Abstract
Facioscapulohumeral dystrophy (FSHD; MIM158900, MIM158901) is caused by misexpression of the DUX4 transcription factor in skeletal muscle1. Animal models of FSHD are hindered by incomplete knowledge regarding the conservation of the DUX4 transcriptional program in other species2,3,4,5. Despite the divergence of their binding motifs, both mouse DUX and human DUX4 in mouse and human muscle cells, respectively, activate genes associated with cleavage-stage embryos, including MERVL and ERVL–MaLR retrotransposons. We found that human DUX4 expressed in mouse cells maintained modest activation of cleavage-stage genes driven by conventional promoters but did not activate MERVL-promoted genes. Thus, the ancestral DUX4-regulated genes are characteristic of cleavage-stage embryos and are driven by conventional promoters, whereas divergence of the DUX4 and DUX homeodomains correlates with retrotransposon specificity. These results provide insight into how species balance conservation of a core transcriptional program with innovation at retrotransposon promoters, and establish a basis for animal models recreating the FSHD transcriptome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Tawil, R., van der Maarel, S.M. & Tapscott, S.J. Facioscapulohumeral dystrophy: the path to consensus on pathophysiology. Skelet. Muscle 4, 12 (2014).
Lek, A., Rahimov, F., Jones, P.L. & Kunkel, L.M. Emerging preclinical animal models for FSHD. Trends Mol. Med. 21, 295–306 (2015).
Wallace, L.M. et al. DUX4, a candidate gene for facioscapulohumeral muscular dystrophy, causes p53-dependent myopathy in vivo. Ann. Neurol. 69, 540–552 (2011).
Krom, Y.D. et al. Intrinsic epigenetic regulation of the D4Z4 macrosatellite repeat in a transgenic mouse model for FSHD. PLoS Genet. 9, e1003415 (2013).
Dandapat, A. et al. Dominant lethal pathologies in male mice engineered to contain an X-linked DUX4 transgene. Cell Rep. 8, 1484–1496 (2014).
Geng, L.N. et al. DUX4 activates germline genes, retroelements, and immune mediators: implications for facioscapulohumeral dystrophy. Dev. Cell 22, 38–51 (2012).
Young, J.M. et al. DUX4 binding to retroelements creates promoters that are active in FSHD muscle and testis. PLoS Genet. 9, e1003947 (2013).
Bosnakovski, D., Daughters, R.S., Xu, Z., Slack, J.M. & Kyba, M. Biphasic myopathic phenotype of mouse DUX, an ORF within conserved FSHD-related repeats. PLoS One 4, e7003 (2009).
Leidenroth, A. et al. Evolution of DUX gene macrosatellites in placental mammals. Chromosoma 121, 489–497 (2012).
Leidenroth, A. & Hewitt, J.E. A family history of DUX4: phylogenetic analysis of DUXA, B, C and Duxbl reveals the ancestral DUX gene. BMC Evol. Biol. 10, 364 (2010).
Clapp, J. et al. Evolutionary conservation of a coding function for D4Z4, the tandem DNA repeat mutated in facioscapulohumeral muscular dystrophy. Am. J. Hum. Genet. 81, 264–279 (2007).
Eidahl, J.O. et al. Mouse Dux is myotoxic and shares partial functional homology with its human paralog DUX4. Hum. Mol. Genet. 25, 4577–4589 (2016).
Knopp, P. et al. DUX4 induces a transcriptome more characteristic of a less-differentiated cell state and inhibits myogenesis. J. Cell Sci. 129, 3816–3831 (2016).
Falco, G. et al. Zscan4: a novel gene expressed exclusively in late 2-cell embryos and embryonic stem cells. Dev. Biol. 307, 539–550 (2007).
Zhang, W. et al. Zfp206 regulates ES cell gene expression and differentiation. Nucleic Acids Res. 34, 4780–4790 (2006).
Macfarlan, T.S. et al. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487, 57–63 (2012).
