Abstract
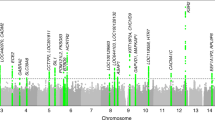
Chronic sleep disturbances, associated with cardiometabolic diseases, psychiatric disorders and all-cause mortality1,2, affect 25–30% of adults worldwide3. Although environmental factors contribute substantially to self-reported habitual sleep duration and disruption, these traits are heritable4,5,6,7,8,9 and identification of the genes involved should improve understanding of sleep, mechanisms linking sleep to disease and development of new therapies. We report single- and multiple-trait genome-wide association analyses of self-reported sleep duration, insomnia symptoms and excessive daytime sleepiness in the UK Biobank (n = 112,586). We discover loci associated with insomnia symptoms (near MEIS1, TMEM132E, CYCL1 and TGFBI in females and WDR27 in males), excessive daytime sleepiness (near AR–OPHN1) and a composite sleep trait (near PATJ (INADL) and HCRTR2) and replicate a locus associated with sleep duration (at PAX8). We also observe genetic correlation between longer sleep duration and schizophrenia risk (rg = 0.29, P = 1.90 × 10−13) and between increased levels of excessive daytime sleepiness and increased measures for adiposity traits (body mass index (BMI): rg = 0.20, P = 3.12 × 10−9; waist circumference: rg = 0.20, P = 2.12 × 10−7).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Fernandez-Mendoza, J. & Vgontzas, A.N. Insomnia and its impact on physical and mental health. Curr. Psychiatry Rep. 15, 418 (2013).
Luyster, F.S., Strollo, P.J. Jr., Zee, P.C. & Walsh, J.K. Sleep: a health imperative. Sleep 35, 727–734 (2012).
Stranges, S., Tigbe, W., Gómez-Olivé, F.X., Thorogood, M. & Kandala, N.B. Sleep problems: an emerging global epidemic? Findings from the INDEPTH WHO-SAGE study among more than 40,000 older adults from 8 countries across Africa and Asia. Sleep 35, 1173–1181 (2012).
de Castro, J.M. The influence of heredity on self-reported sleep patterns in free-living humans. Physiol. Behav. 76, 479–486 (2002).
Evans, D.S. et al. Habitual sleep/wake patterns in the Old Order Amish: heritability and association with non-genetic factors. Sleep 34, 661–669 (2011).
Heath, A.C., Eaves, L.J., Kirk, K.M. & Martin, N.G. Effects of lifestyle, personality, symptoms of anxiety and depression, and genetic predisposition on subjective sleep disturbance and sleep pattern. Twin Res. 1, 176–188 (1998).
Heath, A.C., Kendler, K.S., Eaves, L.J. & Martin, N.G. Evidence for genetic influences on sleep disturbance and sleep pattern in twins. Sleep 13, 318–335 (1990).
Partinen, M., Kaprio, J., Koskenvuo, M., Putkonen, P. & Langinvainio, H. Genetic and environmental determination of human sleep. Sleep 6, 179–185 (1983).
Wing, Y.K. et al. Familial aggregation and heritability of insomnia in a community-based study. Sleep Med. 13, 985–990 (2012).
He, Y. et al. The transcriptional repressor DEC2 regulates sleep length in mammals. Science 325, 866–870 (2009).
Gottlieb, D.J., O'Connor, G.T. & Wilk, J.B. Genome-wide association of sleep and circadian phenotypes. BMC Med. Genet. 8 (Suppl. 1), S9 (2007).
Gottlieb, D.J. et al. Novel loci associated with usual sleep duration: the CHARGE Consortium Genome-Wide Association Study. Mol. Psychiatry 20, 1232–1239 (2015).
Byrne, E.M. et al. A genome-wide association study of sleep habits and insomnia. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 162B, 439–451 (2013).
Allebrandt, K.V. et al. A KATP channel gene effect on sleep duration: from genome-wide association studies to function in Drosophila. Mol. Psychiatry 18, 122–132 (2013).
Gehrman, P.R., Keenan, B.T., Byrne, E.M. & Pack, A.I. Genetics of sleep disorders. Psychiatr. Clin. North Am. 38, 667–681 (2015).
Sehgal, A. & Mignot, E. Genetics of sleep and sleep disorders. Cell 146, 194–207 (2011).
Lane, J.M. et al. Genome-wide association analysis identifies novel loci for chronotype in 100,420 individuals from the UK Biobank. Nat. Commun. 7, 10889 (2016).
Jones, S.E. et al. Genome-wide association analyses in 128,266 individuals identifies new morningness and sleep duration loci. PLoS Genet. 12, e1006125 (2016).
