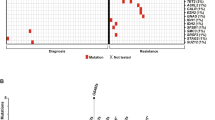
Abstract
TCF3-HLF−positive acute lymphoblastic leukemia (ALL) is currently incurable. Using an integrated approach, we uncovered distinct mutation, gene expression and drug response profiles in TCF3-HLF−positive and treatment-responsive TCF3-PBX1−positive ALL. We identified recurrent intragenic deletions of PAX5 or VPREB1 in constellation with the fusion of TCF3 and HLF. Moreover somatic mutations in the non-translocated allele of TCF3 and a reduction of PAX5 gene dosage in TCF3-HLF ALL suggest cooperation within a restricted genetic context. The enrichment for stem cell and myeloid features in the TCF3-HLF signature may reflect reprogramming by TCF3-HLF of a lymphoid-committed cell of origin toward a hybrid, drug-resistant hematopoietic state. Drug response profiling of matched patient-derived xenografts revealed a distinct profile for TCF3-HLF ALL with resistance to conventional chemotherapeutics but sensitivity to glucocorticoids, anthracyclines and agents in clinical development. Striking on-target sensitivity was achieved with the BCL2-specific inhibitor venetoclax (ABT-199). This integrated approach thus provides alternative treatment options for this deadly disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Mullighan, C.G. et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 446, 758–764 (2007).
Russell, L.J. et al. Deregulated expression of cytokine receptor gene, CRLF2, is involved in lymphoid transformation in B-cell precursor acute lymphoblastic leukemia. Blood 114, 2688–2698 (2009).
Mullighan, C.G. et al. Rearrangement of CRLF2 in B-progenitor- and Down syndrome-associated acute lymphoblastic leukemia. Nat. Genet. 41, 1243–1246 (2009).
Hertzberg, L. et al. Down syndrome acute lymphoblastic leukemia, a highly heterogeneous disease in which aberrant expression of CRLF2 is associated with mutated JAK2: a report from the International BFM Study Group. Blood 115, 1006–1017 (2010).
Holmfeldt, L. et al. The genomic landscape of hypodiploid acute lymphoblastic leukemia. Nat. Genet. 45, 242–252 (2013).
Zhang, J. et al. Key pathways are frequently mutated in high-risk childhood acute lymphoblastic leukemia: a report from the Children's Oncology Group. Blood 118, 3080–3087 (2011).
Irving, J. et al. Ras pathway mutations are highly prevalent in relapsed childhood acute lymphoblastic leukaemia, may act as relapse-drivers and confer sensitivity to MEK inhibition. Blood 124, 3420–3430 (2014).
Inaba, H., Greaves, M. & Mullighan, C.G. Acute lymphoblastic leukaemia. Lancet 381, 1943–1955 (2013).
Kwon, K. et al. Instructive role of the transcription factor E2A in early B lymphopoiesis and germinal center B cell development. Immunity 28, 751–762 (2008).
Felice, M.S. et al. Prognostic impact of t(1;19)/ TCF3–PBX1 in childhood acute lymphoblastic leukemia in the context of Berlin-Frankfurt-Munster-based protocols. Leuk. Lymphoma 52, 1215–1221 (2011).
Hunger, S.P., Ohyashiki, K., Toyama, K. & Cleary, M.L. Hlf, a novel hepatic bZIP protein, shows altered DNA-binding properties following fusion to E2A in t(17;19) acute lymphoblastic leukemia. Genes Dev. 6, 1608–1620 (1992).
Inukai, T. et al. Hypercalcemia in childhood acute lymphoblastic leukemia: frequent implication of parathyroid hormone-related peptide and E2A-HLF from translocation 17;19. Leukemia 21, 288–296 (2007).
Boller, S. & Grosschedl, R. The regulatory network of B-cell differentiation: a focused view of early B-cell factor 1 function. Immunol. Rev. 261, 102–115 (2014).
de Boer, J. et al. The E2A-HLF oncogenic fusion protein acts through Lmo2 and Bcl-2 to immortalize hematopoietic progenitors. Leukemia 25, 321–330 (2011).
Hirose, K. et al. Aberrant induction of LMO2 by the E2A-HLF chimeric transcription factor and its implication in leukemogenesis of B-precursor ALL with t(17;19). Blood 116, 962–970 (2010).