Akiyama, T. et al. Transient bursts of Zscan4 expression are accompanied by the rapid derepression of heterochromatin in mouse embryonic stem cells. DNA Res. 22, 307–318 (2015).
Jagannathan, S. et al. Model systems of DUX4 expression recapitulate the transcriptional profile of FSHD cells. Hum. Mol. Genet. 25, 4419–4431 (2016).
NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 44, D7–D19 (2016).
Bailey, T.L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Noyes, M.B. et al. Analysis of homeodomain specificities allows the family-wide prediction of preferred recognition sites. Cell 133, 1277–1289 (2008).
Peaston, A.E. et al. Retrotransposons regulate host genes in mouse oocytes and preimplantation embryos. Dev. Cell 7, 597–606 (2004).
Hendrickson, P.G. et al. Conserved roles of mouse DUX and human DUX4 in activating cleavage-stage genes and MERVL/HERVL retrotransposons. Nat. Genet. http://dx.doi.org/10.1038/ng.3844 (2017).
Bosnakovski, D. et al. An isogenetic myoblast expression screen identifies DUX4-mediated FSHD-associated molecular pathologies. EMBO J. 27, 2766–2779 (2008).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Reich, M. et al. GenePattern 2.0. Nat. Genet. 38, 500–501 (2006).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Mi, H., Poudel, S., Muruganujan, A., Casagrande, J.T. & Thomas, P.D. PANTHER version 10: expanded protein families and functions, and analysis tools. Nucleic Acids Res. 44 D1, D336–D342 (2016).
Conerly, M.L., Yao, Z., Zhong, J.W., Groudine, M. & Tapscott, S.J. Distinct activities of Myf5 and MyoD indicate separate roles in skeletal muscle lineage specification and differentiation. Dev. Cell 36, 375–385 (2016).
Cao, Y. et al. Genome-wide MyoD binding in skeletal muscle cells: a potential for broad cellular reprogramming. Dev. Cell 18, 662–674 (2010).
Liao, Y., Smyth, G.K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108 (2013).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ma, W., Noble, W.S. & Bailey, T.L. Motif-based analysis of large nucleotide data sets using MEME-ChIP. Nat. Protoc. 9, 1428–1450 (2014).
Bailey, T.L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Edgar, R., Domrachev, M. & Lash, A.E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Acknowledgements
We thank S. Shadle (Fred Hutchinson Cancer Research Center) for making and sharing the pCW57.1-DUX4 virus-like particles used to generate the DUX4-inducible C2C12 clonal cell lines; P. Hendrickson (Huntsman Cancer Institute) for 2C::TdTomato Reporter (Addgene plasmid no. 40281), from which we cloned the MERVL used to create the MERVL reporter in this study; and D. Root (Broad Institute) for the gift of Addgene plasmid no. 41393. We thank G. Kardon for critical guidance in the project and also thank members of the research groups of S. Tapscott and H. Malik for helpful discussions and critical reading of the manuscript. This work was supported by NIH NIAMS R01AR045203 (S.J.T.), National Science Foundation Graduate Research Fellowship Program DGE-1256082 (J.L.W.), University of Washington Interdisciplinary Training in Genome Sciences T32 HG00035 from the National Human Genome Research Institute (J.L.W.), and Friends of FSH Research (J.L.W. and S.J.T.).
Author information
Authors and Affiliations
Contributions
J.L.W. and S.J.T. conceived and designed the study and wrote the manuscript, with the assistance and final approval of all authors. J.L.W. performed and interpreted RNA-seq, ChIP–seq, RT–qPCR, and luciferase experiments. A.T.L. performed RT–qPCR and luciferase experiments. C.-J.W. performed bioinformatic data analyses. J.W.Z. contributed to the creation and characterization of the MMH-chimera cell line.