Hu, Y. et al. GWAS of 89,283 individuals identifies genetic variants associated with self-reporting of being a morning person. Nat. Commun. 7, 10448 (2016).
Pemberton, R. & Fuller Tyszkiewicz, M.D. Factors contributing to depressive mood states in everyday life: a systematic review. J. Affect. Disord. 200, 103–110 (2016).
Foral, P., Knezevich, J., Dewan, N. & Malesker, M. Medication-induced sleep disturbances. Consult Pharm. 26, 414–425 (2011).
Rosenberg, R.P. Clinical assessment of excessive daytime sleepiness in the diagnosis of sleep disorders. J. Clin. Psychiatry 76, e1602 (2015).
Gonnissen, H.K. et al. Sleep duration, sleep quality and body weight: parallel developments. Physiol. Behav. 121, 112–116 (2013).
Kurant, E. et al. Dorsotonals/homothorax, the Drosophila homologue of meis1, interacts with extradenticle in patterning of the embryonic PNS. Development 125, 1037–1048 (1998).
Casares, F. & Mann, R.S. Control of antennal versus leg development in Drosophila. Nature 392, 723–726 (1998).
Hisa, T. et al. Hematopoietic, angiogenic and eye defects in Meis1 mutant animals. EMBO J. 23, 450–459 (2004).
Davidson, S., Miller, K.A., Dowell, A., Gildea, A. & Mackenzie, A. A remote and highly conserved enhancer supports amygdala specific expression of the gene encoding the anxiogenic neuropeptide substance-P. Mol. Psychiatry 11 323, 410–421 (2006).
Oh-hashi, K., Naruse, Y., Amaya, F., Shimosato, G. & Tanaka, M. Cloning and characterization of a novel GRP78-binding protein in the rat brain. J. Biol. Chem. 278, 10531–10537 (2003).
Erhardt, A. et al. Replication and meta-analysis of TMEM132D gene variants in panic disorder. Transl. Psychiatry 2, e156 (2012).
Sklar, P. et al. Whole-genome association study of bipolar disorder. Mol. Psychiatry 13, 558–569 (2008).
Edwards, A.C. et al. Genome-wide association study of comorbid depressive syndrome and alcohol dependence. Psychiatr. Genet. 22, 31–41 (2012).
Han, K.E. et al. Pathogenesis and treatments of TGFBI corneal dystrophies. Prog. Retin. Eye Res. 50, 67–88 (2016).
Bradfield, J.P. et al. A genome-wide meta-analysis of six type 1 diabetes cohorts identifies multiple associated loci. PLoS Genet. 7, e1002293 (2011).
Patry, M. et al. βig-h3 represses T-cell activation in type 1 diabetes. Diabetes 64, 4212–4219 (2015).
Han, B. et al. TGFBI (βIG-H3) is a diabetes-risk gene based on mouse and human genetic studies. Hum. Mol. Genet. 23, 4597–4611 (2014).
Poelmans, G., Buitelaar, J.K., Pauls, D.L. & Franke, B. A theoretical molecular network for dyslexia: integrating available genetic findings. Mol. Psychiatry 16, 365–382 (2011).
Dalal, J. et al. Translational profiling of hypocretin neurons identifies candidate molecules for sleep regulation. Genes Dev. 27, 565–578 (2013).
Yelin-Bekerman, L. et al. Hypocretin neuron–specific transcriptome profiling identifies the sleep modulator Kcnh4a. eLife 4, e08638 (2015).
Mackiewicz, M. et al. Macromolecule biosynthesis: a key function of sleep. Physiol. Genomics 31, 441–457 (2007).
Takahama, K. et al. Pan-neuronal knockdown of the c-Jun N-terminal kinase (JNK) results in a reduction in sleep and longevity in Drosophila. Biochem. Biophys. Res. Commun. 417, 807–811 (2012).
Farh, K.K. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015).
Ward, L.D. & Kellis, M. HaploReg v4: systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 44 D1, D877–D881 (2016).
Hamdan, F.F. et al. De novo mutations in FOXP1 in cases with intellectual disability, autism, and language impairment. Am. J. Hum. Genet. 87, 671–678 (2010).
Fan, Y., Newman, T., Linardopoulou, E. & Trask, B.J. Gene content and function of the ancestral chromosome fusion site in human chromosome 2q13–2q14.1 and paralogous regions. Genome Res. 12, 1663–1672 (2002).
Fan, Y., Linardopoulou, E., Friedman, C., Williams, E. & Trask, B.J. Genomic structure and evolution of the ancestral chromosome fusion site in 2q13–2q14.1 and paralogous regions on other human chromosomes. Genome Res. 12, 1651–1662 (2002).
