Abstract
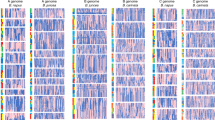
Brassica species, including crops such as cabbage, turnip and oilseed, display enormous phenotypic variation. Brassica genomes have all undergone a whole-genome triplication (WGT) event with unknown effects on phenotype diversification. We resequenced 199 Brassica rapa and 119 Brassica oleracea accessions representing various morphotypes and identified signals of selection at the mesohexaploid subgenome level. For cabbage morphotypes with their typical leaf-heading trait, we identified four subgenome loci that show signs of parallel selection among subgenomes within B. rapa, as well as four such loci within B. oleracea. Fifteen subgenome loci are under selection and are shared by these two species. We also detected strong subgenome parallel selection linked to the domestication of the tuberous morphotypes, turnip (B. rapa) and kohlrabi (B. oleracea). Overall, we demonstrated that the mesohexaploidization of the two Brassica genomes contributed to their diversification into heading and tuber-forming morphotypes through convergent subgenome parallel selection of paralogous genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Nagaharu, U. Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn. J. Bot. 7, 389–452 (1935).
Zhao, J. et al. Genetic relationships within Brassica rapa as inferred from AFLP fingerprints. Theor. Appl. Genet. 110, 1301–1314 (2005).
Cheng, F., Wu, J. & Wang, X. Genome triplication drove the diversification of Brassica plants. Hortic. Res. 1, 14024 (2014).
Lenser, T. & Theißen, G. Molecular mechanisms involved in convergent crop domestication. Trends Plant Sci. 18, 704–714 (2013).
Vavilov, N.I. The law of homologous series in variation. J. Genet. 12, 47–89 (1922).
Hovav, R., Chaudhary, B., Udall, J.A., Flagel, L. & Wendel, J.F. Parallel domestication, convergent evolution and duplicated gene recruitment in allopolyploid cotton. Genetics 179, 1725–1733 (2008).
Fuller, D.Q. et al. Convergent evolution and parallelism in plant domestication revealed by an expanding archaeological record. Proc. Natl. Acad. Sci. USA 111, 6147–6152 (2014).
Wang, M. et al. The genome sequence of African rice (Oryza glaberrima) and evidence for independent domestication. Nat. Genet. 46, 982–988 (2014).
Cheng, F. et al. Deciphering the diploid ancestral genome of the mesohexaploid Brassica rapa. Plant Cell 25, 1541–1554 (2013).
Wang, X. et al. The genome of the mesopolyploid crop species Brassica rapa. Nat. Genet. 43, 1035–1039 (2011).
Liu, S. et al. The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat. Commun. 5, 3930 (2014).
Cheng, F. et al. Biased gene fractionation and dominant gene expression among the subgenomes of Brassica rapa. PLoS One 7, e36442 (2012).
Tang, H. et al. Altered patterns of fractionation and exon deletions in Brassica rapa support a two-step model of paleohexaploidy. Genetics 190, 1563–1574 (2012).
Pritchard, J.K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Prakash, S.H.K. Taxonomy, cytogenetics and origin of crop brassicas, a review. Opera Bot. 55, 1–57 (1980).
Arias, T., Beilstein, M.A., Tang, M., McKain, M.R. & Pires, J.C. Diversification times among Brassica (Brassicaceae) crops suggest hybrid formation after 20 million years of divergence. Am. J. Bot. 101, 86–91 (2014).
Bonnema, G., Carpio, D.P.D. & Zhao, J. in Genetics, Genomics and Breeding of Vegetable Brassicas 1st edn. (eds. Sadowski, J. & Kole, C.) 81–124 (Science Publishers, 2011).
Boswell, V.R. in Our Vegetable Travelers 1st edn, Vol. 96 (ed. Boswell, V.R.) 145–217 (National Geographic Magazine, 1949).
Tajima, F. Evolutionary relationship of DNA sequences in finite populations. Genetics 105, 437–460 (1983).
