Abstract
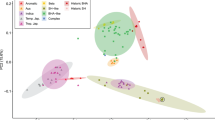
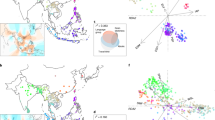
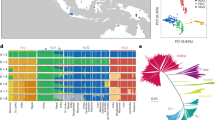
African rice (Oryza glaberrima Steud.) is a cereal crop species closely related to Asian rice (Oryza sativa L.) but was independently domesticated in West Africa ∼3,000 years ago1,2,3. African rice is rarely grown outside sub-Saharan Africa but is of global interest because of its tolerance to abiotic stresses4,5. Here we describe a map of 2.32 million SNPs of African rice from whole-genome resequencing of 93 landraces. Population genomic analysis shows a population bottleneck in this species that began ∼13,000–15,000 years ago with effective population size reaching its minimum value ∼3,500 years ago, suggesting a protracted period of population size reduction likely commencing with predomestication management and/or cultivation. Genome-wide association studies (GWAS) for six salt tolerance traits identify 11 significant loci, 4 of which are within ∼300 kb of genomic regions that possess signatures of positive selection, suggesting adaptive geographical divergence for salt tolerance in this species.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
BioProject
Sequence Read Archive
Referenced accessions
Sequence Read Archive
References
Carney, J.A. Black Rice: The African Origins of Rice Cultivation in the Americas (Harvard University Press, 2002).
Linares, O.F. African rice (Oryza glaberrima): history and future potential. Proc. Natl. Acad. Sci. USA 99, 16360–16365 (2002).
Wang, M. et al. The genome sequence of African rice (Oryza glaberrima) and evidence for independent domestication. Nat. Genet. 46, 982–988 (2014).
Sarla, N. & Mallikarjuna, S.B.P. Oryza glaberrima: a source for improvement of Oryza sativa. Curr. Sci. 89, 955–963 (2005).
Agnoun, A. et al. The African rice Oryza glaberrima Steud: knowledge distribution and prospects. Int. J. Biol. 4, 158–180 (2012).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Kahle, D. & Wickham, H. ggmap: spatial visualization with ggplot2. R J. 5, 144–161 (2013).
Falush, D., Stephens, M. & Pritchard, J.K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567–1587 (2003).
Portères, R. in Papers in African Prehistory (eds. Fage, J.D. & Oliver, R.A.) 43–58 (Cambridge University Press, 1970).
Portères, R. in Origins of African Plant Domestication (eds. Harlan, J.R., De Wet, J.M. & Stemler, A.B.) 409–452 (Mouton, 1976).
Pickrell, J.K. & Pritchard, J.K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, e1002967 (2012).
Tenaillon, M.I., U'Ren, J., Tenaillon, O. & Gaut, B.S. Selection versus demography: a multilocus investigation of the domestication process in maize. Mol. Biol. Evol. 21, 1214–1225 (2004).
Caicedo, A.L. et al. Genome-wide patterns of nucleotide polymorphism in domesticated rice. PLoS Genet. 3, 1745–1756 (2007).
Schiffels, S. & Durbin, R. Inferring human population size and separation history from multiple genome sequences. Nat. Genet. 46, 919–925 (2014).
Thomas, C.G. et al. Full-genome evolutionary histories of selfing, splitting, and selection in Caenorhabditis. Genome Res. 25, 667–678 (2015).
Nabholz, B. et al. Transcriptome population genomics reveals severe bottleneck and domestication cost in the African rice (Oryza glaberrima). Mol. Ecol. 23, 2210–2227 (2014).
Eyre-Walker, A., Gaut, R.L., Hilton, H., Feldman, D.L. & Gaut, B.S. Investigation of the bottleneck leading to the domestication of maize. Proc. Natl. Acad. Sci. USA 95, 4441–4446 (1998).
Hyten, D.L. et al. Impacts of genetic bottlenecks on soybean genome diversity. Proc. Natl. Acad. Sci. USA 103, 16666–16671 (2006).
Zhu, Q., Zheng, X., Luo, J., Gaut, B.S. & Ge, S. Multilocus analysis of nucleotide variation of Oryza sativa and its wild relatives: severe bottleneck during domestication of rice. Mol. Biol. Evol. 24, 875–888 (2007).
Tjallingii, R. et al. Coherent high- and low-latitude control of the northwest African hydrological balance. Nat. Geosci. 1, 670–675 (2008).
Manning, K. & Timpson, A. The demographic response to Holocene climate change in the Sahara. Quat. Sci. Rev. 101, 28–35 (2014).
Murray, S.S. in Fields of Change: Progress in African Archaeobotany (ed. Cappers, R.T.J.) 53–62 (Barkhuis, 2007).