Wang, J., Duncan, D., Shi, Z. & Zhang, B. WEB-based GEne SeT AnaLysis Toolkit (WebGestalt): update 2013. Nucleic Acids Res. 41, W77–W83 (2013).
Cade, B.E. et al. Common variants in DRD2 are associated with sleep duration: the CARe consortium. Hum. Mol. Genet. 25, 167–179 (2016).
Loh, P.R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Bulik-Sullivan, B.K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Finucane, H.K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Zhu, X. et al. Meta-analysis of correlated traits via summary statistics from GWASs with an application in hypertension. Am. J. Hum. Genet. 96, 21–36 (2015).
Mignot, E. Sleep, sleep disorders and hypocretin (orexin). Sleep Med. 5 (Suppl. 1), S2–S8 (2004).
Thompson, M.D., Xhaard, H., Sakurai, T., Rainero, I. & Kukkonen, J.P. OX1 and OX2 orexin/hypocretin receptor pharmacogenetics. Front. Neurosci. 8, 57 (2014).
Herring, W.J. et al. Suvorexant in patients with insomnia: results from two 3-month randomized controlled clinical trials. Biol. Psychiatry 79, 136–148 (2016).
Shieh, B.H. & Niemeyer, B. A novel protein encoded by the InaD gene regulates recovery of visual transduction in Drosophila. Neuron 14, 201–210 (1995).
Peirson, S.N. et al. Microarray analysis and functional genomics identify novel components of melanopsin signaling. Curr. Biol. 17, 1363–1372 (2007).
Bécamel, C. et al. The serotonin 5-HT2A and 5-HT2C receptors interact with specific sets of PDZ proteins. J. Biol. Chem. 279, 20257–20266 (2004).
Sharpley, A.L., Elliott, J.M., Attenburrow, M.J. & Cowen, P.J. Slow wave sleep in humans: role of 5-HT2A and 5-HT2C receptors. Neuropharmacology 33, 467–471 (1994).
Rosenberg, R. et al. APD125, a selective serotonin 5-HT2A receptor inverse agonist, significantly improves sleep maintenance in primary insomnia. Sleep 31, 1663–1671 (2008).
Winkelmann, J. et al. Genome-wide association study of restless legs syndrome identifies common variants in three genomic regions. Nat. Genet. 39, 1000–1006 (2007).
Xiong, L. et al. MEIS1 intronic risk haplotype associated with restless legs syndrome affects its mRNA and protein expression levels. Hum. Mol. Genet. 18, 1065–1074 (2009).
Schulte, E.C. et al. Targeted resequencing and systematic in vivo functional testing identifies rare variants in MEIS1 as significant contributors to restless legs syndrome. Am. J. Hum. Genet. 95, 85–95 (2014).
Spieler, D. et al. Restless legs syndrome–associated intronic common variant in Meis1 alters enhancer function in the developing telencephalon. Genome Res. 24, 592–603 (2014).
Moore, H. IV et al. Periodic leg movements during sleep are associated with polymorphisms in BTBD9, TOX3/BC034767, MEIS1, MAP2K5/SKOR1, and PTPRD. Sleep 37, 1535–1542 (2014).
Winkelmann, J. et al. Genome-wide association study identifies novel restless legs syndrome susceptibility loci on 2p14 and 16q12.1. PLoS Genet. 7, e1002171 (2011).
Allen, R.P., Barker, P.B., Horská, A. & Earley, C.J. Thalamic glutamate/glutamine in restless legs syndrome: increased and related to disturbed sleep. Neurology 80, 2028–2034 (2013).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Byrne, E.M., Gehrman, P.R., Trzaskowski, M., Tiemeier, H. & Pack, A.I. Genetic correlation analysis suggests association between increased self-reported sleep duration in adults and schizophrenia and type 2 diabetes. Sleep 39, 1853–1857 (2016).
Wulff, K., Dijk, D.J., Middleton, B., Foster, R.G. & Joyce, E.M. Sleep and circadian rhythm disruption in schizophrenia. Br. J. Psychiatry 200, 308–316 (2012).
Poulin, J. et al. Sleep habits in middle-aged, non-hospitalized men and women with schizophrenia: a comparison with healthy controls. Psychiatry Res. 179, 274–278 (2010).
Chouinard, S., Poulin, J., Stip, E. & Godbout, R. Sleep in untreated patients with schizophrenia: a meta-analysis. Schizophr. Bull. 30, 957–967 (2004).
