Abstract
The ruff is a Palearctic wader with a spectacular lekking behavior where highly ornamented males compete for females1,2,3,4. This bird has one of the most remarkable mating systems in the animal kingdom, comprising three different male morphs (independents, satellites and faeders) that differ in behavior, plumage color and body size. Remarkably, the satellite and faeder morphs are controlled by dominant alleles5,6. Here we have used whole-genome sequencing and resolved the enigma of how such complex phenotypic differences can have a simple genetic basis. The Satellite and Faeder alleles are both associated with a 4.5-Mb inversion that occurred about 3.8 million years ago. We propose an evolutionary scenario where the Satellite chromosome arose by a rare recombination event about 500,000 years ago. The ruff mating system is the result of an evolutionary process in which multiple genetic changes contributing to phenotypic differences between morphs have accumulated within the inverted region.
Similar content being viewed by others
Main
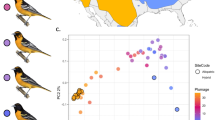
Independents constitute 80–95% of male ruffs and strive to defend territories on leks1,2,3,7. Independent males show a spectacular diversity in the color of their ruff and head tufts (Fig. 1a). Satellites are slightly smaller than independents, usually show white ruff and white tufts (Fig. 1a) and constitute 5–20% of males7,8. Satellites are non-territorial and display submissive behavior, allowing independent males to dominate them at leks (Fig. 1a, middle). Independents clearly recognize satellites as a different kind of male and behave differently with satellites than they do with other independents (see URLs for a link to a video showing the reproductive strategies of the three male morphs). Both independents and satellites may benefit from their interaction by attracting females7. The faeder is a rare third morph (<1% of male ruffs) mimicking females by its smaller size and female-like plumage1,9,10 (Fig. 1a). These disguised males appear on the leks where they attempt to gain access to females that are ready to mate.
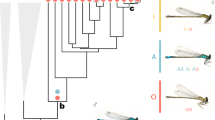
(a) Phenotypic diversity among male ruffs. The middle photograph shows an independent male dominating a satellite. The photographs were provided by Ola Jennersten (left and middle) and F.W. (right). (b) Genome-wide screen for genetic differentiation between independents and satellites using normalized FST values (ZFST) calculated in 15-kb windows. The pattern of differentiation in the associated 4.5-Mb region on scaffold 28 is highlighted below. (c) Neighbor-joining trees for independent and satellite males based on the 4.5-Mb associated region (left) and the rest of the genome (right). (d) The 4.5-Mb inversion disrupts CENPN. (e) Diagnostic test for the inversion: design and genotype results. Primer binding sites are indicated by red arrows, and predicted PCR products are shown below as gray boxes. (f) Conserved synteny between chicken chromosome 11 and ruff scaffold 28 based on an independent male; colored blocks represent individual genes.
A high-quality genome assembly was established using genomic DNA from an independent male kept at the Helsinki zoo in Finland. We estimated the genome size to be 1.23 Gb and generated 139.3 Gb of Illumina HiSeq 2000 sequencing data using fragment libraries with insert sizes ranging from 250 bp to 20 kb (Supplementary Figs. 1, 2, 3 and Supplementary Table 1). The N50 scaffold size was as high as 10.0 Mb (Supplementary Table 2).
We generated ∼8× genome coverage on the basis of 2 × 125-bp paired-end reads from 15 independent and nine satellite males, all from a single location. A screen based on the fixation index (FST) comparing independents and satellites identified a single highly differentiated 4.5-Mb region on scaffold 28 (Fig. 1b). Independents and satellites clustered as two genetically distinct groups in a phylogenetic tree based on this region (Fig. 1c, left). In contrast, there was no significant differentiation between these groups when the tree was constructed on the basis of the rest of the genome (Fig. 1c, right). We hypothesized that the large region of strong differentiation might reflect the presence of an inversion. We used BreakDancer11 to screen for structural changes and identified a 4.5-Mb inversion present in satellites and overlapping perfectly the differentiated region (Fig. 1d). PCR-based sequencing confirmed a proximal breakpoint at 5.8 Mb and a distal breakpoint at 10.3 Mb and identified a 2,108-bp insertion of a repetitive sequence at the distal breakpoint. A diagnostic test (Fig. 1e) showed that all satellites were heterozygous for the inversion and all 112 independents except five were homozygous for the wild-type sequence (Table 1); the latter five were heterozygous for the inversion and most likely reflect phenotype misclassifications in the field. The inversion was also found among adult females and young birds (Table 1). The Independent allele clearly represents the ancestral state, as the inversion disrupts conserved synteny among birds (Fig. 1f).