Zach, B. & Klee, M. Four thousand years of plant exploitation in the Chad Basin of NE Nigeria. II: Discussion on the morphology of caryopses of domesticated Pennisetum and complete catalog of the fruits and seeds of Kursakata. Veg. Hist. Archaeobot. 12, 187–204 (2003).
Fuller, D.Q., Nixon, S., Stevens, C.J. & Murray, M.A. in The Archaeology of African Plant Use (eds. Stevens, C., Nixon, S., Murray, M.A. & Fuller, D.Q.) 17–24 (Left Coast Press, 2014).
Eichhorn, B. & Neumann, K. in Archaeology of African Plant Use (eds. Stevens, C., Nixon, S., Murray, M.A. & Fuller, D.Q.) (Left Coast Press, 2014).
Temudo, M. Planting knowledge, harvesting agro-biodiversity: a case study of Southern Guinea-Bissau rice farming. Hum. Ecol. 39, 301–321 (2011).
Carney, J.A. Landscapes of technology transfer: rice cultivation and African continuities. Technol. Cult. 37, 5–35 (1996).
International Rice Research Institute. Standard Evaluation System for Rice (International Rice Research Institute, 2014).
Matlon, P., Randolph, T. & Guei, R. in Impact of Rice Research (eds. Pingali, P.B. & Hossain, M.) 382–404 (International Rice Research Institute, 1998).
Huang, X. et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 42, 961–967 (2010).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Kang, H.M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Yang, T. et al. The role of a potassium transporter OsHAK5 in potassium acquisition and transport from roots to shoots in rice at low potassium supply levels. Plant Physiol. 166, 945–959 (2014).
Horie, T. et al. Rice sodium-insensitive potassium transporter, OsHAK5, confers increased salt tolerance in tobacco BY2 cells. J. Biosci. Bioeng. 111, 346–356 (2011).
Chen, H., Patterson, N. & Reich, D. Population differentiation as a test for selective sweeps. Genome Res. 20, 393–402 (2010).
Weir, B.S. & Cockerham, C.C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358–1370 (1984).
Lewontin, R.C. & Krakauer, J. Distribution of gene frequency as a test of the theory of the selective neutrality of polymorphisms. Genetics 74, 175–195 (1973).
Sahi, C., Singh, A., Blumwald, E. & Grover, A. Beyond osmolytes and transporters: novel plant salt-stress tolerance-related genes from transcriptional profiling data. Physiol. Plant. 127, 1–9 (2006).
Ruan, S.L. et al. Proteomic identification of OsCYP2, a rice cyclophilin that confers salt tolerance in rice (Oryza sativa L.) seedlings when overexpressed. BMC Plant Biol. 11, 34 (2011).
Allaby, R.G., Fuller, D.Q. & Brown, T.A. The genetic expectations of a protracted model for the origins of domesticated crops. Proc. Natl. Acad. Sci. USA 105, 13982–13986 (2008).
Fuller, D.Q. Contrasting patterns in crop domestication and domestication rates: recent archaeobotanical insights from the Old World. Ann. Bot. 100, 903–924 (2007).
Wilcox, G. in Biodiversity in Agriculture: Domestication, Evolution, and Sustainability (eds. Gepts, P. et al.) 92–109 (Cambridge University Press, 2012).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Flowers, J.M. et al. Whole-genome resequencing reveals extensive natural variation in the model green alga Chlamydomonas reinhardtii. Plant Cell 27, 2353–2369 (2015).
Hazzouri, K.M. et al. Whole genome re-sequencing of date palms yields insights into diversification of a fruit tree crop. Nat. Commun. 6, 8824 (2015).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Korneliussen, T.S., Albrechtsen, A. & Nielsen, R. ANGSD: analysis of next generation sequencing data. BMC Bioinformatics 15, 356 (2014).
Earl, D.A. & vonHoldt, B.M. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4, 359–361 (2012).
Jakobsson, M. & Rosenberg, N.A. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23, 1801–1806 (2007).
Rosenberg, N. Distruct: a program for the graphical display of population structure. Mol. Ecol. Notes 4, 137–138 (2004).
Gronau, I., Hubisz, M.J., Gulko, B., Danko, C.G. & Siepel, A. Bayesian inference of ancient human demography from individual genome sequences. Nat. Genet. 43, 1031–1034 (2011).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Gaut, B.S., Morton, B.R., McCaig, B.C. & Clegg, M.T. Substitution rate comparisons between grasses and palms: synonymous rate differences at the nuclear gene Adh parallel rate differences at the plastid gene rbcL. Proc. Natl. Acad. Sci. USA 93, 10274–10279 (1996).
