Abstract
RELATIVE OF EARLY FLOWERING 6 (REF6, also known as JMJ12) counteracts Polycomb-mediated gene silencing by removing methyl groups from trimethylated histone H3 lysine 27 (H3K27me3) in hundreds of genes in Arabidopsis thaliana1. Here we show that REF6 function and genome-wide targeting require its four Cys2His2 zinc fingers, which directly recognize a CTCTGYTY motif. Motifs bound by REF6 tend to cluster and reside in loci with active chromatin states. Furthermore, REF6 targets CUP-SHAPED COTYLEDON 1 (CUC1), which harbors CTCTGYTY motifs, to modulate H3K27me3 levels and activate CUC1 expression. Loss of REF6 causes CUC1 repression and defects in cotyledon separation. In contrast, REF6 does not bind CUC2, encoding a close homolog of CUC1, which lacks the CTCTGYTY motif. Collectively, these results identify a new targeting mechanism of an H3K27 demethylase to counteract Polycomb-mediated gene silencing that regulates plant development, including organ boundary formation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Lu, F., Cui, X., Zhang, S., Jenuwein, T. & Cao, X. Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat. Genet. 43, 715–719 (2011).
Holec, S. & Berger, F. Polycomb group complexes mediate developmental transitions in plants. Plant Physiol. 158, 35–43 (2012).
Margueron, R. & Reinberg, D. The Polycomb complex PRC2 and its mark in life. Nature 469, 343–349 (2011).
Steffen, P.A. & Ringrose, L. What are memories made of? How Polycomb and Trithorax proteins mediate epigenetic memory. Nat. Rev. Mol. Cell Biol. 15, 340–356 (2014).
Lafos, M. et al. Dynamic regulation of H3K27 trimethylation during Arabidopsis differentiation. PLoS Genet. 7, e1002040 (2011).
Sun, B. et al. Timing mechanism dependent on cell division is invoked by Polycomb eviction in plant stem cells. Science 343, 1248559 (2014).
Wu, M.F. et al. SWI2/SNF2 chromatin remodeling ATPases overcome Polycomb repression and control floral organ identity with the LEAFY and SEPALLATA3 transcription factors. Proc. Natl. Acad. Sci. USA 109, 3576–3581 (2012).
Zhang, X. et al. Whole-genome analysis of histone H3 lysine 27 trimethylation in Arabidopsis. PLoS Biol. 5, e129 (2007).
Hou, X. et al. Nuclear factor Y–mediated H3K27me3 demethylation of the SOC1 locus orchestrates flowering responses of Arabidopsis. Nat. Commun. 5, 4601 (2014).
Noh, B. et al. Divergent roles of a pair of homologous jumonji/zinc-finger-class transcription factor proteins in the regulation of Arabidopsis flowering time. Plant Cell 16, 2601–2613 (2004).
Yu, X. et al. Modulation of brassinosteroid-regulated gene expression by Jumonji domain–containing proteins ELF6 and REF6 in Arabidopsis. Proc. Natl. Acad. Sci. USA 105, 7618–7623 (2008).
Sun, Y. et al. Integration of brassinosteroid signal transduction with the transcription network for plant growth regulation in Arabidopsis. Dev. Cell 19, 765–777 (2010).
Yu, X. et al. A brassinosteroid transcriptional network revealed by genome-wide identification of BESI target genes in Arabidopsis thaliana. Plant J. 65, 634–646 (2011).
Lu, F., Cui, X., Zhang, S., Liu, C. & Cao, X. JMJ14 is an H3K4 demethylase regulating flowering time in Arabidopsis. Cell Res. 20, 387–390 (2010).
Larsson, A.S., Landberg, K. & Meeks-Wagner, D.R. The TERMINAL FLOWER2 (TFL2) gene controls the reproductive transition and meristem identity in Arabidopsis thaliana. Genetics 149, 597–605 (1998).
Klug, A. The discovery of zinc fingers and their applications in gene regulation and genome manipulation. Annu. Rev. Biochem. 79, 213–231 (2010).
Machanick, P. & Bailey, T.L. MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics 27, 1696–1697 (2011).
Bailey, T.L. & Machanick, P. Inferring direct DNA binding from ChIP-seq. Nucleic Acids Res. 40, e128 (2012).
Zhang, S. et al. C-terminal domains of a histone demethylase interact with a pair of transcription factors and mediate specific chromatin association. Cell Discovery 1, 15003 (2015).
van Dijk, K. et al. Dynamic changes in genome-wide histone H3 lysine 4 methylation patterns in response to dehydration stress in Arabidopsis thaliana. BMC Plant Biol. 10, 238 (2010).
