Abstract
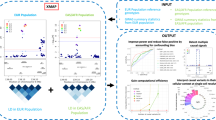
When cellular traits are measured using high-throughput DNA sequencing, quantitative trait loci (QTLs) manifest as fragment count differences between individuals and allelic differences within individuals. We present RASQUAL (Robust Allele-Specific Quantitation and Quality Control), a new statistical approach for association mapping that models genetic effects and accounts for biases in sequencing data using a single, probabilistic framework. RASQUAL substantially improves fine-mapping accuracy and sensitivity relative to existing methods in RNA-seq, DNase-seq and ChIP-seq data. We illustrate how RASQUAL can be used to maximize association detection by generating the first map of chromatin accessibility QTLs (caQTLs) in a European population using ATAC-seq. Despite a modest sample size, we identified 2,707 independent caQTLs (at a false discovery rate of 10%) and demonstrated how RASQUAL and ATAC-seq can provide powerful information for fine-mapping gene-regulatory variants and for linking distal regulatory elements with gene promoters. Our results highlight how combining between-individual and allele-specific genetic signals improves the functional interpretation of noncoding variation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
08 February 2016
In the version of this article initially published, the accession code for the ATAC-seq data was omitted. These data have been deposited in the European Nucleotide Archive under accession ERP011141. The error has been corrected in the HTML and PDF versions of the article.
References
Pickrell, J.K. et al. Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature 464, 768–772 (2010).
Montgomery, S.B. et al. Transcriptome genetics using second generation sequencing in a Caucasian population. Nature 464, 773–777 (2010).
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511 (2013).
Battle, A. et al. Characterizing the genetic basis of transcriptome diversity through RNA-sequencing of 922 individuals. Genome Res. 24, 14–24 (2014).
Degner, J.F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012).
McVicker, G. et al. Identification of genetic variants that affect histone modifications in human cells. Science 342, 747–749 (2013).
Kilpinen, H. et al. Coordinated effects of sequence variation on DNA binding, chromatin structure, and transcription. Science 342, 744–747 (2013).
Kasowski, M. et al. Extensive variation in chromatin states across humans. Science 342, 750–752 (2013).
Ding, Z. et al. Quantitative genetics of CTCF binding reveal local sequence effects and different modes of X-chromosome association. PLoS Genet. 10, e1004798 (2014).
Banovich, N.E. et al. Methylation QTLs are associated with coordinated changes in transcription factor binding, histone modifications, and gene expression levels. PLoS Genet. 10, e1004663 (2014).
Pastinen, T. Genome-wide allele-specific analysis: insights into regulatory variation. Nat. Rev. Genet. 11, 533–538 (2010).
Lefebvre, J.F. et al. Genotype-based test in mapping cis-regulatory variants from allele-specific expression data. PLoS One 7, e38667 (2012).
Degner, J.F. et al. Effect of read-mapping biases on detecting allele-specific expression from RNA-sequencing data. Bioinformatics 25, 3207–3212 (2009).
Pickrell, J.K., Gaffney, D.J., Gilad, Y. & Pritchard, J.K. False positive peaks in ChIP-seq and other sequencing-based functional assays caused by unannotated high copy number regions. Bioinformatics 27, 2144–2146 (2011).
DeVeale, B., van der Kooy, D. & Babak, T. Critical evaluation of imprinted gene expression by RNA-Seq: a new perspective. PLoS Genet. 8, e1002600 (2012).
Waszak, S.M. et al. Identification and removal of low-complexity sites in allele-specific analysis of ChIP-seq data. Bioinformatics 30, 165–171 (2014).
Seoighe, C., Nembaware, V. & Scheffler, K. Maximum likelihood inference of imprinting and allele-specific expression from EST data. Bioinformatics 22, 3032–3039 (2006).
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Dempster, A.P., Laird, N.M. & Rubin, D.B. Maximum likelihood from incomplete data via EM algorithm. J. R. Stat. Soc. B 39, 1–38 (1977).
Weirauch, M.T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Sun, W. A statistical framework for eQTL mapping using RNA-seq data. Biometrics 68, 1–11 (2012).