Pearson, G., Ayers, S. & Eberhard, D. Relative salt tolerance of rice during germination and early seedling development. Soil Sci. 102, 151–156 (1966).
Gregorio, G.B., Senadhira, D. & Mendoza, R.D. Screening Rice for Salinity Tolerance IRRI Discussion Paper Series 22 (IRRI, 1997).
Platten, J.D., Egdane, J.A. & Ismail, A.M. Salinity tolerance, Na+ exclusion and allele mining of HKT1;5 in Oryza sativa and O. glaberrima: many sources, many genes, one mechanism? BMC Plant Biol. 13, 32 (2013).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2015).
Wu, J. et al. Physical maps and recombination frequency of six rice chromosomes. Plant J. 36, 720–730 (2003).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Turner, S.D. qqman: an R package for visualizing GWAS results using QQ and Manhattan plots. Preprint at bioRxiv http://dx.doi.org/10.1101/005165 (2014).
Kersey, P.J. et al. Ensembl Genomes 2016: more genomes, more complexity. Nucleic Acids Res. 44 D1, D574–D580 (2016).
Untergasser, A. et al. Primer3—new capabilities and interfaces. Nucleic Acids Res. 40, e115 (2012).
Thornton, B. & Basu, C. Real-time PCR (qPCR) primer design using free online software. Biochem. Mol. Biol. Educ. 39, 145–154 (2011).
Fischer, G. et al. Global Agro-ecological Zones Assessment for Agriculture (IASA and FAO, 2008).
Fritz, S. et al. Cropland for sub-Saharan Africa: a synergistic approach using five land cover data sets. Geophys. Res. Lett. 38, L04404 (2011).
Acknowledgements
We would like to thank E. Septiningsih for critical discussions. We are grateful to M. Sock and B. Fonton for field assistance, to International Rice Research Institute staff for phenotyping assistance, and to J. Maritz and Z. Joly-Lopez for laboratory assistance. We thank the US Department of Agriculture and International Rice Research Institute for providing germplasm. This work was funded in part by grants from the National Science Foundation Plant Genome Research Program (IOS-1126971), the Zegar Family Foundation and the New York University Abu Dhabi Research Institute to M.D.P., as well as by a National Science Foundation Plant Genome Postdoctoral Fellowship (IOS-1202803) to R.S.M.
Author information
Authors and Affiliations
Contributions
R.S.M., G.B.G. and M.D.P. designed the experiments and analyses. I.K.B. and M.-N.N. helped in design and execution of the fieldwork in Senegal and Togo, respectively. R.S.M., M.S., A.P., J.A., A.B., K.D., B.G. and G.B.G. collected the data. R.S.M., J.Y.C., J.M.F., K.M.H. and M.D.P. analyzed the data. R.S.M., J.Y.C., D.Q.F. and M.D.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 4 and 11, and Supplementary Note (PDF 530 kb)
Supplementary Table 1
Sample data set. (XLSX 500 kb)
Supplementary Table 2
Sanger sequencing to genotype SNPs. (XLSX 494 kb)
Supplementary Table 3
Principal-component analysis of SNP variation. (XLSX 496 kb)
Supplementary Table 5
Salt tolerance phenotypes. (XLSX 35 kb)
Supplementary Table 6
Kruskal–Wallis results of phenotypes. (XLSX 75 kb)
Supplementary Table 7
Kruskal–Wallis pairwise comparisons of phenotypes between geographic populations. (XLSX 72 kb)
Supplementary Table 8
GWAS results. (XLSX 43 kb)
Supplementary Table 9
Significant 10-kb window coordinates and their maximum XPCLR values. (XLSX 40 kb)
Supplementary Table 10
Significant FST regions between NW coast and SW coast populations. (XLSX 488 kb)
Rights and permissions
About this article
Cite this article
Meyer, R., Choi, J., Sanches, M. et al. Domestication history and geographical adaptation inferred from a SNP map of African rice. Nat Genet 48, 1083–1088 (2016). https://doi.org/10.1038/ng.3633
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3633
This article is cited by
-
Resequencing of Rosa rugosa accessions revealed the history of population dynamics, breed origin, and domestication pathways
BMC Plant Biology (2023)
-
Genome sequencing reveals the genetic architecture of heterostyly and domestication history of common buckwheat
Nature Plants (2023)
-
Multiple domestications of Asian rice
Nature Plants (2023)
-
Population-genomic analyses reveal bottlenecks and asymmetric introgression from Persian into iron walnut during domestication
Genome Biology (2022)
-
The spinach YY genome reveals sex chromosome evolution, domestication, and introgression history of the species
Genome Biology (2022)