Inukai, T. et al. SLUG, a ces-1-related zinc finger transcription factor gene with antiapoptotic activity, is a downstream target of the E2A-HLF oncoprotein. Mol. Cell 4, 343–352 (1999).
Inoue, A. et al. Slug, a highly conserved zinc finger transcriptional repressor, protects hematopoietic progenitor cells from radiation-induced apoptosis in vivo. Cancer Cell 2, 279–288 (2002).
Honda, H. et al. Expression of E2A-HLF chimeric protein induced T-cell apoptosis, B-cell maturation arrest, and development of acute lymphoblastic leukemia. Blood 93, 2780–2790 (1999).
Smith, K.S., Rhee, J.W., Naumovski, L. & Cleary, M.L. Disrupted differentiation and oncogenic transformation of lymphoid progenitors in E2A-HLF transgenic mice. Mol. Cell. Biol. 19, 4443–4451 (1999).
Hunger, S.P. Chromosomal translocations involving the E2A gene in acute lymphoblastic leukemia: clinical features and molecular pathogenesis. Blood 87, 1211–1224 (1996).
Wiemels, J.L. et al. Site-specific translocation and evidence of postnatal origin of the t(1;19) E2A–PBX1 fusion in childhood acute lymphoblastic leukemia. Proc. Natl. Acad. Sci. USA 99, 15101–15106 (2002).
Tsai, A.G. et al. Human chromosomal translocations at CpG sites and a theoretical basis for their lineage and stage specificity. Cell 135, 1130–1142 (2008).
Moorman, A.V. et al. A novel integrated cytogenetic and genomic classification refines risk stratification in pediatric acute lymphoblastic leukemia. Blood 124, 1434–1444 (2014).
Mangum, D.S. et al. VPREB1 deletions occur independent of lambda light chain rearrangement in childhood acute lymphoblastic leukemia. Leukemia 28, 216–220 (2014).
Waanders, E. et al. The origin and nature of tightly clustered BTG1 deletions in precursor B-cell acute lymphoblastic leukemia support a model of multiclonal evolution. PLoS Genet. 8, e1002533 (2012).
Tijchon, E., Havinga, J., van Leeuwen, F.N. & Scheijen, B. B-lineage transcription factors and cooperating gene lesions required for leukemia development. Leukemia 27, 541–552 (2013).
Balbin, O.A. et al. Reconstructing targetable pathways in lung cancer by integrating diverse omics data. Nat. Commun. 4, 2617 (2013).
Wright, D.D., Sefton, B.M. & Kamps, M.P. Oncogenic activation of the Lck protein accompanies translocation of the LCK gene in the human HSB2 T-cell leukemia. Mol. Cell. Biol. 14, 2429–2437 (1994).
Schmitz, R. et al. Burkitt lymphoma pathogenesis and therapeutic targets from structural and functional genomics. Nature 490, 116–120 (2012).
Ma, X. et al. Rise and fall of subclones from diagnosis to relapse in pediatric B-acute lymphoblastic leukaemia. Nat. Commun. 6, 6604 (2015).
Laurenti, E. et al. The transcriptional architecture of early human hematopoiesis identifies multilevel control of lymphoid commitment. Nat. Immunol. 14, 756–763 (2013).
Subramanian, A., Kuehn, H., Gould, J., Tamayo, P. & Mesirov, J.P. GSEA-P: a desktop application for Gene Set Enrichment Analysis. Bioinformatics 23, 3251–3253 (2007).
Schepers, A.G. et al. Lineage tracing reveals Lgr5+ stem cell activity in mouse intestinal adenomas. Science 337, 730–735 (2012).
Akahane, K. et al. Specific induction of CD33 expression by E2A-HLF: the first evidence for aberrant myeloid antigen expression in ALL by a fusion transcription factor. Leukemia 24, 865–869 (2010).
Nissim, S. et al. Prostaglandin E2 regulates liver versus pancreas cell-fate decisions and endodermal outgrowth. Dev. Cell 28, 423–437 (2014).
Schmitz, M. et al. Xenografts of highly resistant leukemia recapitulate the clonal composition of the leukemogenic compartment. Blood 118, 1854–1864 (2011).
Bonapace, L. et al. Induction of autophagy-dependent necroptosis is required for childhood acute lymphoblastic leukemia cells to overcome glucocorticoid resistance. J. Clin. Invest. 120, 1310–1323 (2010).
