
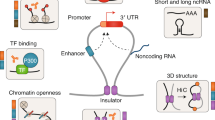
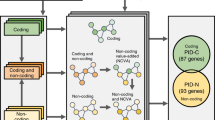
The contributions of coding mutations to tumorigenesis are relatively well known; however, little is known about somatic alterations in noncoding DNA. Here we describe GECCO (Genomic Enrichment Computational Clustering Operation) to analyze somatic noncoding alterations in 308 pancreatic ductal adenocarcinomas (PDAs) and identify commonly mutated regulatory regions. We find recurrent noncoding mutations to be enriched in PDA pathways, including axon guidance and cell adhesion, and newly identified processes, including transcription and homeobox genes. We identified mutations in protein binding sites correlating with differential expression of proximal genes and experimentally validated effects of mutations on expression. We developed an expression modulation score that quantifies the strength of gene regulation imposed by each class of regulatory elements, and found the strongest elements were most frequently mutated, suggesting a selective advantage. Our detailed single-cancer analysis of noncoding alterations identifies regulatory mutations as candidates for diagnostic and prognostic markers, and suggests new mechanisms for tumor evolution.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Siegel, R., Naishadham, D. & Jemal, A. Cancer statistics, 2013. CA Cancer J. Clin. 63, 11–30 (2013).
Jones, S. et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 321, 1801–1806 (2008).
Biankin, A.V. et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 491, 399–405 (2012).
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518, 495–501 (2015).
Huang, F.W. et al. Highly recurrent TERT promoter mutations in human melanoma. Science 339, 957–959 (2013).
Horn, S. et al. TERT promoter mutations in familial and sporadic melanoma. Science 339, 959–961 (2013).
Bell, R.J. et al. The transcription factor GABP selectively binds and activates the mutant TERT promoter in cancer. Science 348, 1036–1039 (2015).
Killela, P.J. et al. TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc. Natl. Acad. Sci. USA 110, 6021–6026 (2013).
Rachakonda, P.S. et al. TERT promoter mutations in bladder cancer affect patient survival and disease recurrence through modification by a common polymorphism. Proc. Natl. Acad. Sci. USA 110, 17426–17431 (2013).
Mansour, M.R. et al. Oncogene regulation. An oncogenic super-enhancer formed through somatic mutation of a noncoding intergenic element. Science 346, 1373–1377 (2014).
Weinhold, N., Jacobsen, A., Schultz, N., Sander, C. & Lee, W. Genome-wide analysis of noncoding regulatory mutations in cancer. Nat. Genet. 46, 1160–1165 (2014).
Fredriksson, N.J., Ny, L., Nilsson, J.A. & Larsson, E. Systematic analysis of noncoding somatic mutations and gene expression alterations across 14 tumor types. Nat. Genet. 46, 1258–1263 (2014).
Melton, C., Reuter, J.A., Spacek, D.V. & Snyder, M. Recurrent somatic mutations in regulatory regions of human cancer genomes. Nat. Genet. 47, 710–716 (2015).
Mathelier, A. et al. Cis-regulatory somatic mutations and gene-expression alteration in B-cell lymphomas. Genome Biol. 16, 84 (2015).
Araya, C.L. et al. Identification of significantly mutated regions across cancer types highlights a rich landscape of functional molecular alterations. Nat. Genet. 48, 117–125 (2016).
Fujimoto, A. et al. Whole-genome mutational landscape and characterization of noncoding and structural mutations in liver cancer. Nat. Genet. 48, 500–509 (2016).
Hudson, T.J. et al. International network of cancer genome projects. Nature 464, 993–998 (2010).
Khurana, E. et al. Integrative annotation of variants from 1092 humans: application to cancer genomics. Science 342, 1235587 (2013).
Fu, Y. et al. FunSeq2: a framework for prioritizing noncoding regulatory variants in cancer. Genome Biol. 15, 480 (2014).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Teng, Y., Mei, Y., Hawthorn, L. & Cowell, J.K. WASF3 regulates miR-200 inactivation by ZEB1 through suppression of KISS1 leading to increased invasiveness in breast cancer cells. Oncogene 33, 203–211 (2014).