Xu, X. et al. Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat. Biotechnol. 30, 105–111 (2012).
Voight, B.F., Kudaravalli, S., Wen, X. & Pritchard, J.K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Axelsson, E. et al. The genomic signature of dog domestication reveals adaptation to a starch-rich diet. Nature 495, 360–364 (2013).
Stamm, P. & Kumar, P.P. The phytohormone signal network regulating elongation growth during shade avoidance. J. Exp. Bot. 61, 2889–2903 (2010).
Santner, A. & Estelle, M. Recent advances and emerging trends in plant hormone signalling. Nature 459, 1071–1078 (2009).
Gazzarrini, S. & McCourt, P. Cross-talk in plant hormone signalling: what Arabidopsis mutants are telling us. Ann. Bot. 91, 605–612 (2003).
Manju, R.V., Abida, P.S., Sudarshana, L., Nataraja, K.N. & Sashidhar, V.R. Unusual discrimination against carrier protein antibodies during partial purification of hapten-protein polyclonal antibodies to plant stress hormones. Indian J. Exp. Biol. 33, 1–5 (1995).
Wang, J., Guo, H., Jin, D., Wang, X. & Paterson, A.H. in The Brassica rapa Genome (ed. Wang, X.) 121–130 (Springer, 2015).
Hake, S. et al. The role of knox genes in plant development. Annu. Rev. Cell Dev. Biol. 20, 125–151 (2004).
Kidner, C.A. & Timmermans, M.C. Mixing and matching pathways in leaf polarity. Curr. Opin. Plant Biol. 10, 13–20 (2007).
Husbands, A.Y., Chitwood, D.H., Plavskin, Y. & Timmermans, M.C. Signals and prepatterns: new insights into organ polarity in plants. Genes Dev. 23, 1986–1997 (2009).
Byrne, M.E. Networks in leaf development. Curr. Opin. Plant Biol. 8, 59–66 (2005).
Bowman, J.L., Eshed, Y. & Baum, S.F. Establishment of polarity in angiosperm lateral organs. Trends Genet. 18, 134–141 (2002).
Eshed, Y., Baum, S.F., Perea, J.V. & Bowman, J.L. Establishment of polarity in lateral organs of plants. Curr. Biol. 11, 1251–1260 (2001).
Izhaki, A. & Bowman, J.L. KANADI and class III HD-Zip gene families regulate embryo patterning and modulate auxin flow during embryogenesis in Arabidopsis. Plant Cell 19, 495–508 (2007).
Wu, G. et al. KANADI1 regulates adaxial–abaxial polarity in Arabidopsis by directly repressing the transcription of ASYMMETRIC LEAVES2. Proc. Natl. Acad. Sci. USA 105, 16392–16397 (2008).
Liu, Z., Jia, L., Wang, H. & He, Y. HYL1 regulates the balance between adaxial and abaxial identity for leaf flattening via miRNA-mediated pathways. J. Exp. Bot. 62, 4367–4381 (2011).
Wu, F. et al. The N-terminal double-stranded RNA binding domains of Arabidopsis HYPONASTIC LEAVES1 are sufficient for pre-microRNA processing. Plant Cell 19, 914–925 (2007).
Yu, X. et al. Cloning and structural and expressional characterization of BcpLH gene preferentially expressed in folding leaf of Chinese cabbage. Sci. China C Life Sci. 43, 321–329 (2000).
Townsley, B.T. & Sinha, N.R. A new development: evolving concepts in leaf ontogeny. Annu. Rev. Plant Biol. 63, 535–562 (2012).
Barkoulas, M., Galinha, C., Grigg, S.P. & Tsiantis, M. From genes to shape: regulatory interactions in leaf development. Curr. Opin. Plant Biol. 10, 660–666 (2007).
Piazza, P., Jasinski, S. & Tsiantis, M. Evolution of leaf developmental mechanisms. New Phytol. 167, 693–710 (2005).
