Abstract
A more complete understanding of the genetic basis of drug resistance in Mycobacterium tuberculosis is critical for prompt diagnosis and optimal treatment, particularly for toxic second-line drugs such as D-cycloserine. Here we used the whole-genome sequences from 498 strains of M. tuberculosis to identify new resistance-conferring genotypes. By combining association and correlated evolution tests with strategies for amplifying signal from rare variants, we found that loss-of-function mutations in ald (Rv2780), encoding L-alanine dehydrogenase, were associated with unexplained drug resistance. Convergent evolution of this loss of function was observed exclusively among multidrug-resistant strains. Drug susceptibility testing established that ald loss of function conferred resistance to D-cycloserine, and susceptibility to the drug was partially restored by complementation of ald. Clinical strains with mutations in ald and alr exhibited increased resistance to D-cycloserine when cultured in vitro. Incorporation of D-cycloserine resistance in novel molecular diagnostics could allow for targeted use of this toxic drug among patients with susceptible infections.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Accessions
BioProject
NCBI Reference Sequence
Sequence Read Archive
References
Bauer, K.A., Perez, K.K., Forrest, G.N. & Goff, D.A. Review of rapid diagnostic tests used by antimicrobial stewardship programs. Clin. Infect. Dis. 59 (suppl. 3), S134–S145 (2014).
Boehme, C.C. et al. Rapid molecular detection of tuberculosis and rifampin resistance. N. Engl. J. Med. 363, 1005–1015 (2010).
Kiet, V.S. et al. Evaluation of the MTBDRsl test for detection of second-line-drug resistance in Mycobacterium tuberculosis. J. Clin. Microbiol. 48, 2934–2939 (2010).
Bastos, M.L. et al. Treatment outcomes of patients with multidrug-resistant and extensively drug-resistant tuberculosis according to drug susceptibility testing to first- and second-line drugs: an individual patient data meta-analysis. Clin. Infect. Dis. 59, 1364–1374 (2014).
Cegielski, J.P. et al. Extensive drug resistance acquired during treatment of multidrug-resistant tuberculosis. Clin. Infect. Dis. 59, 1049–1063 (2014).
Lange, C. et al. Management of patients with multidrug-resistant/extensively drug-resistant tuberculosis in Europe: a TBNET consensus statement. Eur. Respir. J. 44, 23–63 (2014).
Theron, G. et al. The diagnostic accuracy of the GenoType(®) MTBDRsl assay for the detection of resistance to second-line anti-tuberculosis drugs. Cochrane Database Syst. Rev. 10, CD010705 (2014).
Finegold, S.M. Kanamycin. AMA Arch. Intern. Med. 104, 15–28 (1959).
Murray, F.J. A PILOT study of cycloserine toxicity; a United States Public Health Service Cooperative Clinical Investigation. Am. Rev. Tuberc. 74, 196–209 (1956).
Bankier, R.G. Psychosis associated with cycloserine. Can. Med. Assoc. J. 93, 35–37 (1965).
Boston Collaborative Drug Surveillance Program. Drug-induced deafness. JAMA 224, 515–516 (1973).
Yew, W.W., Wong, C.F., Wong, P.C., Lee, J. & Chau, C.H. Adverse neurological reactions in patients with multidrug-resistant pulmonary tuberculosis after coadministration of cycloserine and ofloxacin. Clin. Infect. Dis. 17, 288–289 (1993).
Woods, G.L. Susceptibility testing for mycobacteria. Clin. Infect. Dis. 31, 1209–1215 (2000).
Kam, K.M. et al. Determination of critical concentrations of second-line anti-tuberculosis drugs with clinical and microbiological relevance. Int. J. Tuberc. Lung Dis. 14, 282–288 (2010).
World Health Organization. Companion Handbook to the WHO Guidelines for the Programmatic Management of Drug-Resistant Tuberculosis (World Health Organization, 2014).
Lambert, M.P. & Neuhaus, F.C. Mechanism of D-cycloserine action: alanine racemase from Escherichia coli W. J. Bacteriol. 110, 978–987 (1972).