The single faeder in our material was also heterozygous for the 4.5-Mb inversion (Table 1). We sequenced this individual to 30× coverage, and FST analysis indicated striking genetic differentiation between the faeder and both independents and satellites within the inverted region (Fig. 2a). The differentiation between the faeder and independents was equally strong across the 4.5-Mb region, whereas the pattern of differentiation between the faeder and satellites was a mirror image of the pattern between satellites and independents (Fig. 2a). We phased haplotypes using Beagle12 and constructed haplotype trees separately for region A showing high FST between satellites and independents and for region B showing low FST between satellites and independents (Fig. 2a,c). In region A, the Satellite and Faeder chromosomes were closely related and divergent from Independent chromosomes. In contrast, in region B, the Satellite and Independent chromosomes were more closely related, whereas the Faeder chromosome was divergent (Fig. 2c, bottom). Because we only had access to a single faeder, we genotyped the entire material (>200 birds) using two SNPs diagnostic for the Faeder chromosome. The Faeder chromosome should not be present in independent or satellite males but should occur at a low frequency in adult females and young birds. The results confirmed this prediction, as we identified the Faeder chromosome in only a single adult female (Supplementary Table 3). This female was clearly an outlier with regard to body size (Supplementary Fig. 4), consistent with the observation that females heterozygous for the Faeder allele are smaller than other females6.
(a) Genetic differentiation among independent, satellite and faeder males based on FST analysis. (b) Box plots showing estimated divergence times between the Independent and Satellite chromosomes for regions A and B, calculated using the substitution rates estimated for 48 different bird genomes18. The central rectangle spans the first to third quartiles of the distribution, and the 'whiskers' above and below the box show the maximum and minimum estimates. The line inside the rectangle shows the median. MYA, million years ago. (c) Phylogenetic trees for haplotypes carried by independent, satellite and faeder males for region A (red) and region B (blue) showing different patterns of differentiation. Hap, haplotype. (d) Three deletions (blue) present in satellites and faeders within the 4.5-Mb inversion are all located in the vicinity of HSD17B2 and SDR42E1. Two duplications (magenta) are only present in faeder. (e) The 5.2-kb deletion shows 50% reduced sequence coverage in satellites (top); the deletion encompasses evolutionarily conserved sequences among birds18 (bottom). (f) Proposed evolutionary history of the three alleles underlying the mating system in ruff.
We examined the genetic consequences of the inversion and searched for candidate mutations that might contribute to phenotypic differences. This is challenging because the inverted region contains about 90 genes (Supplementary Fig. 5 and Supplementary Table 4). First, we note that the inversion disrupts the CENPN gene (encoding centromere protein N) (Fig. 1d). The inversion may be recessive lethal, as data from human cells13 and zebrafish14 with mutations in the orthologous gene show that CENPN inactivation has severe deleterious effects. In fact, ruff pedigree data have confirmed that the inversion is recessive lethal15. Birds heterozygous for the Satellite allele must have about 5% higher fitness to maintain an allele frequency of about 5% in compensating for the lethality of the homozygote. Second, we identified a large number of missense mutations present on Satellite and/or Faeder chromosomes (Supplementary Table 5). Third, BreakDancer11 and depth-of-coverage analysis identified three deletions ranging in size from 3.3 to 17.6 kb (Fig. 2d) present in the heterozygous condition in all satellites and in the faeder but not in independent males (Supplementary Table 3). Two of these (5.2 kb and 17.6 kb in length) delete evolutionarily conserved sequences (Fig. 2e and Supplementary Fig. 6), and all three cluster in the vicinity of HSD17B2 (hydroxysteroid (17-β) dehydrogenase 2) and SDR42E1 (short-chain dehydrogenase/reductase family 42E, member 1) (Fig. 2d). HSD17B2 and SDR42E1 both have important roles in the metabolism of sex hormones: HSD17B2 catalyzes conversion of the 17β-hydroxy forms of estrogen and androgens (including testosterone and dihydrotestosterone) into their less active 17-keto forms16. We postulate that one or more of these deletions constitute cis-acting regulatory mutations that alter the expression pattern of HSD17B2 and/or SDR42E1 and contribute to phenotypic differences among male morphs. We identified two deletions and two duplications that were unique to the faeder and may thus contribute to the faeder phenotype (Supplementary Fig. 7).