Corresponding author
Ethics declarations
Competing interests
J.L.W. and S.J.T., together with B. Cairns, E. Grow, and P. Hendrickson, have filed a provisional patent application, ‘Compositions and methods for reprogramming cells and for somatic cell nuclear transfer using DUXC expression’ (US provisional application no. 62/410,078, US Patent and Trademark Office), which is based in part on this work.
Integrated supplementary information
Supplementary Figure 1 Negative control for GSEA.
(a) As a critical negative control, we used GSEA to assess enrichment of the 2C-like state gene signature in a transcriptome where one does not expect to find enrichment. The transcriptome we used was a published dataset representing the MyoD transcriptome when expressed lentivirally in mouse embryonic fibroblasts1. MyoD has no known role in the 2C mouse embryo, rather it is the master regulator of muscle lineage specification2,3,4. That this graph peaks near the center of the x-axis indicates that the majority of the 2C-like state genes are unaffected by MyoD (vertical hash mark). This contrasts distinctly with the taller, left-shifted peak seen in Figure 1b, for example. GSEA p-values were determined by permuting the transcriptome 1,000 times, hence our report of "p-value<0.001". 1. Conerly, M.L., Yao, Z., Zhong, J.W., Groudine, M. & Tapscott, S.J. Distinct Activities of Myf5 and MyoD Indicate Separate Roles in Skeletal Muscle Lineage Specification and Differentiation. Dev Cell 36, 375-85 (2016). 2. Choi, J. et al. MyoD converts primary dermal fibroblasts, chondroblasts, smooth muscle, and retinal pigmented epithelial cells into striated mononucleated myoblasts and multinucleated myotubes. Proc Natl Acad Sci U S A 87, 7988-92 (1990). 3. Davis, R.L., Weintraub, H. & Lassar, A.B. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 51, 987-1000 (1987). 4. Weintraub, H. et al. Activation of muscle-specific genes in pigment, nerve, fat, liver, and fibroblast cell lines by forced expression of MyoD. Proc Natl Acad Sci U S A 86, 5434-8 (1989).
Supplementary Figure 2 Pramef25 is a direct target of Dux.
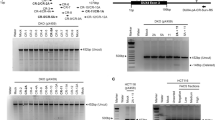
(a) ChIP-seq and RNA-seq coverage near the Pramef25 locus. Black rectangle shows location of 750bp sequence (mm10; chr4: 143,954,684-143,955,431) that was synthesized and cloned upstream of firefly luciferase to create the Pramef25 reporter. Note that Dux regulates an upstream, unannotated start site of Pramef25. Find Individual Motif Occurrences (FIMO) identified three Dux binding motifs that overlap the Pramef25 reporter region. Figure prepared with UCSC Genome Browser5; track heights given in square brackets are read counts. (b) Luciferase assay comparing the activation of the Pramef25 reporter by either Dux or an empty vector. The original sequences of three predicted Dux binding motifs and the sequences to which they were mutated are shown in cartoon to the right. Data shown are mean fold change over empty vector of 3 cell cultures prepared in parallel for each condition. Error bars are s.e.m. The non-mutated Pramef25 reporter activation experiment was repeated on three separate occasions with consistent results. The mutated Pramef25 reporter experiment was performed on one occasion. 5. Kent, W.J. et al. The human genome browser at UCSC. Genome Res 12, 996-1006 (2002).
Supplementary Figure 3 Zscan4c is a direct target of Dux, and each Zscan4-cluster gene contains a Dux ChIP–seq peak at its TSS.