Coleman-Derr, D. & Zilberman, D. Deposition of histone variant H2A.Z within gene bodies regulates responsive genes. PLoS Genet. 8, e1002988 (2012).
Sullivan, A.M. et al. Mapping and dynamics of regulatory DNA and transcription factor networks in A. thaliana. Cell Rep. 8, 2015–2030 (2014).
Inagaki, S. et al. Autocatalytic differentiation of epigenetic modifications within the Arabidopsis genome. EMBO J. 29, 3496–3506 (2010).
Aida, M., Ishida, T., Fukaki, H., Fujisawa, H. & Tasaka, M. Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell 9, 841–857 (1997).
Hibara, K. et al. Arabidopsis CUP-SHAPED COTYLEDON3 regulates postembryonic shoot meristem and organ boundary formation. Plant Cell 18, 2946–2957 (2006).
Takada, S., Hibara, K., Ishida, T. & Tasaka, M. The CUP-SHAPED COTYLEDON1 gene of Arabidopsis regulates shoot apical meristem formation. Development 128, 1127–1135 (2001).
Vroemen, C.W., Mordhorst, A.P., Albrecht, C., Kwaaitaal, M.A. & de Vries, S.C. The CUP-SHAPED COTYLEDON3 gene is required for boundary and shoot meristem formation in Arabidopsis. Plant Cell 15, 1563–1577 (2003).
Crevillén, P. et al. Epigenetic reprogramming that prevents transgenerational inheritance of the vernalized state. Nature 515, 587–590 (2014).
Lu, F. et al. Comparative analysis of JmjC domain–containing proteins reveals the potential histone demethylases in Arabidopsis and rice. J. Integr. Plant Biol. 50, 886–896 (2008).
Amborella Genome Project. The Amborella genome and the evolution of flowering plants. Science 342, 1241089 (2013).
Najafabadi, H.S. et al. C2H2 zinc finger proteins greatly expand the human regulatory lexicon. Nat. Biotechnol. 33, 555–562 (2015).
Clough, S.J. & Bent, A.F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Bowler, C. et al. Chromatin techniques for plant cells. Plant J. 39, 776–789 (2004).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Grant, C.E., Bailey, T.L. & Noble, W.S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Finnegan, E.J. & Dennis, E.S. Vernalization-induced trimethylation of histone H3 lysine 27 at FLC is not maintained in mitotically quiescent cells. Curr. Biol. 17, 1978–1983 (2007).
Lin, R. et al. Transposase-derived transcription factors regulate light signaling in Arabidopsis. Science 318, 1302–1305 (2007).
Johnson, M. et al. NCBI BLAST: a better web interface. Nucleic Acids Res. 36, W5–W9 (2008).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Acknowledgements
We thank Q. Zhu for technical assistance and W. Qian for discussion. We thank the Arabidopsis Biological Resource Center for providing T-DNA insertion lines. This work was supported by the National Basic Research Program of China (grants 2013CB967300 to Xia Cui and 2013CB835200 to X. Cao), the National Natural Science Foundation of China (grants 31210103901 to X. Cao, 31271363 to Xia Cui, and 31428011 to X.Z.), and the State Key Laboratory of Plant Genomics (2014B0227-01 and 2015B0129-01). Research in the laboratory of X.Z. was supported by National Science Foundation grant 0960425. B.Z., Xiekui Cui, and X.M. were supported by the China Postdoctoral Science Foundation (2012M520020, 2014M550874, and 2014M550101, respectively).
Author information
Authors and Affiliations
Contributions
Xia Cui, F.L., Q.Q., and X. Cao conceived and designed the study. Xia Cui, F.L., and Q.Q. performed most of the experiments with help from S.Z., Y.K., Xiekui Cui, and Q.Y. High-throughput sequencing data were analyzed by B.Z. with help from L.G. and X.M. J.M. provided essential reagents for the EMSA analysis. Xia Cui, F.L., Q.Q., B.Z., X.Z., and X. Cao interpreted the data. F.L., Q.Q., B.Z., X.Z., and X. Cao wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 REF6 expression in transgenic lines.