There is a striking diversity in plumage color among male ruffs (Fig. 1a). One of the most obvious candidate genes for variation in pigmentation, MC1R (encoding melanocortin 1 receptor), is located within the inverted region at position 10.2 Mb (Supplementary Fig. 5). Whole-genome sequencing showed that satellites are heterozygous for four derived MC1R missense mutations (encoding p.Val105Leu, p.Arg149His, p.His207Arg and p.Arg303Trp) at residues that are conserved among birds and mammals (Fig. 3). We performed Sanger sequencing of the single MC1R exon in all satellites, the faeder and a subset of the independents. This analysis confirmed complete association between the Satellite allele and these four missense mutations, whereas the faeder was heterozygous for the variant encoding p.His207Arg and three other missense mutations (Supplementary Table 6). The p.His207Arg substitution most likely has functional consequences because the same variant is associated with light color in the red-footed booby17. We propose that the MC1R allele on Satellite chromosomes, possibly together with altered metabolism of sex hormones, underlies the white color of ornamental feathers in satellites. To be causal, this allele must have a dominant effect, as satellites are always heterozygous. This implies a dominant-negative effect or, more likely, a combination of regulatory and coding changes leading to overexpression of a variant form of MC1R specifically in ornamental feathers. The latter mechanism would explain why satellite males, despite their spectacular light color during the breeding season (Fig. 1a), are almost indistinguishable from independents outside the breeding season and why females carrying the Satellite allele are not markedly lighter in color than other females.
Missense variants detected in satellites are represented by red circles, and a selected set of variants associated with plumage variation in other birds29 is represented by yellow circles. Lower left, the amino acids associated with the Independent allele are conserved among birds and mammals.
We estimated the time since divergence of the Satellite and Independent alleles on the basis of region A, showing high FST between the two types of males, to 3.87 ± 0.15 million years ago (Fig. 2a,b), using the nucleotide divergence (1.4%) and estimated mutation rates for birds18. A similar estimate of 4.09 ± 0.16 million years ago was obtained for divergence of the Faeder and Independent alleles on the basis of sequence divergence for the entire 4.5-Mb region. There can be no recombination between the Satellite and Faeder chromosomes, as the Faeder/Satellite genotype is not viable. Furthermore, an inversion is expected to cause suppression of recombination within the inverted region between the wild-type and mutant alleles. Comparison of the faeder and independents across the inversion is consistent with this lack of recombination, as FST values are equally strong across the 4.5-Mb region (Fig. 2a). Remarkably, the strong differentiation between satellites and independents is disrupted in two regions (1.5 and 0.7 Mb in size) that show FST values that are much lower than those for other segments of the inversion but still markedly higher than the background level (Fig. 2a). We postulate that the Satellite chromosome arose by one or two rare recombination events between an Independent and a Faeder-like chromosome (Fig. 2f). This would explain why the pattern of genetic differentiation between the Faeder and Satellite chromosomes is a mirror image of the differences between the Independent and Satellite chromosomes (Fig. 2a). A similar rare recombination event between an inversion and a wild-type chromosome created a third allele at the Rose-comb locus in chicken19. Our model predicts that the pairwise genetic distances between the Satellite and Independent chromosomes for region B should all be equal, as they reflect divergence since the recombination event happened. Our data are consistent with this hypothesis (Supplementary Table 7). Similarly, the pairwise genetic distances between the Satellite and Faeder chromosomes in region A are also equal (Supplementary Table 7). We estimated that this recombination event occurred 520,000 ± 20,000 years ago, on the basis of sequence divergence (0.2%) between the Satellite and Independent chromosomes in region B (Fig. 2b). We constructed 5-kb and 10-kb mate-pair libraries from one satellite male, but analysis of these data did not identify any additional inversions in the vicinity of the 4.5-Mb inversion.