Braybrook, S.A. & Kuhlemeier, C. How a plant builds leaves. Plant Cell 22, 1006–1018 (2010).
Kelley, D.R., Arreola, A., Gallagher, T.L. & Gasser, C.S. ETTIN (ARF3) physically interacts with KANADI proteins to form a functional complex essential for integument development and polarity determination in Arabidopsis. Development 139, 1105–1109 (2012).
Pekker, I., Alvarez, J.P. & Eshed, Y. Auxin response factors mediate Arabidopsis organ asymmetry via modulation of KANADI activity. Plant Cell 17, 2899–2910 (2005).
Beuchat, J. et al. BRX promotes Arabidopsis shoot growth. New Phytol. 188, 23–29 (2010).
Santuari, L. et al. Positional information by differential endocytosis splits auxin response to drive Arabidopsis root meristem growth. Curr. Biol. 21, 1918–1923 (2011).
Li, J. et al. BREVIS RADIX is involved in cytokinin-mediated inhibition of lateral root initiation in Arabidopsis. Planta 229, 593–603 (2009).
Mouchel, C.F., Osmont, K.S. & Hardtke, C.S. BRX mediates feedback between brassinosteroid levels and auxin signalling in root growth. Nature 443, 458–461 (2006).
Scacchi, E. et al. Dynamic, auxin-responsive plasma membrane–to-nucleus movement of Arabidopsis BRX. Development 136, 2059–2067 (2009).
Zhang, N. et al. Morphology, carbohydrate composition and vernalization response in a genetically diverse collection of Asian and European turnips (Brassica rapa subsp. rapa). PLoS One 9, e114241 (2014).
Gepts, P. The contribution of genetic and genomic approaches to plant domestication studies. Curr. Opin. Plant Biol. 18, 51–59 (2014).
Kwak, M., Toro, O., Debouck, D.G. & Gepts, P. Multiple origins of the determinate growth habit in domesticated common bean (Phaseolus vulgaris). Ann. Bot. 110, 1573–1580 (2012).
Lin, Z. et al. Parallel domestication of the Shattering1 genes in cereals. Nat. Genet. 44, 720–724 (2012).
Li, W. & Gill, B.S. Multiple genetic pathways for seed shattering in the grasses. Funct. Integr. Genomics 6, 300–309 (2006).
Tang, H. et al. Seed shattering in a wild sorghum is conferred by a locus unrelated to domestication. Proc. Natl. Acad. Sci. USA 110, 15824–15829 (2013).
Bancroft, I. et al. Dissecting the genome of the polyploid crop oilseed rape by transcriptome sequencing. Nat. Biotechnol. 29, 762–766 (2011).
Cheng, F. et al. BRAD, the genetics and genomics database for Brassica plants. BMC Plant Biol. 11, 136 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Yu, X. et al. QTL mapping of leafy heads by genome resequencing in the RIL population of Brassica rapa. PLoS One 8, e76059 (2013).
Cheng, F., Wu, J., Fang, L. & Wang, X. Syntenic gene analysis between Brassica rapa and other Brassicaceae species. Front. Plant Sci. 3, 198 (2012).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Retief, J.D. Phylogenetic analysis using PHYLIP. Methods Mol. Biol. 132, 243–258 (2000).
Akey, J.M., Zhang, G., Zhang, K., Jin, L. & Shriver, M.D. Interrogating a high-density SNP map for signatures of natural selection. Genome Res. 12, 1805–1814 (2002).
Sabeti, P.C. et al. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002).
Berardini, T.Z. et al. Functional annotation of the Arabidopsis genome using controlled vocabularies. Plant Physiol. 135, 745–755 (2004).
Wang, X. et al. Linkage mapping of a dominant male sterility gene Ms-cd1 in Brassica oleracea. Genome 48, 848–854 (2005).
Morrissy, A.S. et al. Next-generation tag sequencing for cancer gene expression profiling. Genome Res. 19, 1825–1835 (2009).