(a,b) Expression of REF6 in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants detected by RT–qPCR (a) and immunoblot (b). Gene expression was normalized to that of the control gene TUBULIN2 (AT5G62960). RT-qPCR was performed with four technical replicates. Data are shown as means ± s.e. (n = 4). LHP1 was used as the loading control for the immunoblot. (c) pREF6::REF6ΔZnF-HA cannot rescue the short-petiole phenotypes of ref6 mutants. Scale bar, 1 cm.
Supplementary Figure 2 H3K27me3 hypermethylation in REF6ΔZnF-HA ref6 is similar to that in ref6.
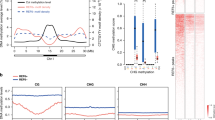
(a) Number of genes showing H3K27me3 hypermethylation in ref6 and REF6ΔZnF-HA ref6 plants. (b) Diagram showing the comparison of H3K27me3-hypermethylated genes in ref6-1 and ref6-3 (ref. 1). The greater number of H3K27me3-hypermethylated genes called in ref6-1 compared to ref6-3 is mainly due to greater sequencing depth in ChIP-seq experiments. (c) Density profile of H3K27me3 in 1,000 randomly selected genes. The H3K27me3 signal was summarized in fixed 1-kb regions around the transcriptional start site (TSS), transcriptional termination site (TTS), and center of the gene.
Supplementary Figure 3 The C2H2-ZnF domains are dispensable for the H3K27me3 and H3K27me2 demethylase activity of REF6.
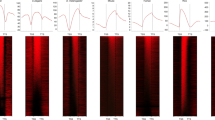
(a) Overexpression of REF6ΔZnF-YFP-HA reduced the levels of H3K27me3 and H3K27me2 in vivo. REF6ΔZnF-YFP-HA fusion protein was transiently expressed in tobacco, and nuclei were isolated for immunostaining. More than 25 pairs of non-transfected nuclei versus transfected nuclei in the same field of view were analyzed. Arrows indicate transfected nuclei. Scale bars, 2 μm. (b) Quantitative analysis of a. Data are shown as means ± s.e. (n = 20). WT, wild type (non-transfected nuclei). (c) REF6ΔZnFox plants showed similar phenotypes to REF6ox plants. The plants showed upward-curling leaves with deeper serrations in the margin, early flowering, and terminal flower phenotypes of various degrees. Scale bar, 1 cm. (d) Activation of Polycomb target genes in REF6ΔZnFox plants. Expression of Polycomb target genes was analyzed by RT–qPCR. Gene expression is normalized to that of the control gene TUBULIN2 (AT5G62960). RT-qPCR was performed with four technical replicates. Data are shown as means ± s.e. (n = 4). (e) H3K27me3 and H3K27me2 levels are reduced in REF6ΔZnFox plants. H3 lysine methylation status is shown in two REF6ΔZnFox lines and one REF6ox line as detected by immunoblotting with the antibodies specified on the right. Immunoblotting with antibody to H3 showed equal loading.
Supplementary Figure 4 REF6 direct targets identified by ChIP-seq with antibody to HA.
(a) Density profile of REF6 binding signals in 1,000 randomly selected genes. REF6 binding signal was summarized in fixed 1-kb regions around the transcriptional start site (TSS), transcriptional termination site (TTS), and center of the gene. (b) Number of genes showing enrichment in REF6-HA ref6 and REF6ΔZnF-HA ref6 plants by ChIP-seq using antibody to HA.
Supplementary Figure 5 Genome Browser view of ChIP-seq data and ChIP–qPCR validation.
(a,b) ChIP-seq data and ChIP–qPCR validation for NAC004 (a) and SUS3 (b) loci. ChIP–qPCR used another biological replicate of samples. Data from ChIP–qPCR analysis are shown as assay-site fold enrichment of the signal from immunoprecipitation over the background. Col was used as the negative-control sample. ChIP–qPCR was performed with four technical replicates. Data are shown as means ± s.e. (n = 4). (c,d) ChIP-seq data at another two genes, HB23 (AT1G26960) (c) and CER3 (AT5G57800) (d). Regions validated by ChIP–qPCR in a and b are marked by black lines on top of the gene model. The locations of CTCTGYTY motifs are indicated by blue bars above the gene models. NA, not analyzed.
Supplementary Figure 6 EMSA showing that the CTCTGYTY motif but not the flanking sequence is important for protein–DNA interaction.
(a,d) Sequences of 50-bp DNA fragments of the NAC04 (a) and CUC1 (d) loci, containing one and two CTCTGTTT motifs, respectively, and their mutant versions used in b, c, and e. WT, wild type. (b) The flanking sequence has minimal effect on DNA–protein interaction. (c) GST-REF6C interacts with probes containing the other three variants of the CTCTGYTY motif. (e) EMSA showing that one CTCTGTTT motif is sufficient for DNA–protein interaction.