The genetic basis for the satellite and faeder morphs constitutes a combination of genetic changes that have accumulated within the inverted region over a period of about 3.8 million years. This resembles the situation in white-throated sparrows in which at least two pericentric inversions involving about ∼100 Mb are associated with altered plumage (brown-and-tan stripes versus white-and-black stripes on the crown) and altered territorial and parental behavior20,21. A recent study suggested that sequence differences in the promoter of ESR1 (encoding estrogen receptor α) are causally related to behavioral differences between morphs22. When an inversion is associated with a complex phenotype, it is challenging to pinpoint causal mutations because many sequence polymorphisms within the inverted region show an equally strong association to the phenotype. An additional complication in ruff is that recessive variants located within the inverted region will never be exposed to purifying selection because homozygosity for the inversion is lethal. Only sequence variants on the Satellite and Faeder chromosomes that show some degree of dominance can contribute to phenotypic differences among the morphs. We propose that the inversion itself caused the first phenotypic effects that constituted the starting point for an evolutionary process eventually resulting in the current mating system in ruff. An inversion may cause phenotypic effects as a result of changes affecting the coding sequence or the regulation of genes, primarily in the vicinity of the breakpoints, as illustrated by the Rose-comb inversion in chicken where translocation of MNR2 leads to ectopic expression of the MNR2 transcription factor and altered comb development19. Similarly, translocation of CYB5B, located at the breakpoint at 10.3 Mb in ruff (Fig. 1d), may cause altered expression of this gene that has a role in the biosynthesis of glucocorticoids and sex steroids23. Furthermore, it appears highly plausible that one or more of the three deletions in the near vicinity of HSD17B2 and SDR42E1 lead to altered metabolism of testosterone and other steroids, which may affect both behavior and plumage. In fact, independents have higher circulating levels of testosterone, whereas satellites and faeders have higher concentrations of androstenedione15. A possible explanation for this difference is that upregulation and/or ectopic expression of HSD17B2 lead to conversion of testosterone into androstenedione in individuals heterozygous for the Satellite or Faeder allele, a model consistent with the dominant inheritance of these alleles. This is a testable hypothesis because it predicts allelic imbalance in HSD17B2 expression in satellites and faeders.
Our study has demonstrated how an inversion followed by subsequent accumulation of several adaptive changes within the inverted region led to the evolution of a spectacular mating system comprising three alleles at a single locus maintained by balancing selection. The presence of an inversion that allowed the evolution of a non-recombining 'supergene' (ref. 24) was critical for this process. Other examples where inversions are associated with complex phenotypes include mimicry in butterflies25,26,27 and colony organization in fire ants28.
Methods
Sample collection and DNA extraction.
Blood was collected form the wing vein of a captive male ruff (further referred to as the reference individual) kept at the Helsinki zoo, Finland, and mixed with EDTA as anticoagulant. DNA was immediately isolated using a standard salt precipitation method. The quantity and quality of the sample were evaluated using the Qubit dsDNA BR assay (Life Technologies) as well as by pulsed-field gel electrophoresis (CHEF Mapper XA, Bio-Rad).
The samples for whole-genome resequencing and further genetic analysis were collected from a ruff population that was studied during the breeding seasons of 1990–2002 on the island of Gotland in the Baltic Sea (57° 10′ N, 18° 20′ E)30,31,32. In each year, males and females were caught on leks using cannon nets. In addition, females and newly hatched young were caught on nests. Individuals were ringed, and morphological measurements including tarsus length (in mm) and wing length (in mm; maximum chord) were collected (all by F.W.). A blood sample was drawn from the wing or brachial vein of each bird and later used for DNA extraction. The birds were released after handling. Birds were caught and handled according to ethical permits and permissions guiding Swedish research and animal welfare (Stockholms Södra Djurförsöksetiska Nämnd S54-99).
Male strategies were determined from plumage pattern and from behavioral observations at leks for ringed independents and satellites7. Most males (>90%) in nuptial plumage were scored as independents or satellites from their plumage alone. Whether female-colored birds were faeder males could only be preliminarily determined from morphological measurements and later verified through sexing using a DNA test33.
Genome assembly.
We generated 139.32 Gb of high-quality next-generation sequencing data with fragment lengths ranging from 250 bp to 20 kb for the reference individual using Illumina HiSeq 2000 sequencing (Supplementary Table 1). We estimated the ruff genome size using k-mer analysis to be about 1.23 Gb (Supplementary Fig. 1), suggesting genome sequence coverage of 113.7-fold. Using SOAPdenovo (version v2.04)34, we obtained an assembly spanning 1.25 Gb, with contig N50 and scaffold N50 sizes of 106.46 kb and 10.00 Mb, respectively (Supplementary Table 2); 96.3% of the assembly was non-gap sequences. The distribution of sequencing depth, calculated on the basis of reads from all sequencing libraries, and the distribution of GC content are presented in Supplementary Figures 2 and 3, respectively.
