Abstract
Plasmodium vivax is a major public health burden, responsible for the majority of malaria infections outside Africa. We explored the impact of demographic history and selective pressures on the P. vivax genome by sequencing 182 clinical isolates sampled from 11 countries across the globe, using hybrid selection to overcome human DNA contamination. We confirmed previous reports of high genomic diversity in P. vivax relative to the more virulent Plasmodium falciparum species; regional populations of P. vivax exhibited greater diversity than the global P. falciparum population, indicating a large and/or stable population. Signals of natural selection suggest that P. vivax is evolving in response to antimalarial drugs and is adapting to regional differences in the human host and the mosquito vector. These findings underline the variable epidemiology of this parasite species and highlight the breadth of approaches that may be required to eliminate P. vivax globally.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
World Health Organization, World Malaria Report (2014).
Neafsey, D.E. et al. The malaria parasite Plasmodium vivax exhibits greater genetic diversity than Plasmodium falciparum. Nat. Genet. 44, 1046–1050 (2012).
Carter, R. Speculations on the origins of Plasmodium vivax malaria. Trends Parasitol. 19, 214–219 (2003).
Melnikov, A. et al. Hybrid selection for sequencing pathogen genomes from clinical samples. Genome Biol. 12, R73 (2011).
Carlton, J.M. et al. Population genetics, evolutionary genomics, and genome-wide studies of malaria: a view across the International Centers of Excellence for Malaria Research. Am. J. Trop. Med. Hyg. 93 (suppl.), 87–98 (2015).
Carlton, J.M. et al. Comparative genomics of the neglected human malaria parasite Plasmodium vivax. Nature 455, 757–763 (2008).
Koepfli, C. et al. Plasmodium vivax diversity and population structure across four continents. PLoS Negl. Trop. Dis. 9, e0003872 (2015).
Liu, W. et al. African origin of the malaria parasite Plasmodium vivax. Nat. Commun. 5, 3346 (2014).
Culleton, R. et al. The origins of African Plasmodium vivax; insights from mitochondrial genome sequencing. PLoS One 6, e29137 (2011).
Rodrigues, P.T. et al. Using mitochondrial genome sequences to track the origin of imported Plasmodium vivax infections diagnosed in the United States. Am. J. Trop. Med. Hyg. 90, 1102–1108 (2014).
Baniecki, M.L. et al. Development of a single nucleotide polymorphism barcode to genotype Plasmodium vivax infections. PLoS Negl. Trop. Dis. 9, e0003539 (2015).
Taylor, J.E. et al. The evolutionary history of Plasmodium vivax as inferred from mitochondrial genomes: parasite genetic diversity in the Americas. Mol. Biol. Evol. 30, 2050–2064 (2013).
Miller, L.H., Mason, S.J., Clyde, D.F. & McGinniss, M.H. The resistance factor to Plasmodium vivax in blacks. The Duffy-blood-group genotype, FyFy. N. Engl. J. Med. 295, 302–304 (1976).
Reich, D., Thangaraj, K., Patterson, N., Price, A.L. & Singh, L. Reconstructing Indian population history. Nature 461, 489–494 (2009).
Ménard, D. et al. Plasmodium vivax clinical malaria is commonly observed in Duffy-negative Malagasy people. Proc. Natl. Acad. Sci. USA 107, 5967–5971 (2010).
Rice, B.L. et al. The origin and diversification of the merozoite surface protein 3 (msp3) multi-gene family in Plasmodium vivax and related parasites. Mol. Phylogenet. Evol. 78, 172–184 (2014).
Arisue, N., Hirai, M., Arai, M., Matsuoka, H. & Horii, T. Phylogeny and evolution of the SERA multigene family in the genus Plasmodium. J. Mol. Evol. 65, 82–91 (2007).
Tachibana, S. et al. Plasmodium cynomolgi genome sequences provide insight into Plasmodium vivax and the monkey malaria clade. Nat. Genet. 44, 1051–1055 (2012).
Molina-Cruz, A. et al. The human malaria parasite Pfs47 gene mediates evasion of the mosquito immune system. Science 340, 984–987 (2013).
Molina-Cruz, A. & Barillas-Mury, C. The remarkable journey of adaptation of the Plasmodium falciparum malaria parasite to New World anopheline mosquitoes. Mem. Inst. Oswaldo Cruz 109, 662–667 (2014).