Supplementary Figure 8 REF6 binds CUC1 and CUC3 in shoot apical tissues.
(a,b) The CTCTGYTY motifs at the CUC1 (a) and CUC3 (b) loci. The numbers indicate base positions from the TSS. The motifs in the sense strand are labeled in red, and the motifs in the anti-sense strand are labeled in blue. Nucleotides in exons and introns are shown in uppercase and lowercase letters, respectively. Nucleotides in qPCR-detected regions are shown in bold letters. (c,d,f) In shoot apical tissues (Online Methods), HA ChIP–qPCR (c) and RT–qPCR (d,f) showed that REF6 binds CUC3 but does not affect the transcript level of CUC3. (e) Transcript levels of CUC2 and CUC3 are not changed in ref6 mutants (14-d-old seedlings). Data from ChIP–qPCR analysis are shown as assay-site fold enrichment of the signal from immunoprecipitation over the background. Col was used as the negative control. Gene expression is normalized to that of the control gene ACTIN7 (AT5G09810). ChIP–qPCR and RT-qPCR were performed with four technical replicates. Data are shown as means ± s.e. (n = 4). NA, not analyzed.
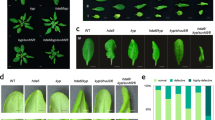
Supplementary Figure 9 Phenotype of ref6 cuc double and triple mutants.
Representative 10-d-old seedlings with different phenotypes are shown. Scale bars, 2 mm.
Supplementary Figure 10 Role of the CTCTGYTY motif and chromatin states in REF6 targeting to activate gene expression through H3K27me3 demethylation.
REF6 binds the CTCTGYTY motif through the C2H2-ZnF cluster and removes local H3K27me3 methylation. Genes containing clusters of CTCTGYTY motifs recruit REF6 more efficiently (top), whereas motifs in heterochromatic regions cannot recruit REF6 (bottom).
Supplementary Figure 11 Phylogenetic tree of REF6 homologs in plants.
The phylogenetic tree was constructed with MEGA (ver.5.05)44 using the neighbor-joining method. The red branches mark the monocot species, and the blue branches mark the eudicots. Amborella trichopoda is the most basal lineage in the clade of angiosperms (‘basal angiosperms’)30. Selaginella moellendorffii and Physcomitrella patens are outgroups.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Table 1. (PDF 2056 kb)
Supplementary Table 2
Chromosomal regions in which H3K27me3 sites were hypermethylated by more than 2-fold in ref6. (XLS 345 kb)
Supplementary Table 3
Chromosomal regions in which H3K27me3 sites were hypermethylated by more than 2-fold in REF6ΔZnF-HA ref6. (XLS 326 kb)
Supplementary Table 4
Chromosomal regions of REF6 binding peaks. (XLS 611 kb)
Supplementary Table 5
REF6 binding peaks for MEME-ChIP analysis. (XLSX 198 kb)
Supplementary Table 6
CTCTGYTY motifs in the Arabidopsis genome. (XLSX 1556 kb)
Supplementary Table 7
List of CTCTGYTY motif clusters. (XLSX 99 kb)
Supplementary Table 8
Primers used in this study. (XLSX 21 kb)
Rights and permissions
About this article
Cite this article
Cui, X., Lu, F., Qiu, Q. et al. REF6 recognizes a specific DNA sequence to demethylate H3K27me3 and regulate organ boundary formation in Arabidopsis. Nat Genet 48, 694–699 (2016). https://doi.org/10.1038/ng.3556
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3556
This article is cited by
-
Gametophytic epigenetic regulators, MEDEA and DEMETER, synergistically suppress ectopic shoot formation in Arabidopsis
Plant Cell Reports (2024)
-
Nearby transposable elements impact plant stress gene regulatory networks: a meta-analysis in A. thaliana and S. lycopersicum
BMC Genomics (2022)
-
Paternal imprinting of dosage-effect defective1 contributes to seed weight xenia in maize
Nature Communications (2022)
-
Bifurcation Analysis of a Heat-Sensitive Epigenetic Regulatory Network
Bulletin of Mathematical Biology (2022)
-
H3K27me3 demethylases alter HSP22 and HSP17.6C expression in response to recurring heat in Arabidopsis
Nature Communications (2021)