Annotation of the region associated with an inversion.
A preliminary annotation for the inverted region was generated with the Maker package (version 2.31-8)35. We first composed a set of high-confidence reference sequences from UniProt by selecting bird proteins that were classified as full length and supported by either proteomics or transcriptomics (74,138 sequences). As additional input, we collected available 454 sequencing data from the SRA available under accession SRA049313. Reads were assembled with the Trinity package (release 2014-07-07)36 into 16,746 transcripts. Finally, to improve the accuracy of the annotation process, we modeled new repeat sequences from a preliminary genome assembly and used this library in combination with a curated repeat library for vertebrates included in the RepeatMasker package. From these data, we generated two complementary gene builds: one based directly on the aligned sequences to most accurately reflect the evidence data ('evidence build') and a second set of gene models seeded from ab initio (de novo) gene predictions generated by the chicken reference profile model, included with the Augustus package (version 2.7)37. Both builds were compared and reconciled for the target region using the WebApollo curation platform38. Functional annotation of candidate transcript models was performed through similarity searches39 against the UniProt/SWISS-PROT reference protein set (downloaded May 2014) in combination with the prediction of functional motifs and domains via the InterProScan package (release 5.7-48.0)40.
Whole-genome resequencing and SNP calling.
Sequencing libraries (average fragment size of about 500 bp) were constructed for 15 independents, nine satellites and one faeder, and 2 × 125-bp paired-end reads were generated using Illumina HiSeq 2000 sequencers. For the independents and satellites, we generated ∼8-fold coverage based on high-quality reads, after strict filtering of low-quality and adaptor-contaminated reads, and ∼30-fold coverage was generated for the single faeder male. The genomic reads were mapped against the ruff genome assembly using Burrows-Wheeler Aligner (BWA)41 (version 0.6.2) with default parameters. PCR duplicates were filtered from the alignments using Picard. Further, we performed base quality recalibration and indel realignment using the Genome Analysis Toolkit (GATK)42 and performed SNP discovery and genotyping across the 25 samples according to GATK best-practices recommendations43,44. Low-quality SNP calls were filtered out by an in-house filtering pipeline that excluded a SNP if it did not satisfy the threshold of a combination of various quality parameters (for example, SNP quality, base quality, mapping quality, haplotype score, Fisher strand bias, minimum read depth and maximum read depth). Thresholds were chosen on the basis of the distribution of each of these parameters from the raw variant calls.
Genome-wide screen for genetic differentiation among male morphs.
We divided the genome into non-overlapping 15-kb windows and estimated the genetic divergence (FST) between independents and satellites using VCFtools (v.0.1.11)45. The 4.5-Mb region in scaffold 28 that showed strong genetic differentiation between independents and satellites was further subdivided into smaller windows (5 kb in length) to refine the pattern of differentiation. A similar analysis was carried out when comparing data from the single faeder male and the other morphs.
Phylogenetic analysis.
We used PLINK46 to calculate pairwise genetic distances between individuals. These distances were used to generate neighbor-joining trees with PHYLIP47. Phased haplotypes for the 4.5-Mb inversion region were generated using Beagle12, and these haplotypes were used to estimate Jukes-Cantor corrected nucleotide distances among the Independent, Satellite and Faeder alleles with PHYLIP47. The net frequency of nucleotide substitutions (dA) was calculated according to the method of Nei48, and the time since divergence (t) of alleles was calculated as t = dA/2λ, where λ is the genomic substitution rate. As a substitution rate in ruff is not yet available, we used the substitution rates estimated for each of 48 other bird genomes18 to calculate a confidence interval for t, and the data are presented as a box plot.
Structural variants.
We used the paired-end sequence data for detection of structural variants with BreakDancer11. Information on read pairs that mapped with unexpected separation distances or orientation was used to predict inversions, insertions, duplications and deletions. Two-sided Fisher's exact test was used to identify structural variants showing significant frequency differences between morphs. Sequence alignments around the detected structural variants were also manually inspected using the Integrative Genomics Viewer (IGV)49, to exclude false positives. Normalized read coverage was compared between morphs to check the consistency of the deletions predicted by BreakDancer11. We also generated 5-kb and 10-kb mate-pair data from a single satellite individual to confirm the structural variants detected using paired-end data.
Functional annotation of genetic variants.
We used SNPeff (v.3.4)50 to annotate the genetic variants and categorized the variants into coding (synonymous and nonsynonymous), upstream/downstream and intronic/intergenic classes.