Strominger, J.L., Ito, E. & Threnn, R.H. Competitive inhibition of enzymatic reactions by oxamycin. J. Am. Chem. Soc. 82, 998–999 (1960).
Prosser, G.A. & de Carvalho, L.P.S. Metabolomics reveal D-alanine: D-alanine ligase as the target of D-cycloserine in Mycobacterium tuberculosis. ACS Med. Chem. Lett. 4, 1233–1237 (2013).
Cáceres, N.E. et al. Overexpression of the D-alanine racemase gene confers resistance to D-cycloserine in Mycobacterium smegmatis. J. Bacteriol. 179, 5046–5055 (1997).
Feng, Z. & Barletta, R.G. Roles of Mycobacterium smegmatis D-alanine: D-alanine ligase and D-alanine racemase in the mechanisms of action of and resistance to the peptidoglycan inhibitor D-cycloserine. Antimicrob. Agents Chemother. 47, 283–291 (2003).
Köser, C.U. et al. Whole-genome sequencing for rapid susceptibility testing of M. tuberculosis. N. Engl. J. Med. 369, 290–292 (2013).
Merker, M. et al. Whole genome sequencing reveals complex evolution patterns of multidrug-resistant Mycobacterium tuberculosis Beijing strains in patients. PLoS One 8, e82551 (2013).
Rist, N., Canetti, G., Boisvert, H. & Le Lirzin, M. The BCG antibiogram. Diagnostic value of resistance to cycloserine. Rev. Tuberc. Pneumol. (Paris) 31, 1060–1065 (1967).
Chen, J.M., Uplekar, S., Gordon, S.V. & Cole, S.T. A point mutation in cycA partially contributes to the D-cycloserine resistance trait of Mycobacterium bovis BCG vaccine strains. PLoS One 7, e43467 (2012).
Farhat, M.R. et al. Genomic analysis identifies targets of convergent positive selection in drug-resistant Mycobacterium tuberculosis. Nat. Genet. 45, 1183–1189 (2013).
Zhang, H. et al. Genome sequencing of 161 Mycobacterium tuberculosis isolates from China identifies genes and intergenic regions associated with drug resistance. Nat. Genet. 45, 1255–1260 (2013).
Casali, N. et al. Evolution and transmission of drug-resistant tuberculosis in a Russian population. Nat. Genet. 46, 279–286 (2014).
Cohen, K.A. et al. Evolution of extensively drug-resistant tuberculosis over four decades: whole genome sequencing and dating analysis of Mycobacterium tuberculosis isolates from KwaZulu-Natal. PLoS Med. 12, e1001880 (2015).
Wong, S.Y. et al. Mutations in gidB confer low-level streptomycin resistance in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 55, 2515–2522 (2011).
Maus, C.E., Plikaytis, B.B. & Shinnick, T.M. Mutation of tlyA confers capreomycin resistance in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 49, 571–577 (2005).
Morlock, G.P., Metchock, B., Sikes, D., Crawford, J.T. & Cooksey, R.C. ethA, inhA, and katG loci of ethionamide-resistant clinical Mycobacterium tuberculosis isolates. Antimicrob. Agents Chemother. 47, 3799–3805 (2003).
Heym, B., Alzari, P.M., Honoré, N. & Cole, S.T. Missense mutations in the catalase-peroxidase gene, katG, are associated with isoniazid resistance in Mycobacterium tuberculosis. Mol. Microbiol. 15, 235–245 (1995).
Scorpio, A. & Zhang, Y. Mutations in pncA, a gene encoding pyrazinamidase/nicotinamidase, cause resistance to the antituberculous drug pyrazinamide in tubercle bacillus. Nat. Med. 2, 662–667 (1996).
Pagel, M. Detecting correlated evolution on phylogenies: a general method for the comparative analysis of discrete characters. Proc. R. Soc. Lond. B 255, 37–45 (1994).
Lamichhane, G. et al. A postgenomic method for predicting essential genes at subsaturation levels of mutagenesis: application to Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 100, 7213–7218 (2003).
Li, G. et al. Efflux pump gene expression in multidrug-resistant Mycobacterium tuberculosis clinical isolates. PLoS One 10, e0119013 (2015).