Moreno, M. et al. Complete mtDNA genomes of Anopheles darlingi and an approach to anopheline divergence time. Malar. J. 9, 127 (2010).
Anthony, T.G., Polley, S.D., Vogler, A.P. & Conway, D.J. Evidence of non-neutral polymorphism in Plasmodium falciparum gamete surface protein genes Pfs47 and Pfs48/45. Mol. Biochem. Parasitol. 156, 117–123 (2007).
Kwiatkowski, D.P. How malaria has affected the human genome and what human genetics can teach us about malaria. Am. J. Hum. Genet. 77, 171–192 (2005).
Gething, P.W. et al. A long neglected world malaria map: Plasmodium vivax endemicity in 2010. PLoS Negl. Trop. Dis. 6, e1814 (2012).
Lacroix, C. & Ménard, R. TRAP-like protein of Plasmodium sporozoites: linking gliding motility to host-cell traversal. Trends Parasitol. 24, 431–434 (2008).
Thompson, J. et al. Plasmodium cysteine repeat modular proteins 1-4: complex proteins with roles throughout the malaria parasite life cycle. Cell. Microbiol. 9, 1466–1480 (2007).
Chuquiyauri, R. et al. Genome-scale protein microarray comparison of human antibody responses in Plasmodium vivax relapse and reinfection. Am. J. Trop. Med. Hyg. 93, 801–809 (2015).
Pacheco, M.A. et al. Evidence of purifying selection on merozoite surface protein 8 (MSP8) and 10 (MSP10) in Plasmodium spp. Infect. Genet. Evol. 12, 978–986 (2012).
Mbengue, A. et al. A molecular mechanism of artemisinin resistance in Plasmodium falciparum malaria. Nature 520, 683–687 (2015).
Schousboe, M.L. et al. Multiple origins of mutations in the mdr1 gene—a putative marker of chloroquine resistance in P. vivax. PLoS Negl. Trop. Dis. 9, e0004196 (2015).
Sidhu, A.B., Verdier-Pinard, D. & Fidock, D.A. Chloroquine resistance in Plasmodium falciparum malaria parasites conferred by pfcrt mutations. Science 298, 210–213 (2002).
Ariey, F. et al. A molecular marker of artemisinin-resistant Plasmodium falciparum malaria. Nature 505, 50–55 (2014).
Rubio, J.M. et al. Semi-nested, multiplex polymerase chain reaction for detection of human malaria parasites and evidence of Plasmodium vivax infection in Equatorial Guinea. Am. J. Trop. Med. Hyg. 60, 183–187 (1999).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Tournamille, C., Colin, Y., Cartron, J.P. & Le Van Kim, C. Disruption of a GATA motif in the Duffy gene promoter abolishes erythroid gene expression in Duffy-negative individuals. Nat. Genet. 10, 224–228 (1995).
Ye, K., Schulz, M.H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Menard, D. et al. Whole genome sequencing of field isolates reveals a common duplication of the Duffy binding protein gene in Malagasy Plasmodium vivax strains. PLoS Negl. Trop. Dis. 7, e2489 (2013).
Li, H. et al. 1000 Genome Project Data Processing Subgroup. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Bertels, F., Silander, O.K., Pachkov, M., Rainey, P.B. & van Nimwegen, E. Automated reconstruction of whole-genome phylogenies from short-sequence reads. Mol. Biol. Evol. 31, 1077–1088 (2014).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Van der Auwera, G.A. et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinformatics 43, 11.10.1–11.10.33 (2013).
Zheng, X. et al. A high-performance computing toolset for relatedness and principal component analysis of SNP data. Bioinformatics 28, 3326–3328 (2012).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Alexander, D.H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Daniels, R.F. et al. Modeling malaria genomics reveals transmission decline and rebound in Senegal. Proc. Natl. Acad. Sci. USA 112, 7067–7072 (2015).
McDonald, J.H. & Kreitman, M. Adaptive protein evolution at the Adh locus in Drosophila. Nature 351, 652–654 (1991).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Storey, J.D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 100, 9440–9445 (2003).
Smith, N.G. & Eyre-Walker, A. Adaptive protein evolution in Drosophila. Nature 415, 1022–1024 (2002).