Diagnostic PCR tests, Sanger sequencing and SNP genotyping.
PCR was used to amplify the regions around the identified inversion and deletion breakpoints and the MC1R coding sequence; all primer sequences are given in Supplementary Table 8. Amplified fragments were either analyzed by agarose gel electrophoresis or subjected to Sanger sequencing using standard methods. DNA sequences were analyzed using CodonCode Aligner 5.1.4 software. Four TaqMan SNP genotyping assays (Life Technologies) diagnostic for the Satellite or Faeder haplotype were designed (Supplementary Table 8). Standard TaqMan allele discrimination assays were performed using an Applied Biosystems 7900 HT real-time PCR instrument.
URLs.
NCBI Sequence Read Archive (SRA), http://www.ncbi.nlm.nih.gov/sra/; RepeatMasker, http://repeatmasker.org/; Picard, http://broadinstitute.github.io/picard/; video of ruff reproductive strategies, http://www.scilifelab.se/research/scientific-highlights/ruff.
Accession codes.
The Illumina reads have been submitted to the Sequence Read Archive (SRA) under accession SRA266458. The assembly and annotation are available under accession PRJNA281024. DNA sequences reported in this manuscript have been submitted to GenBank under accessions KT202232–KT202235 and KT428875.
References
Hogan-Warburg, A.J. Social behaviour of the Ruff Philomachus pugnax (L.). Ardea 13, 109–229 (1966).
Van Rhijn, J.G. The Ruff (Poyser, 1991).
Höglund, J. & Alatalo, R.V. Leks (Princeton Univ. Press, 1995).
Lank, D.B. & Dale, J. Visual signals for individual identification: the silent “song” of ruffs. Auk 118, 759–765 (2001).
Lank, D.B., Smith, C.M., Hanotte, O., Burke, T. & Cooke, F. Genetic polymorphism for alternative mating behaviour in lekking male ruff Philomachus pugnax. Nature 378, 59–62 (1995).
Lank, D.B., Farrell, L.L., Burke, T., Piersma, T. & McRae, S.B. A dominant allele controls development into female mimic male and diminutive female ruffs. Biol. Lett. 9, 20130653 (2013).
Widemo, F. Alternative reproductive strategies in the ruff: a mixed ESS? Anim. Behav. 56, 329–336 (1998).
Höglund, J. & Lundberg, A. Plumage color correlates with body size in the ruff (Philomachus pugnax). Auk 13, 306–308 (1989).
Van Rhijn, J.G. Behavioural dimorphism in male ruffs Philomachus pugnax (L.). Behaviour 47, 153–227 (1973).
Jukema, J. & Piersma, T. Permanent female mimics in a lekking shorebird. Biol. Lett. 2, 161–164 (2006).
Chen, K. et al. BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat. Methods 6, 677–681 (2009).
Browning, S.R. & Browning, B.L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Foltz, D.R. et al. The human CENP-A centromeric nucleosome-associated complex. Nat. Cell Biol. 8, 458–469 (2006).
Amsterdam, A. et al. Identification of 315 genes essential for early zebrafish development. Proc. Natl. Acad. Sci. USA 101, 12792–12797 (2004).
Küpper, C. et al. A supergene determines highly divergent male reproductive morphs in the ruff. Nat. Genet. 10.1038/ng.3443 (16 November 2015).
Saloniemi, T., Jokela, H., Strauss, L., Pakarinen, P. & Poutanen, M. The diversity of sex steroid action: novel functions of hydroxysteroid (17β) dehydrogenases as revealed by genetically modified mouse models. J. Endocrinol. 212, 27–40 (2012).
Baião, P.C., Schreiber, E. & Parker, P.G. The genetic basis of the plumage polymorphism in red-footed boobies (Sula sula): a melanocortin-1 receptor (MC1R) analysis. J. Hered. 98, 287–292 (2007).
Zhang, G. et al. Comparative genomics reveals insights into avian genome evolution and adaptation. Science 346, 1311–1320 (2014).
Imsland, F. et al. The Rose-comb mutation in chickens constitutes a structural rearrangement causing both altered comb morphology and defective sperm motility. PLoS Genet. 8, e1002775 (2012).
Thomas, J.W. et al. The chromosomal polymorphism linked to variation in social behavior in the white-throated sparrow (Zonotrichia albicollis) is a complex rearrangement and suppressor of recombination. Genetics 179, 1455–1468 (2008).