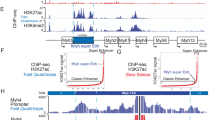
(a) ChIP-seq and RNA-seq coverage near the Zscan4c locus. Black rectangle shows location of 450bp sequence (chr7:11,005,309-11,005,758) that was synthesized and cloned upstream of luciferase to create the Zscan4c reporter. Find Individual Motif Occurrences (FIMO) identified four Dux binding motifs that overlap the Zscan4c reporter region. Figure prepared with Integrative Genomics Viewer6,7; track heights given in square brackets are read counts. (b) Luciferase assay comparing the activation of the Zscan4c reporter by either Dux or an empty vector. The original sequences of four predicted Dux binding motifs and the sequences to which they were mutated are shown in cartoon to the right. Data shown are mean fold change over empty vector of 3 cell cultures prepared in parallel for each condition. Error bars are s.e.m. The non-mutated Zscan4c reporter activation experiment was repeated on three separate occasions with consistent results. The mutated Zscan4c reporter experiment was performed on two occasions. (c) UCSC genome browser shot of Zscan4 cluster, showing RNA-seq and ChIP-seq coverage tracks. FIMO track shows locations of predicted Dux binding motifs and MERV-L track shows RepeatMasker MT2_Mm and MERVL-int locations. mm10 genomic coordinates: chr7:10,788,877-11,408,611. Note: The two loci with RNA-seq and ChIP-seq coverage in the absence of a UCSC-annotated Zscan4 gene are annotated as Zscan4 genes in other annotation models. (d) UCSC genome browser shot of Zscan4a, mm10 genomic coordinates: chr7:10792200-10801100. (e) UCSC genome browser shot of Zscan4b, mm10 genomic coordinates: chr7:10898700-10907000. (f) UCSC genome browser shot of Zscan4c and a MERV-L, mm10 genomic coordinates: chr7:11003700-11030500. We did not find any RNA-seq reads that support splicing between this MERV-L and Zscan4c. (g) UCSC genome browser shot of Zscan4d and a MERV-L, mm10 genomic coordinates: chr7:11159600-11186100. We did not find any RNA-seq reads that support splicing between this MERV-L and Zscan4d. (h) UCSC genome browser shot of Zscan4f, mm10 genomic coordinates: chr7:11395900-11404300. (i) UCSC genome browser shot of MERV-L downstream of Zscan4c (mm10 genomic coordinates: chr7: 11,019,863-11,029,599), zoomed in and rescaled to show ChIP-seq peaks at the LTR portion of the element and RNA-seq read coverage of the internal sequence. Note the scale differences between panels 3i-j and the remainder of Supplementary Figure 3. (j) UCSC genome browser shot of MERV-L upstream of Zscan4d (mm10 genomic coordinates: chr7: 11,168,315-11,178,031), zoomed in and rescaled to show ChIP-seq peaks and RNA-seq read coverage. Note the scale differences in panels 3i-j and the remainder of Supplementary Figure 3. 6. Robinson, J.T. et al. Integrative genomics viewer. Nat Biotechnol 29, 24-6 (2011). 7. Thorvaldsdottir, H., Robinson, J.T. & Mesirov, J.P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14, 178-92 (2013).
Supplementary Figure 4 Reciprocal GSEA showing that Dux and DUX4 activate orthologous genes in their respective species.
(a) Making the converse comparison as the graph in main text Figure 1d, this GSEA shows that the 500 genes most upregulated by Dux were significantly enriched in the genes most upregulated by DUX4. The x-axis is the log2FoldChange-ranked DUX4 transcriptome. This analysis compared Dux-expressing mouse cells to DUX4-expressing human cells. Since this comparison is between species, we limited both gene set and transcriptome to genes with simple 1:1 mouse-to-human orthologs.
Supplementary Figure 5 RNA-seq and ChIP–seq data for DUX4 expressed in mouse muscle cells.
(a) Comparison of DUX4 binding motifs in mouse and human muscle cells as determined by MEME. E-values listed reflect an estimate of the expected number of motifs, with the given motif's log likelihood ratio (or higher) and with the same width and site count, that one would find in a similarly sized set of random sequences (sequences where each position is independent and letters are chosen according to the background letter frequencies). (b) DUX4 transcriptome in mouse muscle cells as determined by comparing induced and un-induced cell cultures. Red dots are genes affected more than absolute(log2FoldChange)>=2 and adjusted p-value<=0.05. (c) Comparison of transcriptome induced by DUX4 and Dux in mouse muscle cells. Genes absent from one dataset, but present in the other dataset, were set to log2FoldChange of 0 in order to display on the graph. Spearman's rank correlation coefficient is 0.1569. (d) This is a subset of graph in Supplementary Figure 5c; each dot is a gene that contributed to the core enrichment of the 2C-like gene signature in the Dux transcriptome. MERV-L-promoted genes that were bound and activated by each factor are highlighted as determined by the "Peak-Associated Gene" method. Note that DUX4 did not bind and activate any of the 2C-like MERV-L promoted genes and thus there are no black dots on the graph.