Acknowledgements
We acknowledge J. Bochicchio and S. Chapman for project management, A. Gnirke for technical support, and members of the Broad Institute Genomics Platform and NYU's Genomics Core for data generation. We thank F. Santillan and P. Michon for technical assistance and MR4 for providing us with malaria parasites deposited by W.E. Collins. The following grants supported this work: National Institute of Allergy and Infectious Diseases (NIAID)/National Institutes of Health (NIH) International Centers of Excellence for Malaria Research U19AI089676 to J.M.C.; U19AI089681, K24AI068903 and D43TW007120 to J.M.V.; U19AI089672 to L.C.; São Paulo Research Foundation 2009/52729-9 to M.U.F.; National Council for Science and Technology Mexico 29005-M SALUD-2004-119 and National Institute of Public Health Mexico project 476191 to L.G.-C.; Victorian State Government Operational Infrastructure Support and Australian Government National Health and Medical Research Council Independent Medical Research Institutes Infrastructure Support Scheme (NHMRC IRIISS) to A.B. and I.M.; 5U19AI089702 to S.H. and M.A.-H.; Armed Forces Health Surveillance Center, Global Emerging Infections Surveillance and Response System and US NIH grant D43TW007393 to A.G.L.; NIH U19AI089686 to J.W.K.; and Bill and Melinda Gates Foundation grant to J.S. Sequencing and analysis work at the Broad Institute was supported by federal funds from the NIAID, NIH, US Department of Health and Human Services, under contract HHSN272200900018C. M.U.F. is supported by a senior research scholarship from the Conselho Nacional de Desenvolvimento Científico e Tecnológico of Brazil, I.M. is supported by NHMRC senior research fellowship 1043345 and D.N.H. is supported by NIH training grant T32AI007180. The content of this manuscript is solely the responsibility of the authors and does not necessarily represent the official policy or position of the US Department of the Navy, the US Department of Defense, the US government or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
J.M.C., I.M. and D.E.N. conceived and conducted the study. A.M., P.L.S., P.R., A.F.V., Q.F., Y.W., C.M.L., S.D., J.F.S., M.L., C.B., D.K., W.R., W.N. and M.K. undertook field and/or wet-lab work and sequencing of the samples. D.N.H., Z.L., J.M.C. and D.E.N. analyzed data. D.N.H., J.M.C., Z.L. and D.E.N. wrote the manuscript, and A.E., S.H., M.A.-H., L.C., G.C.B., A.G.L., A.B., I.M., J.W.K., A.E., N.V., M.U.F., J.S., D.G., J.M.V., L.G.-C. and B.W.B. revised the manuscript and made comments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Evaluation of the performance of different hybrid selection baits on a sample with 0.27% initial P. vivax mappable reads.
(a) Enrichment of P. vivax reads using six different baits, including synthetic oligonucleotides (far left column), ‘whole-genome baits’ (WGB) constructed from individual sources (Brazil I, India VII, N. Korean and Mauritania) of genomic DNA, and a WGB mixture (far right column). The vertical axis indicates percent mappable reads following hybrid selection, and numerals above bars indicate fold enrichment in mappable reads. (b) The fold enrichment of P. vivax DNA using WGB (vertical axis) was compromised by contamination of genomic DNA with host (in this case, monkey) material (horizontal axis).
Supplementary Figure 3 Determination of complexity of infection using variant calls from 195 P. vivax isolates.
Each column along the x axis represents one isolate, and the scale along the y axis represents the number of high-quality variants annotated as heterozygous within that isolate. Isolates exhibiting more than 1,236 heterozygous calls, which represents twice the median observed in the population (vertical dotted line), were classified as complex infections containing more than one haploid parasite lineage.
Supplementary Figure 4 Region-specific projections of the variation data that use two principal components and are limited to Old World and New World populations.
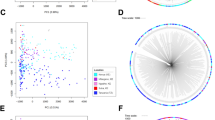
(a) Two-eigenvector PCA limited to Old World isolates. (b) Two-eigenvector PCA limited to New World isolates. Both analyses show similar results to the PCA in Figure 3a that uses the global population of 195 P. vivax isolates.
Supplementary Figure 5 Analysis of the incidence and extent of genomic regions exhibiting identity by descent (IBD) within subpopulations.
This analysis was limited to populations with at least 12 single-infection isolates that were also classified as high quality as listed in Supplementary Table 1.
Supplementary Figure 6 Three maximum-likelihood trees computed by RAxML and REALPHY software that use three different reference genomes (Salvador I, Mauritania I and North Korean) and a subset of high-quality sequenced P. vivax isolates.
