Abstract
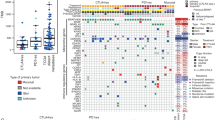
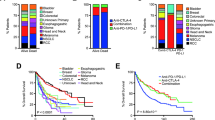
Immune checkpoint blockade has shown significant promise as an anticancer treatment, yet the determinants of response are not completely understood. Here we show that somatic mutations in SERPINB3 and SERPINB4 are associated with survival after anti-CTLA4 immunotherapy in two independent cohorts of patients with melanoma (n = 174). Interestingly, serpins are homologs of the well-known ovalbumin antigen and are associated with autoimmunity. Our findings have implications for the personalization of immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Hodi, F.S. et al. N. Engl. J. Med. 363, 711–723 (2010).
Robert, C. et al. N. Engl. J. Med. 364, 2517–2526 (2011).
Brahmer, J. et al. N. Engl. J. Med. 373, 123–135 (2015).
Motzer, R.J. et al. N. Engl. J. Med. 373, 1803–1813 (2015).
Robert, C. et al. N. Engl. J. Med. 372, 320–330 (2015).
Snyder, A. et al. N. Engl. J. Med. 371, 2189–2199 (2014).
Rizvi, N.A. et al. Science 348, 124–128 (2015).
Le, D.T. et al. N. Engl. J. Med. 372, 2509–2520 (2015).
Van Allen, E.M. et al. Science 350, 207–211 (2015).
McGranahan, N. et al. Science 351, 1463–1469 (2016).
Cancer Genome Atlas Network. Cell 161, 1681–1696 (2015).
Vidalino, L. et al. Autoimmun. Rev. 9, 108–112 (2009).
Gatto, M. et al. Clin. Rev. Allergy Immunol. 45, 267–280 (2013).
Sivaprasad, U. et al. J. Invest. Dermatol. 135, 160–169 (2015).
Schneider, S.S. et al. Proc. Natl. Acad. Sci. USA 92, 3147–3151 (1995).
Catanzaro, J.M. et al. Nat. Commun. 5, 3729 (2014).
Guan, J., Gupta, R. & Filipp, F.V. Sci. Rep. 5, 7857 (2015).
Katagiri, C., Nakanishi, J., Kadoya, K. & Hibino, T. J. Cell Biol. 172, 983–990 (2006).
El-Rachkidy, R.G., Young, H.S., Griffiths, C.E. & Camp, R.D. J. Invest. Dermatol. 128, 2219–2224 (2008).
Lysvand, H., Hagen, L., Klubicka, L., Slupphaug, G. & Iversen, O.J. Biochim. Biophys. Acta 1842, 734–738 (2014).
Münz, C. Front. Immunol. 3, 9 (2012).
Schumacher, T. et al. Nature 512, 324–327 (2014).
Mine, Y. & Yang, M. J. Agric. Food Chem. 56, 4874–4900 (2008).
Holen, E. & Elsayed, S. Clin. Exp. Allergy 26, 1080–1088 (1996).
DePristo, M.A. et al. Nat. Genet. 43, 491–498 (2011).
Li, H. & Durbin, R. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. Genome Res. 20, 1297–1303 (2010).
Cibulskis, K. et al. Nat. Biotechnol. 31, 213–219 (2013).
Koboldt, D.C. et al. Genome Res. 22, 568–576 (2012).
Larson, D.E. et al. Bioinformatics 28, 311–317 (2012).
Saunders, C.T. et al. Bioinformatics 28, 1811–1817 (2012).
Strona, G., Nappo, D., Boccacci, F., Fattorini, S. & San-Miguel-Ayanz, J. Nat. Commun. 5, 4114 (2014).
Kim, J. et al. Nat. Genet. 48, 600–606 (2016).
Schellens, I.M. et al. PLoS One 10, e0136417 (2015).
Snyder, A. & Chan, T.A. Curr. Opin. Genet. Dev. 30, 7–16 (2015).
Nielsen, M. et al. Protein Sci. 12, 1007–1017 (2003).
Andreatta, M. et al. Immunogenetics 67, 641–650 (2015).