Supplementary Figure 6 Distribution of transcribed LTR repeats according to repFamily.
(a) Expression levels of repeats during Dux expression in mouse cells compared to un-induced cells of the same cell line, broken down by repeat family. Each dot is a repeatName as defined by RepeatMasker. Red color indicates differential expression at absolute(log2-Foldchange)>=1 and adjusted p-value<=0.05. (b) DUX4-expressing mouse muscle cells compared to un-induced cells of the same cell line. (c) Re-analyzed data from DUX4-expressing human muscle cells compared to un-induced cells of the same cell line. (d) The MERVL_LTR consensus sequence from RepBase carries a match to the Dux binding motif (q-value = 0.0132, determined by FIMO).
Supplementary Figure 7 Browser screenshots of peak-associated genes in the 2C-like signature that start in MERV-L elements.
(a) AF067061. Note that we defined "Peak-associated genes" algorithmically as genes that have a ChIP-seq peak and at least one RNA-seq read that connects the peak location to an annotated exon of the gene. All RNA-seq tracks in this panel have 10,500 read track height. All ChIP-seq tracks in this panel have 153 read track height. (b) B020004J07Rik. All RNA-seq tracks in this panel have 550 read track height. All ChIP-seq tracks in this panel have 90 read track height. (c) Gm8994. All RNA-seq tracks in this panel have 175 read track height. All ChIP-seq tracks in this panel have 80 read track height.
Supplementary Figure 8 Distribution of ChIP–seq peak locations according to repeat family in mouse muscle cells expressing either Dux or DUX4.
(a) Stacked bar chart shows the distribution of ChIP-seq peak locations for the top 10,000 peaks for each condition. Dux ChIP-seq peaks occurred 2.4-fold more often in LTR elements than expected if these binding sites were evenly distributed across the genome; ERVL elements contributed the most to this overrepresentation with 4.2-fold more peaks in ERVL than expected by chance (see Panel C). DUX4 binding sites were 1.5-fold overrepresented in LTR elements in mouse cells and ERVL-MaLR elements contributed the most to this enrichment with 2.6-fold more peaks in ERVL-MaLR than expected by chance. Note that the vast majority of DUX4-bound ERVL-MaLRs are not shared with Dux. Only 4% of bound ERVL-MaLRs are shared (334/ 8027 total peak locations). Shown for comparison is DUX4 ChIP-seq peak distribution in human muscle cells, based on re-analysis of previously published data to match computational methods of this study8,9. (b) Grouped bar chart shows the fold enrichment for the ChIP-seq peak distribution shown in (a) compared to genomic distribution of each LTR family as reported by RepeatMasker. 8. Geng, L.N. et al. DUX4 activates germline genes, retroelements, and immune mediators: implications for facioscapulohumeral dystrophy. Dev Cell 22, 38-51 (2012). 9. Young, J.M. et al. DUX4 binding to retroelements creates promoters that are active in FSHD muscle and testis. PLoS Genet 9, e1003947 (2013).
Supplementary Figure 9 Luciferase assay comparing activation of the human ZSCAN4 reporter by Dux, DUX4, and several chimeras.
(a) We tested the activity of DUX4, Dux, and the indicated chimeras on the published human ZSCAN4 promoter driving luciferase, which has four good matches to the DUX4 binding motif and two good matches to the Dux binding motif and is activated by both DUX4 and Dux. Data shown are mean fold change over empty vector of 3 cell cultures prepared in parallel for each condition. Error bars are s.e.m. This experiment was completed on one occasion.
