Abstract
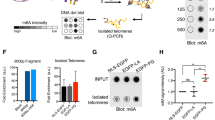
Mutations in the PARN gene (encoding poly(A)-specific ribonuclease) cause telomere diseases including familial idiopathic pulmonary fibrosis (IPF) and dyskeratosis congenita1,2, but how PARN deficiency impairs telomere maintenance is unclear. Here, using somatic cells and induced pluripotent stem cells (iPSCs) from patients with dyskeratosis congenita with PARN mutations, we show that PARN is required for the 3′-end maturation of the telomerase RNA component (TERC). Patient-derived cells as well as immortalized cells in which PARN is disrupted show decreased levels of TERC. Deep sequencing of TERC RNA 3′ termini shows that PARN is required for removal of post-transcriptionally acquired oligo(A) tails that target nuclear RNAs for degradation. Diminished TERC levels and the increased proportion of oligo(A) forms of TERC are normalized by restoring PARN, which is limiting for TERC maturation in cells. Our results demonstrate a new role for PARN in the biogenesis of TERC and provide a mechanism linking PARN mutations to telomere diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Stuart, B.D. et al. Exome sequencing links mutations in PARN and RTEL1 with familial pulmonary fibrosis and telomere shortening. Nat. Genet. 47, 512–517 (2015).
Tummala, H. et al. Poly(A)-specific ribonuclease deficiency impacts telomere biology and causes dyskeratosis congenita. J. Clin. Invest. 125, 2151–2160 (2015).
Dehlin, E., Wormington, M., Korner, C.G. & Wahle, E. Cap-dependent deadenylation of mRNA. EMBO J. 19, 1079–1086 (2000).
Körner, C.G. & Wahle, E. Poly(A) tail shortening by a mammalian poly(A)-specific 3′-exoribonuclease. J. Biol. Chem. 272, 10448–10456 (1997).
Körner, C.G. et al. The deadenylating nuclease (DAN) is involved in poly(A) tail removal during the meiotic maturation of Xenopus oocytes. EMBO J. 17, 5427–5437 (1998).
Virtanen, A., Henriksson, N., Nilsson, P. & Nissbeck, M. Poly(A)-specific ribonuclease (PARN): an allosterically regulated, processive and mRNA cap–interacting deadenylase. Crit. Rev. Biochem. Mol. Biol. 48, 192–209 (2013).
Yoda, M. et al. Poly(A)-specific ribonuclease mediates 3′-end trimming of Argonaute2-cleaved precursor microRNAs. Cell Rep. 5, 715–726 (2013).
Berndt, H. et al. Maturation of mammalian H/ACA box snoRNAs: PAPD5-dependent adenylation and PARN-dependent trimming. RNA 18, 958–972 (2012).
Mason, P.J. & Bessler, M. mRNA deadenylation and telomere disease. J. Clin. Invest. 125, 1796–1798 (2015).
Egan, E.D. & Collins, K. An enhanced H/ACA RNP assembly mechanism for human telomerase RNA. Mol. Cell. Biol. 32, 2428–2439 (2012).
Feng, J. et al. The RNA component of human telomerase. Science 269, 1236–1241 (1995).
Venteicher, A.S. et al. A human telomerase holoenzyme protein required for Cajal body localization and telomere synthesis. Science 323, 644–648 (2009).
Greider, C.W. Telomerase RNA levels limit the telomere length equilibrium. Cold Spring Harb. Symp. Quant. Biol. 71, 225–229 (2006).
Cristofari, G. & Lingner, J. Telomere length homeostasis requires that telomerase levels are limiting. EMBO J. 25, 565–574 (2006).
Cao, Y., Bryan, T.M. & Reddel, R.R. Increased copy number of the TERT and TERC telomerase subunit genes in cancer cells. Cancer Sci. 99, 1092–1099 (2008).
Soder, A.I. et al. Amplification, increased dosage and in situ expression of the telomerase RNA gene in human cancer. Oncogene 14, 1013–1021 (1997).