Winham, S.J. et al. Genome-wide investigation of regional blood-based DNA methylation adjusted for complete blood counts implicates BNC2 in ovarian cancer. Genet. Epidemiol. 38, 457–466 (2014).
Dulak, A.M. et al. Exome and whole-genome sequencing of esophageal adenocarcinoma identifies recurrent driver events and mutational complexity. Nat. Genet. 45, 478–486 (2013).
Sherman, S.K. et al. Gastric inhibitory polypeptide receptor (GIPR) is a promising target for imaging and therapy in neuroendocrine tumors. Surgery 154, 1206–1213, discussion 1214 (2013).
Uzawa, K. et al. Targeting phosphodiesterase 3B enhances cisplatin sensitivity in human cancer cells. Cancer Med. 2, 40–49 (2013).
Renjie, W. & Haiqian, L. MiR-132, miR-15a and miR-16 synergistically inhibit pituitary tumor cell proliferation, invasion and migration by targeting Sox5. Cancer Lett. 356, 568–578 (2015).
Sandelin, A., Alkema, W., Engström, P., Wasserman, W.W. & Lenhard, B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res. 32, D91–D94 (2004).
Flandin, P. et al. Lhx6 and Lhx8 coordinately induce neuronal expression of Shh that controls the generation of interneuron progenitors. Neuron 70, 939–950 (2011).
Boon, M.R. et al. Bone morphogenetic protein 7: a broad-spectrum growth factor with multiple target therapeutic potency. Cytokine Growth Factor Rev. 22, 221–229 (2011).
Gutschner, T. et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 73, 1180–1189 (2013).
Moriyama, T. et al. MicroRNA-21 modulates biological functions of pancreatic cancer cells including their proliferation, invasion, and chemoresistance. Mol. Cancer Ther. 8, 1067–1074 (2009).
Cheung, H.W. et al. Systematic investigation of genetic vulnerabilities across cancer cell lines reveals lineage-specific dependencies in ovarian cancer. Proc. Natl. Acad. Sci. USA 108, 12372–12377 (2011).
Lan, Q. et al. Genetic susceptibility for chronic lymphocytic leukemia among Chinese in Hong Kong. Eur. J. Haematol. 85, 492–495 (2010).
Sun, H.T., Cheng, S.X., Tu, Y., Li, X.H. & Zhang, S. FoxQ1 promotes glioma cells proliferation and migration by regulating NRXN3 expression. PLoS One 8, e55693 (2013).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Mascarenhas, J.B. et al. PAX6 is expressed in pancreatic cancer and actively participates in cancer progression through activation of the MET tyrosine kinase receptor gene. J. Biol. Chem. 284, 27524–27532 (2009).
Segara, D. et al. Expression of HOXB2, a retinoic acid signaling target in pancreatic cancer and pancreatic intraepithelial neoplasia. Clin. Cancer Res. 11, 3587–3596 (2005).
Chile, T. et al. HOXB7 mRNA is overexpressed in pancreatic ductal adenocarcinomas and its knockdown induces cell cycle arrest and apoptosis. BMC Cancer 13, 451 (2013).
Whittle, M.C. et al. RUNX3 controls a metastatic switch in pancreatic ductal adenocarcinoma. Cell 161, 1345–1360 (2015).
Than, B.L. et al. The role of KCNQ1 in mouse and human gastrointestinal cancers. Oncogene 33, 3861–3868 (2014).
Geimer Le Lay, A.S. et al. The tumor suppressor Ikaros shapes the repertoire of notch target genes in T cells. Sci. Signal. 7, ra28 (2014).
Anglim, P.P. et al. Identification of a panel of sensitive and specific DNA methylation markers for squamous cell lung cancer. Mol. Cancer 7, 62 (2008).
Benetatos, L. et al. CpG methylation analysis of the MEG3 and SNRPN imprinted genes in acute myeloid leukemia and myelodysplastic syndromes. Leuk. Res. 34, 148–153 (2010).
Kelley, D.R., Snoek, J. & Rinn, J.L. Basset: learning the regulatory code of the accessible genome with deep convolutional neural networks. Genome Res. 26, 990–999 (2016).