The tree is rooted at the node between the Old World and Indian/African isolates in each case. Highlighted in red is the reference genome used, and the bootstrap support is shown at each node (100 bootstraps).
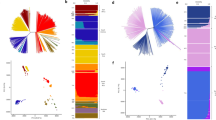
Supplementary Figure 7 Admixture analysis of 195 isolates.
Admixture plots under six different K cluster values. Colors correspond to K clusters within each graph, whereas columns are consistent across graphs and correspond to geographical population. The tenfold cross-validation error associated with each admixture analysis was minimized at K = 5.
Supplementary Figure 8 A comparison of the per-gene fixation index (FST) between New World and Old World isolates and per-gene nucleotide diversity calculated in each case from 73 single-infection, high-quality samples.
In blue are genes annotated as antigens, including members of the P. vivax serine-repeat antigen (SERA) family; members of the merozoite surface protein (MSP) superfamily, including single-copy genes MSP1, MSP4, MSP5, MSP8, MSP9, MSP10 and several members of the MSP3 multigene family and the MSP7 multigene family; members of the variant interspersed repeat (vir) gene family (excluding the most highly conserved and potential founder gene PVX_113230); members of the Pv-fam-a (PvTRAG), Pv-fam-b, Pv-fam-c, Pv-fam-d (HYPB), Pv-fam-e (RAD), Pv-fam-g, Pv-fam-h (HYP16) and Pv-fam-i (HYP11) gene families; and any gene annotated as an antigen in the Salvador I reference genome annotation. In green are genes identified as being putatively involved in antimalarial drug resistance, including dihydrofolate reductase–thymidylate synthase (DHFR-TS), dihydropteroate synthase (DHPS), multidrug resistance 1 protein (MDR1) and chloroquine resistance transporter gene (PvCRT). Gray dots represent all other annotated genes for which the two statistics could be calculated.
Supplementary Figure 9 Results from the McDonald–Kreitman (MK) test using a subset of single-infection and high-quality isolates from Colombia, Peru, Mexico, Thailand, Myanmar and Papua New Guinea.
Codon gapped-nucleotide alignments for each gene in the P. vivax genome were created and aligned to the nearest P. cynomolgi ortholog where available; genes without a one-to-one ortholog were excluded. Plotted along the y axis is the χ2 value associated with the MK test for each gene within the P. vivax alignment set. From this χ2 value, a P value was calculated. A q-value FDR correction was added to account for the effects of multiple sampling. Genes that were found to be significant (q value < 0.05) are shown in red. α, an estimate of the proportion of bases fixed by positive selection, is plotted along the x axis and ranges from ∞ (infinity) to 1. Values above 0 are inferred to be under positive selection, whereas values below zero are subject to purifying selection and/or balancing selection. Significant results are also listed in Supplementary Table 3.
Supplementary Figure 10 Plots of five different population genetic values across chromosome 5 of P. vivax.
In each case, the statistics were calculated using high-quality, single-infection isolates. Values of linkage disequilibrium (LD) are collapsed into a mean value across a window of 50 kb for each nucleotide. In the case of nucleotide diversity (π), Tajima’s D and FST, values represent the mean value across all nucleotides within a 1-kb window. Plotted along the y axis in each figure is a trendline representing a moving average (period 10) for each population genetic statistic. The vertical red line indicates genes of interest identified through high FST values as labeled in Figure 5 or genes with significant McDonald–Kreitman test results as listed in Supplementary Table 3. The thickness of the red line is an approximation of gene length.
Supplementary Figure 11 Plots of five different population genetic values across chromosome 7 of P. vivax.
In each case, the statistics were calculated using high-quality, single-infection isolates. Values of linkage disequilibrium (LD) are collapsed into a mean value across a window of 50 kb for each nucleotide. In the case of nucleotide diversity (π), Tajima’s D and FST, values represent the mean value across all nucleotides within a 1-kb window. Plotted along the y axis in each figure is a trendline representing a moving average (period 10) for each population genetic statistic. The vertical red line indicates genes of interest identified through high FST values as labeled in Figure 5 or genes with significant McDonald–Kreitman test results as listed in Supplementary Table 3. The thickness of the red line is an approximation of gene length.
Supplementary Figure 12 Plots of six different population genetic values across chromosome 9 of P. vivax.