Converse, S.E. et al. MmpL8 is required for sulfolipid-1 biosynthesis and Mycobacterium tuberculosis virulence. Proc. Natl. Acad. Sci. USA 100, 6121–6126 (2003).
Reeves, A.Z. et al. Aminoglycoside cross-resistance in Mycobacterium tuberculosis due to mutations in the 5′ untranslated region of whiB7. Antimicrob. Agents Chemother. 57, 1857–1865 (2013).
Köser, C.U., Bryant, J.M., Parkhill, J. & Peacock, S.J. Consequences of whiB7 (Rv3197A) mutations in Beijing genotype isolates of the Mycobacterium tuberculosis complex. Antimicrob. Agents Chemother. 57, 3461 (2013).
Morris, R.P. et al. Ancestral antibiotic resistance in Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 102, 12200–12205 (2005).
Warit, S. et al. Genetic characterisation of a whiB7 mutant of a Mycobacterium tuberculosis clinical strain. J. Glob. Antimicrob. Resist. 3, 262–266 (2015).
Wiame, J.M. & Piérard, A. Occurrence of an L(+)-alanine-dehydrogenase in Bacillus subtilis. Nature 176, 1073–1075 (1955).
Department of Health, South Africa. Management of Drug-Resistant Tuberculosis: Policy Guidelines http://www.health-e.org.za/wp-content/uploads/2014/06/MDR-TB-Clinical-Guidelines-Updated-Jan-2013.pdf (2013).
Durek, C., Rüsch-Gerdes, S., Jocham, D. & Böhle, A. Sensitivity of BCG to modern antibiotics. Eur. Urol. 37 (suppl. 1), 21–25 (2000).
Goh, K.S. & Rastogi, N. Rapid preliminary differentiation of species within the Mycobacterium tuberculosis complex: proposition of a radiometric method. Res. Microbiol. 142, 659–665 (1991).
Garcia Pelayo, M.C. et al. A comprehensive survey of single nucleotide polymorphisms (SNPs) across Mycobacterium bovis strains and M. bovis BCG vaccine strains refines the genealogy and defines a minimal set of SNPs that separate virulent M. bovis strains and M. bovis BCG strains. Infect. Immun. 77, 2230–2238 (2009).
Chen, J.M., Alexander, D.C., Behr, M.A. & Liu, J. Mycobacterium bovis BCG vaccines exhibit defects in alanine and serine catabolism. Infect. Immun. 71, 708–716 (2003).
Garnier, T. et al. The complete genome sequence of Mycobacterium bovis. Proc. Natl. Acad. Sci. USA 100, 7877–7882 (2003).
Sassetti, C.M., Boyd, D.H. & Rubin, E.J. Genes required for mycobacterial growth defined by high density mutagenesis. Mol. Microbiol. 48, 77–84 (2003).
Choi, Y., Sims, G.E., Murphy, S., Miller, J.R. & Chan, A.P. Predicting the functional effect of amino acid substitutions and indels. PLoS One 7, e46688 (2012).
Falush, D. & Bowden, R. Genome-wide association mapping in bacteria? Trends Microbiol. 14, 353–355 (2006).
Shell, S.S. et al. DNA methylation impacts gene expression and ensures hypoxic survival of Mycobacterium tuberculosis. PLoS Pathog. 9, e1003419 (2013).
Brosch, R. et al. A new evolutionary scenario for the Mycobacterium tuberculosis complex. Proc. Natl. Acad. Sci. USA 99, 3684–3689 (2002).
Chavadi, S. et al. Global effects of inactivation of the pyruvate kinase gene in the Mycobacterium tuberculosis complex. J. Bacteriol. 191, 7545–7553 (2009).
Keating, L.A. et al. The pyruvate requirement of some members of the Mycobacterium tuberculosis complex is due to an inactive pyruvate kinase: implications for in vivo growth. Mol. Microbiol. 56, 163–174 (2005).
Ängeby, K., Juréen, P., Kahlmeter, G., Hoffner, S.E. & Schön, T. Challenging a dogma: antimicrobial susceptibility testing breakpoints for Mycobacterium tuberculosis. Bull. World Health Organ. 90, 693–698 (2012).