Xue, J. et al. Transcriptome analysis of the brown planthopper Nilaparvata lugens. PLoS One 5, e14233 (2010).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Zhang, Z. et al. KaKs_Calculator: calculating Ka and Ks through model selection and model averaging. Genomics Proteomics Bioinformatics 4, 259–263 (2006).
Schnable, J.C., Springer, N.M. & Freeling, M. Differentiation of the maize subgenomes by genome dominance and both ancient and ongoing gene loss. Proc. Natl. Acad. Sci. USA 108, 4069–4074 (2011).
Acknowledgements
This study was funded by National Program on Key Basic Research Projects (973 Program: 2012CB113900 to Xiaowu Wang and F.C. and 2013CB127000 to Xiaowu Wang), the National High-Technology R&D Program of China (2012AA100201 to J.W.), a National Program on Key Research Project (2016YFD0100307) and the National Natural Science Foundation of China (NSFC grants 31301771 to F.C., 31272179 to J.W. and 31301784 to J. Liang). Research was supported by the Program for Strategic Scientific Alliances between China and the Netherlands, the Key Laboratory of Biology and Genetic Improvement of Horticultural Crops, the Ministry of Agriculture, China, and the Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences. The authors gratefully acknowledge funding from the European Community under the Seventh Framework Programme for Research, Technological Development and Demonstration Activities, for Integrated Project NUE-CROPS FP7-CP-IP 222645. B. oleracea resequencing data and analyses were funded by a grant from the Technological Top Institute (TTI) Green Genetics, Rijk Zwaan and Bejo Zaden.
Author information
Authors and Affiliations
Contributions
Xiaowu Wang, F.C., G.B. and J.W. conceived and designed the experiments. J.W., G.B., Y.Z., M.Z., Yunxia Liu, Yumei Liu, T.B., X.Z., L.F., Y.H., Shujiang Zhang, Shifan Zhang, F.L., Hui Zhang, J.Z., N.G., Z.L., Jin Liu, Y.M., Haijiao Zhang, J.B. and Y.C. contributed materials. Y.W., H.W., N.Z., J.D., Y. Liao and K.W. contributed to phenotyping. N.Z. performed QTL analysis for the RIL populations. R.S., G.B., T.B., H. Zheng, X.H., F.Z., K.L., B.L., D.L., Xiaobo Wang, Jisheng Liu and C.S. contributed to resequencing. F.C., B.L., C.C. and T.B. analyzed the data and performed statistical analysis. P.L., J. Liang and L.F. performed the experiments. F.C. and Xiaowu Wang wrote the manuscript, with help from J. Liang, M.R.F., J.W., G.B., T.B. and P.L.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–24, Supplementary Tables 1–10, 17–21, 23, 24, 26, 31 and 33–36, and Supplementary Note. (PDF 4680 kb)
Supplementary Tables 11–16, 22, 25, 27–30 and 32
Supplementary Tables 11–16, 22, 25, 27–30 and 32. (XLS 85 kb)
Rights and permissions
About this article
Cite this article
Cheng, F., Sun, R., Hou, X. et al. Subgenome parallel selection is associated with morphotype diversification and convergent crop domestication in Brassica rapa and Brassica oleracea. Nat Genet 48, 1218–1224 (2016). https://doi.org/10.1038/ng.3634
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3634
This article is cited by
-
Asymmetric and parallel subgenome selection co-shape common carp domestication
BMC Biology (2024)
-
Large-scale gene expression alterations introduced by structural variation drive morphotype diversification in Brassica oleracea
Nature Genetics (2024)
-
Structural variations fine-tune gene expression to steer Brassica oleracea diversification
Nature Genetics (2024)
-
Differences in the transcriptional immune response to Albugo candida between white rust resistant and susceptible cultivars in Brassica rapa L.
Scientific Reports (2023)
-
Vegetable biology and breeding in the genomics era
Science China Life Sciences (2023)