Stark, Z. et al. Two novel germline KRAS mutations: expanding the molecular and clinical phenotype. Clin. Genet. 81, 590–594 (2012).
Boutter, J. et al. Image-based RNA interference screening reveals an individual dependence of acute lymphoblastic leukemia on stromal cysteine support. Oncotarget 5, 11501–11512 (2014).
Bicocca, V.T. et al. Crosstalk between ROR1 and the Pre-B cell receptor promotes survival of t(1;19) acute lymphoblastic leukemia. Cancer Cell 22, 656–667 (2012).
Heltemes-Harris, L.M. et al. Ebf1 or Pax5 haploinsufficiency synergizes with STAT5 activation to initiate acute lymphoblastic leukemia. J. Exp. Med. 208, 1135–1149 (2011).
Souers, A.J. et al. ABT-199, a potent and selective BCL-2 inhibitor, achieves antitumor activity while sparing platelets. Nat. Med. 19, 202–208 (2013).
Rolink, A.G., Nutt, S.L., Melchers, F. & Busslinger, M. Long-term in vivo reconstitution of T-cell development by Pax5-deficient B-cell progenitors. Nature 401, 603–606 (1999).
Joshi, I. et al. Loss of Ikaros DNA-binding function confers integrin-dependent survival on pre-B cells and progression to acute lymphoblastic leukemia. Nat. Immunol. 15, 294–304 (2014).
Clappier, E. et al. An intragenic ERG deletion is a marker of an oncogenic subtype of B-cell precursor acute lymphoblastic leukemia with a favorable outcome despite frequent IKZF1 deletions. Leukemia 28, 70–77 (2014).
Grawunder, U. et al. Down-regulation of RAG1 and RAG2 gene expression in preB cells after functional immunoglobulin heavy chain rearrangement. Immunity 3, 601–608 (1995).
Den Boer, M.L. et al. A subtype of childhood acute lymphoblastic leukaemia with poor treatment outcome: a genome-wide classification study. Lancet Oncol. 10, 125–134 (2009).
Andersson, A.K. et al. The landscape of somatic mutations in infant MLL-rearranged acute lymphoblastic leukemias. Nat. Genet. 47, 330–337 (2015).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Liu, D. et al. Leucine-rich repeat-containing G-protein-coupled Receptor 5 marks short-term hematopoietic stem and progenitor cells during mouse embryonic development. J. Biol. Chem. 289, 23809–23816 (2014).
Riddell, J. et al. Reprogramming committed murine blood cells to induced hematopoietic stem cells with defined factors. Cell 157, 549–564 (2014).
Sandler, V.M. et al. Reprogramming human endothelial cells to haematopoietic cells requires vascular induction. Nature 511, 312–318 (2014).
Urbánek, P., Wang, Z.Q., Fetka, I., Wagner, E.F. & Busslinger, M. Complete block of early B cell differentiation and altered patterning of the posterior midbrain in mice lacking Pax5/BSAP. Cell 79, 901–912 (1994).
Simmons, S. et al. Biphenotypic B-lymphoid/myeloid cells expressing low levels of Pax5: potential targets of BAL development. Blood 120, 3688–3698 (2012).
Papaemmanuil, E. et al. RAG-mediated recombination is the predominant driver of oncogenic rearrangement in ETV6-RUNX1 acute lymphoblastic leukemia. Nat. Genet. 46, 116–125 (2014).
Peirs, S. et al. ABT-199 mediated inhibition of BCL-2 as a novel therapeutic strategy in T-cell acute lymphoblastic leukemia. Blood 124, 3738–3747 (2014).
Chonghaile, T.N. et al. Maturation stage of T-cell acute lymphoblastic leukemia determines BCL-2 versus BCL-XL dependence and sensitivity to ABT-199. Cancer Discov. 4, 1074–1087 (2014).
El Omari, K. et al. Structural basis for LMO2-driven recruitment of the SCL:E47bHLH heterodimer to hematopoietic-specific transcriptional targets. Cell Rep. 4, 135–147 (2013).
Conter, V. et al. Molecular response to treatment redefines all prognostic factors in children and adolescents with B-cell precursor acute lymphoblastic leukemia: results in 3184 patients of the AIEOP-BFM ALL 2000 study. Blood 115, 3206–3214 (2010).