Heiss, N.S. et al. X-linked dyskeratosis congenita is caused by mutations in a highly conserved gene with putative nucleolar functions. Nat. Genet. 19, 32–38 (1998).
Mitchell, J.R., Wood, E. & Collins, K. A telomerase component is defective in the human disease dyskeratosis congenita. Nature 402, 551–555 (1999).
Vulliamy, T. et al. Mutations in the telomerase component NHP2 cause the premature ageing syndrome dyskeratosis congenita. Proc. Natl. Acad. Sci. USA 105, 8073–8078 (2008).
Vulliamy, T. et al. The RNA component of telomerase is mutated in autosomal dominant dyskeratosis congenita. Nature 413, 432–435 (2001).
Walne, A.J. et al. Genetic heterogeneity in autosomal recessive dyskeratosis congenita with one subtype due to mutations in the telomerase-associated protein NOP10. Hum. Mol. Genet. 16, 1619–1629 (2007).
Mitchell, J.R., Cheng, J. & Collins, K. A box H/ACA small nucleolar RNA–like domain at the human telomerase RNA 3′ end. Mol. Cell. Biol. 19, 567–576 (1999).
Houseley, J. & Tollervey, D. The many pathways of RNA degradation. Cell 136, 763–776 (2009).
Schmidt, K. & Butler, J.S. Nuclear RNA surveillance: role of TRAMP in controlling exosome specificity. Wiley Interdiscip. Rev. RNA 4, 217–231 (2013).
Kiss, T., Fayet-Lebaron, E. & Jady, B.E. Box H/ACA small ribonucleoproteins. Mol. Cell 37, 597–606 (2010).
Rammelt, C., Bilen, B., Zavolan, M. & Keller, W. PAPD5, a noncanonical poly(A) polymerase with an unusual RNA-binding motif. RNA 17, 1737–1746 (2011).
LaCava, J. et al. RNA degradation by the exosome is promoted by a nuclear polyadenylation complex. Cell 121, 713–724 (2005).
Fu, D. & Collins, K. Distinct biogenesis pathways for human telomerase RNA and H/ACA small nucleolar RNAs. Mol. Cell 11, 1361–1372 (2003).
Zaug, A.J., Linger, J. & Cech, T.R. Method for determining RNA 3′ ends and application to human telomerase RNA. Nucleic Acids Res. 24, 532–533 (1996).
Goldfarb, K.C. & Cech, T.R. 3′ terminal diversity of MRP RNA and other human noncoding RNAs revealed by deep sequencing. BMC Mol. Biol. 14, 23 (2013).
Jongmans, M.C. et al. Revertant somatic mosaicism by mitotic recombination in dyskeratosis congenita. Am. J. Hum. Genet. 90, 426–433 (2012).
Kirwan, M. et al. Exogenous TERC alone can enhance proliferative potential, telomerase activity and telomere length in lymphocytes from dyskeratosis congenita patients. Br. J. Haematol. 144, 771–781 (2009).
Westin, E.R. et al. Telomere restoration and extension of proliferative lifespan in dyskeratosis congenita fibroblasts. Aging Cell 6, 383–394 (2007).
Wong, J.M. & Collins, K. Telomerase RNA level limits telomere maintenance in X-linked dyskeratosis congenita. Genes Dev. 20, 2848–2858 (2006).
Hao, L.Y. et al. Short telomeres, even in the presence of telomerase, limit tissue renewal capacity. Cell 123, 1121–1131 (2005).
Alter, B.P., Giri, N., Savage, S.A. & Rosenberg, P.S. Cancer in dyskeratosis congenita. Blood 113, 6549–6557 (2009).
Armanios, M.Y. et al. Telomerase mutations in families with idiopathic pulmonary fibrosis. N. Engl. J. Med. 356, 1317–1326 (2007).
Calado, R.T. et al. Constitutional telomerase mutations are genetic risk factors for cirrhosis. Hepatology 53, 1600–1607 (2011).
Kirwan, M. et al. Defining the pathogenic role of telomerase mutations in myelodysplastic syndrome and acute myeloid leukemia. Hum. Mutat. 30, 1567–1573 (2009).