Supplementary Figure 10 Dux-binding sites identified through two complementary ChIP–seq approaches.
(a) Cartoons of antibodies and chimera combinations used in ChIP-seq. (b) Quantity of overlapping peaks by genomic coordinates for each antibody listed. (c) Top motif is a de novo motif prediction for peaks called from MMH-expressing cells immunoprecipitated with 50:50 mix of MO488 and MO489 antibodies compared to a mock pull-down. Bottom motif is a de novo motif prediction for peaks called from Dux-expressing cells immunoprecipitated with A-19 antibody compared to a mock pull-down. (d) Comparison of MMH transcriptome and Dux transcriptome in mouse muscle cells based on RNA-seq following transgene induction by doxycycline-treatment for 18 hours for MMH_clone6 and 36 hours for Dux_clone15B. These time points were chosen such that they immediately precede the predominant wave of cell death that occurs after prolonged exposure of muscle cells to Dux, so that they are matched functionally if not temporally. Comparator transcriptome for determining differential expression of genes by Dux and MMH was that of firefly luciferase-expressing mouse muscle cells. Data shown are from three cell cultures of each condition. Dux-expressing and luciferase-expressing cells were prepared and sequenced in parallel. MMH-expressing cells were prepared and sequenced at a separate time. Pearson correlation coefficient was 0.7847.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10. (PDF 1199 kb)
Supplementary Table 1
Genes up- or down-regulated in Dux-expressing mouse muscle cells more than absolute(log2-Fold Change >= 2) (XLSX 294 kb)
Supplementary Table 2
ChIP-seq peaks for Dux- and MMH-expressing mouse muscle cells (XLSX 10377 kb)
Supplementary Table 3
List of genes in 2C-like signature (n=469 total,) and ranked by their log2FoldChange in Dux transcriptome (XLSX 62 kb)
Supplementary Table 4
GO analysis of Dux transcriptome (XLSX 13 kb)
Supplementary Table 5
Genes up- or down-regulated in DUX4-expressing mouse muscle cells more than absolute(log2-Fold Change >= 2) (XLSX 297 kb)
Supplementary Table 6
ChIP-seq peaks for DUX4-expressing mouse muscle cells (XLSX 8497 kb)
Supplementary Table 7
List of genes in 2C-like signature and ranked by their log2FoldChange in Dux transcriptome (XLSX 54 kb)
Supplementary Table 8
Repetitive element differential expression analysis in Dux-expressing mouse muscle cells (XLSX 160 kb)
Supplementary Table 9
Repetitive element differential expression analysis in DUX4-expressing mouse muscle cells (XLSX 152 kb)
Supplementary Table 10
Peak associated genes in Dux-expressing mouse muscle cells (XLSX 133 kb)
Supplementary Table 11
Peak associated genes in DUX4-expressing mouse muscle cells (XLSX 93 kb)
Supplementary Table 12
RT-qPCR primers (XLSX 9 kb)
Rights and permissions
About this article
Cite this article
Whiddon, J., Langford, A., Wong, CJ. et al. Conservation and innovation in the DUX4-family gene network. Nat Genet 49, 935–940 (2017). https://doi.org/10.1038/ng.3846
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3846
This article is cited by
-
Jump-starting life: balancing transposable element co-option and genome integrity in the developing mammalian embryo
EMBO Reports (2024)
-
The homeobox transcription factor DUXBL controls exit from totipotency
Nature Genetics (2024)
-
Unreprogrammed H3K9me3 prevents minor zygotic genome activation and lineage commitment in SCNT embryos
Nature Communications (2023)
-
A genome-wide screen reveals new regulators of the 2-cell-like cell state
Nature Structural & Molecular Biology (2023)
-
Maternal TDP-43 interacts with RNA Pol II and regulates zygotic genome activation
Nature Communications (2023)