Harrison, C.J. et al. Detection of prognostically relevant genetic abnormalities in childhood B-cell precursor acute lymphoblastic leukaemia: recommendations from the Biology and Diagnosis Committee of the International Berlin-Frankfurt-Munster study group. Br. J. Haematol. 151, 132–142 (2010).
Case, M. et al. Mutation of genes affecting the RAS pathway is common in childhood acute lymphoblastic leukemia. Cancer Res. 68, 6803–6809 (2008).
Dörge, P. et al. IKZF1 deletion is an independent predictor of outcome in pediatric acute lymphoblastic leukemia treated according to the ALL-BFM 2000 protocol. Haematologica 98, 428–432 (2013).
Bauer, M.J., Cox, A.J. & Evers, D.J. Fast gapped read mapping for Illumina reads. In ISMB, ISBC (2010).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Xi, R. et al. Copy number variation detection in whole-genome sequencing data using the Bayesian information criterion. Proc. Natl. Acad. Sci. USA 108, E1128–E1136 (2011).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
McPherson, A. et al. deFuse: an algorithm for gene fusion discovery in tumor RNA-Seq data. PLOS Comput. Biol. 7, e1001138 (2011).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Forster, M. et al. From next-generation sequencing alignments to accurate comparison and validation of single-nucleotide variants: the pibase software. Nucleic Acids Res. 41, e16 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Koboldt, D.C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Albers, C.A. et al. Dindel: accurate indel calls from short-read data. Genome Res. 21, 961–973 (2011).
Mortazavi, A., Williams, B.A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Frisch, M., Klocke, B., Haltmeier, M. & Frech, K. LitInspector: literature and signal transduction pathway mining in PubMed abstracts. Nucleic Acids Res. 37, W135–W140 (2009).
Ho Sui, S.J. et al. oPOSSUM: identification of over-represented transcription factor binding sites in co-expressed genes. Nucleic Acids Res. 33, 3154–3164 (2005).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Stade, B., Seelow, D., Thomsen, I., Krawczak, M. & Franke, A. GrabBlur–a framework to facilitate the secure exchange of whole-exome and -genome SNV data using VCF files. BMC Genomics 15 (suppl. 4), S8 (2014).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Ratei, R. et al. Lineage classification of childhood acute lymphoblastic leukemia according to the EGIL recommendations: results of the ALL-BFM 2000 trial. Klin. Padiatr. 225 (suppl. 1), S34–S39 (2013).
Festing, M.F. & Altman, D.G. Guidelines for the design and statistical analysis of experiments using laboratory animals. ILAR J. 43, 244–258 (2002).
Acknowledgements
We thank all participants and personnel involved in the clinical trials in Austria, France, Germany, United Kingdom and Switzerland. We thank T. Radimerski and Novartis for providing essential compounds. We thank the Leukaemia & Lymphoma Research (LLR) Childhood Leukaemia Cell Bank in the UK for providing primary patient samples. This work was supported by the German Federal Office for Radiation Protection (grant St.Sch. 3611S70014), by the Swiss National Research Foundation SNF 310030-133108, the foundation 'Kinderkrebsforschung Schweiz', the 'Krebsliga Zurich', the Sassella Foundation, the Fondation Panacée, the clinical research focus program 'Human Hemato-Lymphatic Diseases' of the University of Zurich, the Deutsche Forschungsgemeinschaft (DFG), Clusters of Excellence 'Inflammation at Interfaces', the EU Seventh Framework Program (FP7/2007-2013, grant 262055, ESGI; FP7-HEALTH-F2-2011 grant 261474, ENCCA; ERA-Net Transcan, Validation of biomarkers for personalized cancer medicine, TRANSCALL; Health-F2-2010 grant 260791, EUROCANPLATFORM), the 'Katharina Hardt Stiftung', the 'Deutsche José Carreras Leukämie-Stiftung', the 'Madeleine Schickedanz-Kinderkrebs-Stiftung', the 'Deutsche Krebshilfe – Dr. Mildred Scheel Stiftung' (grants 108613, 102588 and 108588), the Foundation of Experimental Biomedicine in Zurich, the Max Planck Society, and the 'Verein für krebskranke Kinder Hannover e.V.'. We thank A. Dehos, B. Grosche, T. Jung, W. Weiss and G. Ziegelberger, German Federal Office for Radiation Protection, as well as B. Heinzow, State Office for Social Services of Schleswig-Holstein, and A. Böttger, German Federal Ministry for the Environment, Nature Conservation, Building and Nuclear Safety for their support and critical discussions. We are grateful for the excellent technical assistance offered by the sequencing team of the Department of Vertebrate Genomics of the Max Planck Institute for Molecular Genetics (Berlin) and by the team of the Genomics Core facility of the European Molecular Biology Laboratory. We thank K. Alemazkour for excellent technical assistance regarding whole exome sequencing at the Department of Pediatric Oncology, Hematology and Clinical Immunology (Düsseldorf, Germany). We thank N. Forgo, Institute for Legal Informatics, Leibniz University Hannover, and H.-D. Tröger, Hannover Medical School, for legal and ethical counselling.