Horton, B.M., Moore, I.T. & Maney, D.L. New insights into the hormonal and behavioural correlates of polymorphism in white-throated sparrows, Zonotrichia albicollis. Anim. Behav. 93, 207–219 (2014).
Horton, B.M. et al. Estrogen receptor α polymorphism in a species with alternative behavioral phenotypes. Proc. Natl. Acad. Sci. USA 111, 1443–1448 (2014).
Soucy, P. & Luu-The, V. Assessment of the ability of type 2 cytochrome B5 to modulate 17,20-lyase activity of human P450c17. J. Steroid Biochem. Mol. Biol. 80, 71–75 (2002).
Schwander, T., Libbrecht, R. & Keller, L. Supergenes and complex phenotypes. Curr. Biol. 24, R288–R294 (2014).
Joron, M. et al. Chromosomal rearrangements maintain a polymorphic supergene controlling butterfly mimicry. Nature 477, 203–206 (2011).
Kunte, K. et al. doublesex is a mimicry supergene. Nature 507, 229–232 (2014).
Nishikawa, H. et al. A genetic mechanism for female-limited Batesian mimicry in Papilio butterfly. Nat. Genet. 47, 405–409 (2015).
Wang, J. et al. A Y-like social chromosome causes alternative colony organization in fire ants. Nature 493, 664–668 (2013).
Mundy, N.I. A window on the genetics of evolution: MC1R and plumage colouration in birds. Proc. Biol. Sci. 272, 1633–1640 (2005).
Höglund, J., Montgomerie, R. & Widemo, F. Costs and consequences of variation in the size of ruff leks. Behav. Ecol. Sociobiol. 13, 31–39 (1993).
Widemo, F. The social implications of traditional use of lek sites in the ruff (Philomachus pugnax). Behav. Ecol. 8, 211–217 (1997).
Thuman, K. Female Reproductive Strategies in the Ruff. PhD thesis, Uppsala Univ. (2003).
Fridolfsson, A.-K. & Ellegren, H. A simple and universal method for molecular sexing of non-ratite birds. J. Avian Biol. 30, 116–121 (1999).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience 1, 18 (2012).
Cantarel, B.L. et al. MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 18, 188–196 (2008).
Grabherr, M.G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Stanke, M., Steinkamp, R., Waack, S. & Morgenstern, B. AUGUSTUS: a web server for gene finding in eukaryotes. Nucleic Acids Res. 32, W309–W312 (2004).
Lee, E. et al. Web Apollo: a web-based genomic annotation editing platform. Genome Biol. 14, R93 (2013).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Van der Auwera, G.A. et al. From FastQ data to high-confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinformatics 11, 11.10.1–11.10.33 (2002).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Felsenstein, J. PHYLIP—Phylogeny Inference Package (Version 3.2). Cladistics 5, 164–166 (1989).
Nei, M. Molecular Evolutionary Genetics (Columbia Univ. Press, 1987).
Robinson, J.T. et al. Integrative Genomics Viewer. Nat. Biotechnol. 29, 24–26 (2011).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Acknowledgements
We thank K. Pynnönen-Oudman and S. Sainmaa at the Helsinki zoo for kindly providing a fresh blood sample for the genome assembly. Genome sequencing was funded by the Knut and Alice Wallenberg foundation (to L.A.) and by grants of the National Basic Research Program of China (973 Program; 2013CB835204), the Shenzhen municipal government of China (DRC-SZ (2014) 843), the Key Laboratory of Genomics, the Chinese Ministry of Agriculture, the Platform of Whole Genome–Based Molecular Breeding and the ShenZhen Engineering Laboratory for Genomics-Assisted Animal Breeding. The Swedish Research Council funded the field work (grants 1989-2546, 1992-2685 and 2013-5418 to J.H. and grant 2001-6005 to F.W.). D.S.T. benefited from an Erasmus-Mundus fellowship associated with the European Graduate School of Animal Breeding and Genetics. The SNP&SEQ Technology Platform, supported by Uppsala University and Hospital, the Science for Life Laboratory and the Swedish Research Council (80576801 and 70374401), contributed to genome sequencing. Computer resources were provided by UPPMAX, Uppsala University.