In each case, the statistics were calculated using high-quality, single-infection isolates. Values of linkage disequilibrium (LD) are collapsed into a mean value across a window of 50 kb for each nucleotide. In the case of nucleotide diversity (π), Tajima’s D and FST, values represent the mean value across all nucleotides within a 1-kb window. Plotted along the y axis in each figure is a trendline representing a moving average (period 10) for each population genetic statistic. The vertical red line indicates genes of interest identified through high FST values as labeled in Figure 5 or genes with significant McDonald–Kreitman test results as listed in Supplementary Table 3. The thickness of the red line is an approximation of gene length.
Supplementary Figure 13 Plots of six different population genetic values across chromosome 11 of P. vivax.
In each case, the statistics were calculated using high-quality, single-infection isolates. Values of linkage disequilibrium (LD) are collapsed into a mean value across a window of 50 kb for each nucleotide. In the case of nucleotide diversity (π), Tajima’s D and FST, values represent the mean value across all nucleotides within a 1-kb window. Plotted along the y axis in each figure is a trendline representing a moving average (period 10) for each population genetic statistic. The vertical red line indicates genes of interest identified through high FST values as labeled in Figure 5 or genes with significant McDonald–Kreitman test results as listed in Supplementary Table 3. The thickness of the red line is an approximation of gene length.
Supplementary Figure 14 Plots of six different population genetic values across chromosome 12 of P. vivax.
In each case, the statistics were calculated using high-quality, single-infection isolates. Values of linkage disequilibrium (LD) are collapsed into a mean value across a window of 50 kb for each nucleotide. In the case of nucleotide diversity (π), Tajima’s D and FST, values represent the mean value across all nucleotides within a 1-kb window. Plotted along the y axis in each figure is a trendline representing a moving average (period 10) for each population genetic statistic. The vertical red line indicates genes of interest identified through high FST values as labeled in Figure 5 or genes with significant McDonald–Kreitman test results as listed in Supplementary Table 3. The thickness of the red line is an approximation of gene length.
Supplementary Figure 15 Plots of six different population genetic values across chromosome 14 of P. vivax.
In each case, the statistics were calculated using high-quality, single-infection isolates. Values of linkage disequilibrium (LD) are collapsed into a mean value across a window of 50 kb for each nucleotide. In the case of nucleotide diversity (π), Tajima’s D and FST, values represent the mean value across all nucleotides within a 1-kb window. Plotted along the y axis in each figure is a trendline representing a moving average (period 10) for each population genetic statistic. The vertical red line indicates genes of interest identified through high FST values as labeled in Figure 5 or genes with significant McDonald–Kreitman test results as listed in Supplementary Table 3. The thickness of the red line is an approximation of gene length.
Supplementary Figure 16 Haplotype map of three genes found to exhibit signals of positive selection within P. vivax.
Shown in red are nonsynonymous alternative alleles within each gene. In gray are sites that are identical to the reference genome nucleotide. Blank entries (white) indicate missing data.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–16, Supplementary Table 4 and Supplementary Note. (PDF 3319 kb)
Supplementary Table 1
Genome statistics and metadata. Country of origin, GenBank accession number, and various in silico and molecular biology assay results for the 195 P. vivax isolates analyzed in this study. (XLSX 151 kb)
Supplementary Table 2
Genes of interest. List of nonsynonymous SNPs, insertions, and deletions in 22 genes identified as, or being associated with, genes having high FST. (XLSX 71 kb)
Supplementary Table 3
Summary of results from the McDonald–Kreitman test. (XLSX 536 kb)
Source data
Rights and permissions
About this article
Cite this article
Hupalo, D., Luo, Z., Melnikov, A. et al. Population genomics studies identify signatures of global dispersal and drug resistance in Plasmodium vivax. Nat Genet 48, 953–958 (2016). https://doi.org/10.1038/ng.3588
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3588
This article is cited by
-
Genomics of Plasmodium vivax in Colombia reveals evidence of local bottle-necking and inter-country connectivity in the Americas
Scientific Reports (2023)
-
Etiology of acute febrile illness in the peruvian amazon as determined by modular formatted quantitative PCR: a protocol for RIVERA, a health facility-based case-control study
BMC Public Health (2023)
-
Genomic analysis of Plasmodium vivax describes patterns of connectivity and putative drivers of adaptation in Ethiopia
Scientific Reports (2023)
-
Evolutionary history of Plasmodium vivax and Plasmodium simium in the Americas
Malaria Journal (2022)
-
Genetic diversity of Plasmodium vivax reticulocyte binding protein 2b in global parasite populations
Parasites & Vectors (2022)