Author information
Authors and Affiliations
Contributions
A. Borkhardt, A.F., J.O.K., J.-P.B., M.-L.Y. and M. Stanulla jointly designed the project. A. Baruchel, A.V., A.V.M., C.C., C.P.K., F.N., G.B., G.C., G.t.K., H.C., M. Schrappe, M. Stanulla, N.v.d.W., O.A.H., C.E. and R.P.-G. provided samples or clinical data. A. Borkhardt, A.C.M., A.F., A.V., C.E., G.H.-S., H.L., J.-P.B., J.O.K., J.T., M.D., M.F., M.-L.Y., M.P.D., M. Schrappe, M. Stanulla, M. Zimmermann, O.A.H., P.R. and T.Z. contributed reagents, materials or analysis tools. A.M.S., B.B., B.M., C.E., C.L.W., H.-J.W., J.I.H., J.-P.B., M.F., M.G., S. Sungalee and U.F. designed experiments. A.M.S., A.R., B.B., B.R., B.S., B.S.P., C.E., C.K., C.L.W., D.D., D.S., H.-J.W., J.T., K.H., L.Z., M.G., M.R., M. Zaliova, M. Sultan, P.K., S.E., S. Sungalee, T.B., U.F. and V.F. performed experiments. A.R., B.S., B.S.P., C.E., C.K., C.L.W., D.S., E.E., J.I.H., H.-J.W., M.G., M.P.D., M.F., N.B., S.G., G.H.-S., P.H., P.K., M.-L.Y., M.R., M. Stanulla, M. Schütte, M. Zaliova, S. Sungalee, T.R., U.F. and V.A. analyzed data. A. Borkhardt, A.C.M., A.F., B.B., H.L., J.-P.B., J.O.K., M.D., M.F., M.-L.Y., M. Stanulla, O.H., R.T. and S. Schreiber, U.F. supervised research. A.R., C.L.W., D.S., H.-J.W., M.F., M.P.D., M. Stanulla, P.K., S. Sungalee, T.R. and U.F. prepared tables and figures. J.-P.B., M. Stanulla and M.-L.Y. wrote the manuscript. A. Borkhardt, A.F., A.R., A.V.M., H.-J.W., J.O.K., M.F., M.P.D., O.H., S.H., S. Sungalee, T.R. and U.F. contributed to the writing of the manuscript. All authors critically reviewed the manuscript for its content.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 TCF3 breakpoints of TCF3-PBX1 (patients 1a−5a) and TCF3-HLF (patients 6a−9a and11a) translocations.
The CpG motifs closest to the breakpoints are highlighted in red boxes and the closest cac motifs in blue.
Supplementary Figure 2 The TCF3-HLF translocation is only detected in the lymphoid progenitor gate.