Torrea, G. et al. Bedaquiline susceptibility testing of Mycobacterium tuberculosis in an automated liquid culture system. J. Antimicrob. Chemother. 70, 2300–2305 (2015).
Kahlmeter, G. The 2014 Garrod Lecture: EUCAST—are we heading towards international agreement? J. Antimicrob. Chemother. 70, 2427–2439 (2015).
Pholwat, S. et al. Integrated microfluidic card with TaqMan probes and high-resolution melt analysis to detect tuberculosis drug resistance mutations across 10 genes. MBio 6, e02273 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Walker, B.J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One 9, e112963 (2014).
Price, M.N., Dehal, P.S. & Arkin, A.P. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS One 5, e9490 (2010).
George, K.M., Yuan, Y., Sherman, D.R. & Barry, C.E. III. The biosynthesis of cyclopropanated mycolic acids in Mycobacterium tuberculosis. Identification and functional analysis of CMAS-2. J. Biol. Chem. 270, 27292–27298 (1995).
Streicher, E.M. et al. Rapid sequencing of the Mycobacterium tuberculosis pncA gene for detection of pyrazinamide susceptibility. J. Clin. Microbiol. 52, 4056–4057 (2014).
Acknowledgements
The authors would like to express their sincere gratitude to the patients in South Africa and China who provided clinical samples, without which this study would not have been possible. The authors also gratefully acknowledge the collaborative efforts of the following individuals whose significant previous contributions allowed for this study: W.R. Bishai, N. Bantubani, L. Alvarado, S.B. Chapman, N.R. Mvelase, E.Y. Duffy, M.G. Fitzgerald, P. Govender, S. Gujja, S. Hamilton, C. Howarth, J.D. Larimer, M.D. Pearson, M.E. Priest, and Q. Zeng. M. bovis ATCC 19210 was generously provided by R. Warren (Stellenbosch University). The authors would also like to thank C. Cuomo and three anonymous reviewers for providing helpful comments on the manuscript. Additionally, the authors would like to acknowledge the Broad Sequencing Platform for assistance with data acquisition.
This project has been funded in part with federal funds from NIAID, NIH, US Department of Health and Human Services, under contract HHSN272200900018C, IeDEA (NIH grant 5U01AI069924-07), and grant U19AI110818 to the Broad Institute. K.A.C. was supported by NHLBI grant T32HL007633. T.A. was supported by a postdoctoral fellowship from the Research Foundation–Flanders. Additional funding sources included grant U19AI51794 and the US Centers for Disease Control and Prevention. The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
A.M.E., A.S.P., B.W.B., C.A.D., and K.A.C. conceived and designed the project. N.P., M.R.O'D., K.P.M., and A.S.P. provided the clinical isolates. K.A.C., V.M., K.M., J.G., and D.V.A. performed the wet-lab experiments. C.A.D., K.A.C., T.A., T.P.S., A.L.M., and A.S. analyzed the data. B.J.W. contributed analytic tools. C.A.D., K.A.C., A.M.E., and A.S.P. wrote the manuscript. J.W., B.W.B., A.M.E., and A.S.P. supervised and coordinated the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Significance of associations between genotypic variants and drug resistance phenotypes excluding strains with known resistance-conferring genotypes on a per-drug basis.
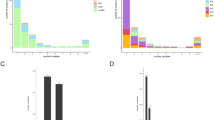
Each circle represents a genotypic feature that is plotted at the intersection of the negative log-transformed P values from the Fisher’s exact and correlated evolution tests for each drug. P values were corrected for multiple comparisons using the Benjamini–Hochberg method. Genotypic variants known to confer resistance are colored according to the drug to which they confer resistance, and genotypes with no known effect on drug resistance are shown in gray. Genotypes scoring well in both tests appear in the upper right quadrants.
Supplementary Figure 2 A single-gene knockout of ald has a shorter time to positivity than wild-type M. tuberculosis when cultured in the presence of d-cycloserine.