Acknowledgements
We thank J. Wolchok, T. Merghoub, J. Yuan, P. Wong, and A. Snyder for collaborative interactions. We thank the Integrated Genomics Operation and the Ludwig Immune Monitoring Facility at Memorial Sloan Kettering for technical assistance. We thank A. Heguy and the Genome Technology Center at New York University and L. Mangarin for assistance with validation sequencing. This work was funded by a Pershing Square Sohn Cancer Research grant (T.A.C.), the Frederick Adler Chair (T.A.C.), Stand Up 2 Cancer (T.A.C.), the STARR Cancer Consortium (T.A.C.), and in part through NIH/NCI Cancer Center Support Grant P30 CA008748. Research supported by a Stand Up To Cancer – Cancer Research Institute Cancer Immunology Translational Cancer Research Grant. Stand Up To Cancer is a program of the Entertainment Industry Foundation administered by the American Association for Cancer Research.
Author information
Authors and Affiliations
Contributions
T.A.C. and N.R. designed and conceived the study. Analysis of mutations in individual genes with outcome was performed by S.M.K., N.W., N.R., V.M., and J.J.H. Neoantigen analysis was performed by J.J.H., S.M.K., and V.M. Analysis of expression data was performed by L.A.W., A.D., and N.R. N.R., J.J.H., and T.A.C. prepared the manuscript. All authors participated in discussion of the final manuscript and interpretation of results.
Corresponding author
Ethics declarations
Competing interests
T.A.C. is a co-founder of Gritstone Oncology.
Integrated supplementary information
Supplementary Figure 1 Example IGV plots of SERPINB3 mutations in cohort 1 (tumor–normal pairs).
Left, sequencing data from the tumor; right, sequencing data from the corresponding normal sample. (a) Patient CR1509 (Tcov = 119, TAF = 0.30). (b) Patient CR3665 (Tcov = 246; TAF = 0.20). (c) Patient NR2137 (Tcov = 191, TAF = 0.24).
Supplementary Figure 2 Alignment of human SERPINB3 and chicken ovalbumin proteins.
Residues in red represent SERPINB3 mutations identified in the presently studied patient cohorts. SERPINB3 regions highlighted in gray represent experimentally validated human HLA-binding peptides35. Ovalbumin regions highlighted in yellow represent functionally validated immunogenic epitopes of human T cells24. (An asterisk indicates an exact amino acid match, a colon indicates alignment of amino acid residues with strongly similar properties, and a period indicates alignment of amino acid residues with weakly similar properties, as described at http://www.ebi.ac.uk/Tools/msa/clustalo/help/faq.html#23 (accessed 28 March 2016).
Supplementary Figure 3 Expression of SERPINB3 and SERPINB4 in primary and metastatic melanomas.
(a) Expression of SERPINB3 in primary melanoma versus regional or distant metastatic samples (P = 1.16 × 10–13, Wilcoxon rank-sum test). (b) Expression of SERPINB4 in primary melanoma versus regional or distant metastatic samples (P = 2.99 × 10–15, Wilcoxon rank-sum test).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3. (PDF 582 kb)
Supplementary Table 1
Recurrently mutated genes in melanoma as identified by InVex analysis performed by TCGA. (XLSX 8 kb)
Supplementary Table 2
Multivariate model of overall survival and SERPINB3 and SERPINB4 mutations. (XLSX 10 kb)
Supplementary Table 3
SERPINB3 and SERPINB4 mutations in both cohorts of patients. (XLSX 12 kb)
Supplementary Table 4
MHC class I predicted neoantigens from SERPINB3 and SERPINB4 mutations. (XLSX 11 kb)
Supplementary Table 5
MHC class II predicted neoantigens from SERPINB3 and SERPINB4 mutations. (XLSX 16 kb)
Rights and permissions
About this article
Cite this article
Riaz, N., Havel, J., Kendall, S. et al. Recurrent SERPINB3 and SERPINB4 mutations in patients who respond to anti-CTLA4 immunotherapy. Nat Genet 48, 1327–1329 (2016). https://doi.org/10.1038/ng.3677
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3677
This article is cited by
-
Exploring immunotherapy in colorectal cancer
Journal of Hematology & Oncology (2022)
-
Non‑small cell lung cancer carrying PBRM1 mutation suggests an immunologically cold phenotype leading to immunotherapy failure even with high TMB
Scientific Reports (2022)
-
NLRC5/CITA expression correlates with efficient response to checkpoint blockade immunotherapy
Scientific Reports (2021)
-
Identification of immune-related genes with prognostic significance in the microenvironment of cutaneous melanoma
Virchows Archiv (2021)
-
ZFHX3 mutation as a protective biomarker for immune checkpoint blockade in non-small cell lung cancer
Cancer Immunology, Immunotherapy (2021)