Experiments, combining sorting of specific cell populations using flow cytometry (a) and subsequent detection of the fusion gene in sorted cells employing FISH probes or real-time PCR (b), were performed using bone marrow from the TCF3-HLF-positive patients 15, 16, and 17 of the validation cohort. Bone marrow cells were sorted into HSC (CD34+CD38-CD90+CD45RA-) and MLP (CD34+CD38-CD90neg-lowCD45RA+) populations as previously described (Doulatov et al. Nature Immunology 11, 585-593 (2010)). Leukemic cells (CD34-CD38+CD19+CD10+) were sorted according to standard diagnostic markers. Sorted cell populations were analyzed by FISH and real-time PCR as described in the Online Methods section. All sorted leukemia cells carried the translocation (FISH signals 2R, 1G, 1B, 2B/G fusion) while none of the scored HSC or MLP (2R, 2G, 2B) populations were positive. The TCF3-HLF fusion was detected only in blast populations and not in myeloid cells indicating that the TCF3-HLF-positive leukemia originated in committed lymphoid cells. For patient 15, the results were confirmed using a TCF3-HLF-fusion-specific quantitative real-time PCR on a fraction of sorted cells as described in the supplementary online methods section. Leukemic cells were 100% positive for the TCF3-HLF-fusion, whereas the MLP and the HSC fractions were negative. As a positive control the reference gene β-globin was employed. The TCF3-HLF fusion was detected by real-time PCR only in the blast population, but not in myeloid cells, indicating that the TCF3-HLF-positive leukemia may have originated from committed lymphoid cells.
Supplementary Figure 3 BTG1 deletions.
(a) Whole genome sequencing (WGS) read ratio plot showing a BTG1 deletion identified in the diagnostic sample of patient 8. This BTG1 deletion was present in a subclonal population of ~14% of leukemia cells. (b) WGS read ratio plot showing a BTG1 deletion identified in the diagnostic sample of patient 9. (c) Exome sequencing read ratio plot for a xenograft (7b_X) derived from the remission sample of patient 7 showing a BTG1 deletion. (d) The BTG1 deletion in xenograft 7b_X ia also visible in the SNP allele frequency data from exome sequencing, inspecting the SNP rs709222 in BTG1 exon 2.
Supplementary Figure 4 KHDRBS1-LCK fusion gene in patient 6a.
(a) A heterozygous deletion encompassing 223 kb of genomic DNA on chromosome 1 fused intron 1 of KHDRBS1 to the promoter region of the LCK proto-oncogene. (b) Transcriptome sequencing showed a decrease of KHDRBS1 expression after exon 1. (c) LCK expression was induced starting from exon 2. (d) Analysis of chimeric fusion reads showed the fusion of KHDRBS1 exon 1 with LCK exon 2, confirmed by 14 chimeric reads spanning the fusion border. (e) Amino acid sequence of the in-frame fusion protein encoded by the KHDRBS1-LCK fusion transcript.
Supplementary Figure 5 Recurrent fusion of the genes KHDRBS1 and LCK in ALL.
RNA was isolated from samples of a cohort of 74 unselected pediatric ALL and transcribed into cDNA. Nested PCR was performed using specific primers for the KHDRBS1-LCK fusion transcript detected in patient 6a. (a) The gel electrophoresis presents the expected specific PCR product of 226 bp in the sample 6a and three patient samples of the validation cohort as well as controls. The negative control is representative of the 71 patient samples of this validation cohort tested negative. (b) The chromatograms display the sequencing result for patient 6a (upper panel) and a patient of the validation cohort (patient D3, lower panel) as an example. The fusion of KHDRBS1 exon 1 to LCK exon 2 was confirmed by Sanger sequencing in all positive cases (3/74).
Supplementary Figure 6 Pattern of recurrent genomic lesions and model of leukemogenesis in TCF3-HLF−positive leukemia.
Left panel: Recurrent genetic lesions identified in TCF3-HLF-positive ALL affect transcription factors regulating lymphoid differentiation, the Ras pathway and cell cycle regulation. Besides the 12 cases analyzed in the present study, a single case reported in a recent study is included (Ma et al. Nature Communications 6, 6604 (2015), DOI: 10.1038/ncomms7604) (x indicates a KRAS mutation that was detected only in a patient derived xenograft sample, not in the primary tissue. Note that the cases 16 and 17 have not been analyzed by next generation sequencing and besides PAX5 deletion, NRAS and KRAS hotspot mutations other genetic alterations possibly present are not yet analyzed). Right panel: The TCF3-HLF gene fusion probably occurs in an early B cell progenitor. B cell differentiation is blocked by mutations in transcription factors playing key roles in lymphoid development. Secondary mutations involving the Ras pathway and cell cycle regulators lead to leukemia development.
Supplementary Figure 7 Engraftment of TCF3-PBX1− and TCF3-HLF−positive ALL samples.