Tsakiri, K.D. et al. Adult-onset pulmonary fibrosis caused by mutations in telomerase. Proc. Natl. Acad. Sci. USA 104, 7552–7557 (2007).
Yamaguchi, H. et al. Mutations of the human telomerase RNA gene (TERC) in aplastic anemia and myelodysplastic syndrome. Blood 102, 916–918 (2003).
Trapp, S. et al. A virus-encoded telomerase RNA promotes malignant T cell lymphomagenesis. J. Exp. Med. 203, 1307–1317 (2006).
Codd, V. et al. Common variants near TERC are associated with mean telomere length. Nat. Genet. 42, 197–199 (2010).
Soerensen, M. et al. Genetic variation in TERT and TERC and human leukocyte telomere length and longevity: a cross-sectional and longitudinal analysis. Aging Cell 11, 223–227 (2012).
Lee, J.E. et al. The PARN deadenylase targets a discrete set of mRNAs for decay and regulates cell motility in mouse myoblasts. PLoS Genet. 8, e1002901 (2012).
Aubert, G., Baerlocher, G.M., Vulto, I., Poon, S.S. & Lansdorp, P.M. Collapse of telomere homeostasis in hematopoietic cells caused by heterozygous mutations in telomerase genes. PLoS Genet. 8, e1002696 (2012).
Park, I.H. et al. Reprogramming of human somatic cells to pluripotency with defined factors. Nature 451, 141–146 (2008).
Park, I.H., Lerou, P.H., Zhao, R., Huo, H. & Daley, G.Q. Generation of human-induced pluripotent stem cells. Nat. Protoc. 3, 1180–1186 (2008).
Warlich, E. et al. Lentiviral vector design and imaging approaches to visualize the early stages of cellular reprogramming. Mol. Ther. 19, 782–789 (2011).
Bolger, A.M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Magocč, T. & Salzberg, S.L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Kim, D., Langmead, B. & Salzberg, S.L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Patro, R., Duggal, G. & Kingsford, C. Salmon: accurate, versatile and ultrafast quantification from RNA-seq data using lightweight-alignment. bioRxiv 10.1101/021592 (2015).
Acknowledgements
We thank the patients and their families for participation in the research; B.A. Croker, G.Q. Daley and L.I. Zon for comments on the manuscript; and K.E. Gagne for technical assistance. The work was funded in part by the Translational Research Program and the Stem Cell Program, Boston Children's Hospital (S.A.); the Manton Center for Orphan Disease Research (D.H.M.); and the Scientific and Technological Research Council of Turkey (B.B.).
Author information
Authors and Affiliations
Contributions
S.A. and D.H.M. conceived the study, executed experiments, analyzed data, prepared figures and wrote the manuscript. B.B. and M.S. executed experiments, analyzed data and prepared figures. E.G. provided patient information. I.H. provided registry infrastructure. P.C. wrote custom bioinformatics scripts and analyzed the RNA-seq data. A.K.T. performed next-generation sequencing, wrote custom bioinformatics scripts and analyzed the 3′ RACE deep sequencing data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1–7. (PDF 4968 kb)
Rights and permissions
About this article
Cite this article
Moon, D., Segal, M., Boyraz, B. et al. Poly(A)-specific ribonuclease (PARN) mediates 3′-end maturation of the telomerase RNA component. Nat Genet 47, 1482–1488 (2015). https://doi.org/10.1038/ng.3423
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3423
This article is cited by
-
Identification of PARN nuclease activity inhibitors by computational-based docking and high-throughput screening
Scientific Reports (2023)
-
Poly (A)-specific ribonuclease deficiency impacts oogenesis in zebrafish
Scientific Reports (2023)
-
Thymidine nucleotide metabolism controls human telomere length
Nature Genetics (2023)
-
Genetics of human telomere biology disorders
Nature Reviews Genetics (2023)
-
Genome-wide association study of lung adenocarcinoma in East Asia and comparison with a European population
Nature Communications (2023)