Four laboratory strains—wild-type M. tuberculosis (WT), Δald (ald knockout), ald knockout complemented with ald (Δald-comp), and BCG—were cultured in varying concentrations of d-cycloserine. The color legend indicates the concentration of d-cycloserine in μg/ml. Strains were set up in triplicate, and the resulting time to positivity in MGIT was recorded as days since inoculation. The mean time to positivity is plotted with error bars to represent s.e.m. P values were calculated using two-way ANOVA.
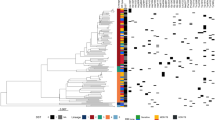
Supplementary Figure 3 Evolution of ald, alr, ddlA, cycA, and pykA across the M. tuberculosis complex (MTBC).
A maximum-likelihood phylogeny was estimated using strains representing the diversity of the MTBC and specifically the vaccine strain BCG. Acquisition of nonsynonymous and loss-of-function mutations in the genes mentioned above was then reconstructed using parsimony and depicted at the appropriate nodes. Clades other than BCG with uniform genotypes were collapsed for visualization. Loss of function of both ald and pykA occurs in conjunction with the loss of region of difference 9 (RD9); pykA loss of function is reverted in both BCG and M. suricattae (indicated above as gain of function, GOF). A cycA loss-of-function mutation also occurs at the base of the M. microti–M. pinnipedii clade. Two mutations were identified in nearly all strains except for the reference H37Rv and close relatives (cycA R93L and ddlA T365A), suggesting that they are phylogenetic mutations and not correlative of resistance to any drugs.
Supplementary Figure 4 Restoration of wild-type ald in BCG inhibits growth in the presence of higher concentrations of d-cycloserine.
As BCG, which is innately resistant to d-cycloserine, contains a frameshift mutation in ald, a wild-type copy of M. tuberculosis ald was used for complementation back into BCG (BCG-ald-comp). Both BCG and BCG-ald-comp were subsequently cultured in varying concentrations of d-cycloserine in MGIT, and the time to positivity was normalized to that for the no-drug control for each strain to calculate the growth inhibition index. P values were calculated using two-way ANOVA.
Supplementary Figure 5 Restoration of wild-type ald in BCG confers longer time to positivity in the presence of higher concentrations of d-cycloserine.
As BCG, which is innately resistant to d-cycloserine, contains a frameshift mutation in ald, a wild-type copy of M. tuberculosis ald was used for complementation back into BCG (BCG-ald-comp). Both BCG and BCG-ald-comp were subsequently cultured in varying concentrations of d-cycloserine in MGIT. Strains were assessed in triplicate, and mean time to positivity in MGIT in days is plotted with the s.e.m. The concentration of d-cycloserine is given in μg/ml. P values were calculated using two-way ANOVA.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1, 2 and 5–12, and Supplementary Note. (PDF 1731 kb)
Supplementary Table 3
Top 20 scoring features from the Fisher's exact and correlated evolution tests for each drug from the analysis of all strains. (XLSX 59 kb)
Supplementary Table 4
Top 20 scoring features from the Fisher's exact and correlated evolution tests for each drug from the analysis of strains without known resistance-onferring mutations. (XLSX 59 kb)
Rights and permissions
About this article
Cite this article
Desjardins, C., Cohen, K., Munsamy, V. et al. Genomic and functional analyses of Mycobacterium tuberculosis strains implicate ald in D-cycloserine resistance. Nat Genet 48, 544–551 (2016). https://doi.org/10.1038/ng.3548
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3548
This article is cited by
-
Heterologous production of the D-cycloserine intermediate O-acetyl-L-serine in a human type II pulmonary cell model
Scientific Reports (2023)
-
Laboratory evolution of Mycobacterium on agar plates for analysis of resistance acquisition and drug sensitivity profiles
Scientific Reports (2021)
-
Phylogenetically informative mutations in genes implicated in antibiotic resistance in Mycobacterium tuberculosis complex
Genome Medicine (2020)
-
Integron gene cassettes harboring novel variants of d-alanine-d-alanine ligase confer high-level resistance to d-cycloserine
Scientific Reports (2020)
-
A biochemically-interpretable machine learning classifier for microbial GWAS
Nature Communications (2020)