Author information
Authors and Affiliations
Contributions
L.A., J.H. and F.W. conceived the study. F.W. and J.H. collected all material and phenotype data. G.F., M.P.H., C.S., H.Z., W.C., X. Lia, L.H., J.W., E.L., Q.W., S.M.-Y.L., X.X. and X. Liu contributed to genome assembly and annotation. S.L., G.F. and L.A. were responsible for bioinformatics analysis of genome resequencing data. U. Gunnarsson, D.S.T., S.K. and U. Gustafson performed experimental work. L.A. and S.L. wrote the manuscript with input from other authors. All authors approved the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Estimation of ruff genome size using k-mer analysis.
The x axis shows depth (×), and the y axis shows proportion, which represents the frequency at that depth divided by the total frequency of all depth. We estimated the genome size on the basis of the total length of used reads divided by sequencing depth as (N × (L – K + 1) – B)/D=G, where N is the total count of used reads, L is the length of used reads, K is k-mer length (K = 17), B is the total count of low-frequency (frequency ≤1) k-mers that are probably caused by sequencing errors, G is the genome size and D is the k-mer depth, which is estimated from the k-mer distribution. Generally, the k-mer distribution should approximate a Poisson distribution.
Supplementary Figure 2 The distribution of sequencing depth based on the mapping results for reads from short-insert libraries (<2 kb).
The x axis represents average depth. We used 10-kb non-overlapping sliding windows to calculate the average depth among windows.
Supplementary Figure 3 GC content distribution for the ruff genome and three other representative bird genomes.
The x axis shows GC content, and the y axis shows the percentage of windows with a certain GC content. We calculated the GC content in 500-bp sliding windows along the genome (with 250-bp overlap).
Supplementary Figure 4 Wing length plotted against tarsus length for I/I (blue), I/S (red) and I/F (black cross) individuals.
All females are in the lower left cluster, and all males in the upper right cluster.
Supplementary Figure 5 Overview of gene content in the region on scaffold 28 encompassing the inversion present on the Satellite and Faeder chromosomes.
Genes marked with an asterisk may represent fragmented annotations due to lack of ruff transcriptome data or the presence of multiple copies.
Supplementary Figure 6 Evidence for the presence of deletions downstream of HSD17B2 on the Satellite and Faeder chromosomes.
The 17.6-kb (a) and 3.3-kb (b) deletions show 50% reduced sequence coverage in satellites, and the 17.6-kb deletion encompasses evolutionarily conserved sequences among birds18 (bottom). Sequence conservation for the 3.3-kb region is not available because the orthologous region is missing in the chicken Galgal3 assembly that was used to deduce sequence conservation.
Supplementary Figure 7 Structural changes unique to the Faeder chromosome on scaffold 28.
(a) 6.1-kb deletion at 7,690,931–7,697,072 bp, upstream of HNF4B. (b) 10.2-kb deletion at 9,343,057–9,353,258 bp, upstream of ZFPM1. (c) 15.0-kb duplication and copy number expansion at 6,126,000–6,141,000 bp, overlapping CMIP. (d) 1.2-kb deletion of noncoding sequence followed by 16.4-kb duplication showing copy number expansion. This duplicated region include in total 11 exons of PLCG2 at 6,209,770–6,227,434 bp. The structural changes expand the ORF of PLCG2 from 3,702 to 4,620 bp. The breakpoints for the 16.4-kb duplication have been verified by PCR analysis and Sanger sequencing. All four structural changes affect sequences that are conserved among birds18 (right).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–3 and 6–8. (PDF 1627 kb)
Supplementary Table 4
Compilation of the genes within the 4.5-Mb inverted region in ruff scaffold 28. (XLSX 57 kb)
Supplementary Table 5
List of missense mutations associated with the Satellite (a) and Faeder (b) alleles (XLSX 52 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the articles Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Lamichhaney, S., Fan, G., Widemo, F. et al. Structural genomic changes underlie alternative reproductive strategies in the ruff (Philomachus pugnax). Nat Genet 48, 84–88 (2016). https://doi.org/10.1038/ng.3430
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3430
This article is cited by
-
The roles of sexual selection and sexual conflict in shaping patterns of genome and transcriptome variation
Nature Ecology & Evolution (2023)
-
Sex-specific morphs: the genetics and evolution of intra-sexual variation
Nature Reviews Genetics (2023)
-
The significant role of post-pairing male behavior on the evolution of male preferences and female traits
Communications Biology (2022)
-
Exploring potentialities of avian genomic research in Nepalese Himalayas
Avian Research (2021)
-
Research progress in the population differentiation and geographical adaptation of cotton
Journal of Cotton Research (2021)