Squazzo, S.L. et al. Suz12 binds to silenced regions of the genome in a cell-type-specific manner. Genome Res. 16, 890–900 (2006).
Weirauch, M.T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Gupta, S., Stamatoyannopoulos, J.A., Bailey, T.L. & Noble, W.S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Acknowledgements
We thank the members of the Tuveson laboratory, C. Vakoc and A. Siepel for discussions. D.A.T. is a distinguished scholar of the Lustgarten Foundation and Director of the Lustgarten Foundation-designated Laboratory of Pancreatic Cancer Research. D.A.T. is also supported by the Cold Spring Harbor Laboratory Association, the V Foundation, PCUK and the David Rubinstein Center for Pancreatic Cancer Research at MSKCC. In addition, we are grateful for support from the following: the STARR Foundation (I7-A718 for D.A.T.), DOD (W81XWH-13-PRCRP-IA for D.A.T.), Louis Morin Charitable Trust (M.E.F.) and NIH (5P30CA45508-26, 5P50CA101955-07, 1U10CA180944-03, 5U01CA168409-5, 1R01CA188134-01A1 and 1R01CA190092-03 for D.A.T. and R01HG006677 for M.C.S.).
Author information
Authors and Affiliations
Contributions
M.E.F., T.G., M.C.S. and D.A.T. wrote the manuscript. M.C.S. and D.A.T. supervised the study. T.G. performed FunSeq analysis and developed GECCO. M.E.F. performed pathway analysis. M.E.F., T.G., S.M.G., A.V.B., E.K., S.S., L.D.S., S.G. and J.D.M. contributed to data analysis. D.K.C. and P.B. performed patient outcome analysis. D.R.K. performed Basset analysis. N.W. performed germline sequence analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Identification of recurrent noncoding mutations in PDA.
Distribution of SNV rates across the patient cohort.
Supplementary Figure 2 Overlap of SNVs and common coding mutations in PDA.
Distribution of SNVs across the patient cohort, with common coding mutations (colored bars) in PDA genes.
Supplementary Figure 3 Overlap of gene-proximal NCMs in CRRs and common coding mutations in PDA.
Distribution of CRR mutation rates across the patient cohort, with common coding mutations (colored bars) in PDA genes.
Supplementary Figure 4 NCMs disrupt transcription factor binding motifs.
(a) A G→A mutation in a regulatory site on chromosome 15 at position 25,200,056 alters a critical nucleotide in an NRF1 binding site. The regulatory site lies in the promoter of SNRPN. At the bottom, the heat map displays the predicted change in binding, considered here as ChIP-seq signal for NRF1 in H1-hESCs. The line plots above measure the maximum (gain) and minimum (loss) predicted change; the loss highlights nucleotides that significantly alter the overall signal upon mutation as this mutation does. (b) A G→T mutation in a regulatory site on chromosome 3 at position 115,757,580 introduces a GATA factor binding site nearby an established PU.1 binding site. The heat map displays the predicted change in accessibility, considered here as DNase-seq signal in K562. In other cells, such as monocytes, the model predicts reduced accessibility, suggesting that GATA binding here may alter the combinatorial logic of the regulatory element in a complex fashion.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Tables 1–4 and Supplementary Note (PDF 2660 kb)
Rights and permissions
About this article
Cite this article
Feigin, M., Garvin, T., Bailey, P. et al. Recurrent noncoding regulatory mutations in pancreatic ductal adenocarcinoma. Nat Genet 49, 825–833 (2017). https://doi.org/10.1038/ng.3861
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3861
This article is cited by
-
Nutrient transporters: connecting cancer metabolism to therapeutic opportunities
Oncogene (2023)
-
ATP13A3 facilitates polyamine transport in human pancreatic cancer cells
Scientific Reports (2022)
-
Pancreatic cancer evolution and heterogeneity: integrating omics and clinical data
Nature Reviews Cancer (2022)
-
The molecular biology of pancreatic adenocarcinoma: translational challenges and clinical perspectives
Signal Transduction and Targeted Therapy (2021)
-
The pancreatic cancer genome revisited
Nature Reviews Gastroenterology & Hepatology (2021)