Hattersley, A.T. & Tooke, J.E. The fetal insulin hypothesis: an alternative explanation of the association of low birthweight with diabetes and vascular disease. Lancet 353, 1789–1792 (1999).
Horikoshi, M. et al. New loci associated with birth weight identify genetic links between intrauterine growth and adult height and metabolism. Nat. Genet. 45, 76–82 (2013).
Ananthakrishnan, A.N. et al. Sleep duration affects risk for ulcerative colitis: a prospective cohort study. Clin. Gastroenterol. Hepatol. 12, 1879–1886 (2014).
Tasali, E., Leproult, R., Ehrmann, D.A. & Van Cauter, E. Slow-wave sleep and the risk of type 2 diabetes in humans. Proc. Natl. Acad. Sci. USA 105, 1044–1049 (2008).
Nedeltcheva, A.V. & Scheer, F.A. Metabolic effects of sleep disruption, links to obesity and diabetes. Curr. Opin. Endocrinol. Diabetes Obes. 21, 293–298 (2014).
Vgontzas, A.N. et al. Obesity without sleep apnea is associated with daytime sleepiness. Arch. Intern. Med. 158, 1333–1337 (1998).
Bixler, E.O. et al. Excessive daytime sleepiness in a general population sample: the role of sleep apnea, age, obesity, diabetes, and depression. J. Clin. Endocrinol. Metab. 90, 4510–4515 (2005).
Swanson, J.M. The UK Biobank and selection bias. Lancet 380, 110 (2012).
Collins, R. What makes UK Biobank special? Lancet 379, 1173–1174 (2012).
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Allen, N.E., Sudlow, C., Peakman, T. & Collins, R. UK Biobank. UK Biobank data: come and get it. Sci. Transl. Med. 6, 224ed4 (2014).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
O'Connell, J. et al. Haplotype estimation for biobank-scale data sets. Nat. Genet. 48, 817–820 (2016).
Loh, P.R., Palamara, P.F. & Price, A.L. Fast and accurate long-range phasing in a UK Biobank cohort. Nat. Genet. 48, 811–816 (2016).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Chang, C.C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
International HapMap Consortium. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Locke, A.E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat. Genet. 42, 105–116 (2010).
Acknowledgements
This research has been conducted using the UK Biobank Resource under application number 6818. We would like to thank the participants and researchers from the UK Biobank who contributed or collected data. This work was supported by US NIH grants R01DK107859 (R.S.), R21HL121728 (R.S.), F32DK102323 (J.M.L.), R01HL113338 (J.M.L., S.R. and R.S.), R01DK102696 (R.S. and F.A.J.L.S.), R01DK105072 (R.S. and F.A.J.L.S.), T32HL007567 (J.L.) and HG003054 (X.Z.), the University of Manchester (Research Infrastructure Fund), the Wellcome Trust (salary support for D.W.R. and A.L.) and UK Medical Research Council MC_UU_12013/5 (D.A.L.). Data on glycemic traits have been contributed by MAGIC investigators and were downloaded from http://www.magicinvestigators.org/. Data on coronary artery disease and myocardial infarction have been contributed by CARDIoGRAMplusC4D investigators and were downloaded from http://www.cardiogramplusc4d.org/. We thank the International Genomics of Alzheimer's Project (IGAP) for providing summary results data for these analyses.
Author information
Authors and Affiliations
Contributions
J.M.L., M.K.R. and R.S. designed the study. J.M.L., J.L., I.V. and R.S. performed genetic analyses. J.M.L. and R.S. wrote the manuscript, and all co-authors helped interpret data and reviewed and edited the manuscript, before approving its submission. R.S. is the guarantor of this work and, as such, had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–12. (PDF 3987 kb)
Rights and permissions
About this article
Cite this article
Lane, J., Liang, J., Vlasac, I. et al. Genome-wide association analyses of sleep disturbance traits identify new loci and highlight shared genetics with neuropsychiatric and metabolic traits. Nat Genet 49, 274–281 (2017). https://doi.org/10.1038/ng.3749
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3749
This article is cited by
-
Sex effects on DNA methylation affect discovery in epigenome-wide association study of schizophrenia
Molecular Psychiatry (2024)
-
Exome sequencing identifies genes associated with sleep-related traits
Nature Human Behaviour (2024)
-
Dance/movement therapy for improving metabolic parameters in long-term veterans with schizophrenia
Schizophrenia (2024)
-
Investigating the causal interplay between sleep traits and risk of acute myocardial infarction: a Mendelian randomization study
BMC Medicine (2023)
-
Nonrestorative sleep is a risk factor for metabolic syndrome in the general Japanese population
Diabetology & Metabolic Syndrome (2023)