Shabalin, A.A. Matrix eQTL: ultra fast eQTL analysis via large matrix operations. Bioinformatics 28, 1353–1358 (2012).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Gregg, C., Zhang, J., Butler, J.E., Haig, D. & Dulac, C. Sex-specific parent-of-origin allelic expression in the mouse brain. Science 329, 682–685 (2010).
Gregg, C. et al. High-resolution analysis of parent-of-origin allelic expression in the mouse brain. Science 329, 643–648 (2010).
Heap, G.A. et al. Genome-wide analysis of allelic expression imbalance in human primary cells by high-throughput transcriptome resequencing. Hum. Mol. Genet. 19, 122–134 (2010).
McDaniell, R. et al. Heritable individual-specific and allele-specific chromatin signatures in humans. Science 328, 235–239 (2010).
Ongen, H. et al. Putative cis-regulatory drivers in colorectal cancer. Nature 512, 87–90 (2014).
Kasowski, M. et al. Variation in transcription factor binding among humans. Science 328, 232–235 (2010).
Li, G. et al. Identification of allele-specific alternative mRNA processing via transcriptome sequencing. Nucleic Acids Res. 40, e104 (2012).
GTEx Consortium. The landscape of genomic imprinting across diverse adult human tissues. Genome Res. 25, 927–936 (2015).
Babak, T. et al. Genetic conflict reflected in tissue-specific maps of genomic imprinting in human and mouse. Nat. Genet. 47, 544–549 (2015).
Leighton, P.A., Saam, J.R., Ingram, R.S., Stewart, C.L. & Tilghman, S.M. An enhancer deletion affects both H19 and Igf2 expression. Genes Dev. 9, 2079–2089 (1995).
Banet, G. et al. Characterization of human and mouse H19 regulatory sequences. Mol. Biol. Rep. 27, 157–165 (2000).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Berndt, S.I. et al. Genome-wide association study identifies multiple risk loci for chronic lymphocytic leukemia. Nat. Genet. 45, 868–876 (2013).
Koren, A. et al. Genetic variation in human DNA replication timing. Cell 159, 1015–1026 (2014).
Panousis, N.I., Gutierrez-Arcelus, M., Dermitzakis, E.T. & Lappalainen, T. Allelic mapping bias in RNA-sequencing is not a major confounder in eQTL studies. Genome Biol. 15, 467 (2014).
del Rosario, R.C. et al. Sensitive detection of chromatin-altering polymorphisms reveals autoimmune disease mechanisms. Nat. Methods 12, 458–464 (2015).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 42, D1001–D1006 (2014).
Acknowledgements
We thank O. Stegle, M. Hemberg, G. Trynka and the three anonymous reviewers for their helpful comments. N.K., A.J.K. and D.J.G. were funded by Wellcome Trust grant 098051.
Author information
Authors and Affiliations
Contributions
D.J.G. and N.K. conceived and designed the experiments. N.K. and A.J.K. performed the experiments. N.K. performed statistical analysis and analyzed the data. N.K. and A.J.K. contributed reagents, materials and analysis tools. D.J.G., N.K. and A.J.K. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–30, Supplementary Tables 1–4 and Supplementary Note. (PDF 9239 kb)
Rights and permissions
About this article
Cite this article
Kumasaka, N., Knights, A. & Gaffney, D. Fine-mapping cellular QTLs with RASQUAL and ATAC-seq. Nat Genet 48, 206–213 (2016). https://doi.org/10.1038/ng.3467
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3467
This article is cited by
-
Haplotype-aware modeling of cis-regulatory effects highlights the gaps remaining in eQTL data
Nature Communications (2024)
-
SEESAW: detecting isoform-level allelic imbalance accounting for inferential uncertainty
Genome Biology (2023)
-
Single-cell multiomics of the human retina reveals hierarchical transcription factor collaboration in mediating cell type-specific effects of genetic variants on gene regulation
Genome Biology (2023)
-
Single-cell allele-specific expression analysis reveals dynamic and cell-type-specific regulatory effects
Nature Communications (2023)
-
Molecular quantitative trait loci
Nature Reviews Methods Primers (2023)