ALL cells were recovered from cryopreserved samples and transplanted into NSG mice. The percentage of mCD45-hCD19+hCD45+ leukemic cells was monitored by flow cytometry every 4 weeks. Upper panels: Primary and secondary transplantation of diagnostic TCF3-PBX1-positive samples. Middle panels: Primary and secondary transplantation of diagnostic TCF3-HLF-positive samples. Lower panels: Primary and secondary transplantation of MRD and relapse TCF3-HLF-positive samples. MRD samples were successfully amplified only for the TCF3-HLF-positive subtype. Patient samples were transplanted intrafemorally (primary transplantation). Xenograft-samples were transplanted intravenously (secondary transplantation). Mean and SD of 2-3 mice is shown when available. The number of living cells transplanted is indicated in the legend.
Supplementary Figure 8 Expression of PAX5 in primary and patient derived xenografts (PDX) of TCF3-PBX1− and TCF3-HLF−positive ALL samples.
The significant differential expression of PAX5 between TCF3-PBX1-positive and TCF3-HLF-positive primary ALL is preserved in patient derived xenograft samples.
Supplementary Figure 9 Cell survival and cell cycle analysis of TCF3-PBX1− and TCF3-HLF−positive ALL cells in coculture on hMSCs.
Upper panel: ALL cell viability comparing co-cultures to ALL cell suspension cultures after 72 h by flow cytometry using 7-AAD staining. The percentage of living cells normalized to the input cells seeded on day 0 is shown (mean with SD of duplicates). Lower panel: Percentage of cells of the population in the G0–G1, S and G2–M phases of the cell cycle. Flow cytometric analysis was performed at day 1 of co-culture with hMSC after 20 h of EdU incubation. Mean and SD of at least 2 independent experiments performed in triplicates are shown.
Supplementary Figure 10 Drug profiling of TCF3-PBX1 and TCF3-HLF ALL xenografts derived from diagnostic, MRD and relapse samples.
Unsupervised clustering of xenograft samples according to their response (log IC50) to 98 compounds in co-culture on hMSC after 72 h. Xenografts derived from diagnostic „a“, MRD „b“ and relapse „c“ samples are included. Fitted values are provided in the supplementary section (absolute IC50). Compounds are arranged based on the family of the targets. Numbers identify the compounds shown in Figure 5c.
Supplementary Figure 11 In vitro dose response curves to ABT-199 (venetoclax) comparing xenografts derived from TCF3-HLF-positive ALL patients at the time point of diagnosis 'a', MRD 'b' and relapse 'c'.
ALL co-cultures were treated with ABT-199 for 72 hours, the responses were normalized against DMSO-treated controls.
Supplementary Figure 12 Drug activity profiling of two additional TCF3-HLF−translocated leukemia cases of the validation cohort (15, 17) reveals similar patterns of drug sensitivities and high responsiveness to ABT-199 (venetoclax) compared to the TCF3-HLF−positive patients of the discovery cohort.
Selection of drugs based on differences in sensitivity or resistance in TCF3-PBX1-positive compared to TCF3-HLF-positive patients (Figure 5c). Similar patterns of drug sensitivity were observed testing patient derived primary (patient 15, 17) and xenograft cells (other TCF3-PBX1- and TCF3-HLF-positive cases shown). Drugs active across both patient cohorts include doxorubicin, idarubicin, mitoxantrone, bortezomib, and ABT-199 (venetoclax).
Supplementary Figure 13 In vitro dose response co-titration curves to ABT-199/dexamethasone (top) and ABT-199/vincristine (bottom) for patient derived xenografts from TCF3-HLF−positive (6a−11a) ALL patients.
Xenograft cells were seeded on a layer of stroma cell and treated with the indicated combinations and concentrations of ABT-199, vincristine and dexamethasone, respectively, for 72 h. Drug concentrations used in these assays are provided in Supplementary Table 26. The response was normalized against DMSO-treated controls. The co-titration curves indicate that ABT-199 combined with either vincristine or dexamethasone likely exhibits synergistic activity in the patients marked with (*); combination indices (see Table) were calculated based on the Chou-Talaly method (Chou and Talaly JBC 252: 6438-6442, (1977)).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 and Supplementary Tables 1–4 and 8. (PDF 2415 kb)
Supplementary Tables 5–7 and 9–27
Supplementary Tables 5–7 and 9–27. (XLSX 685 kb)
Rights and permissions
About this article
Cite this article
Fischer, U., Forster, M., Rinaldi, A. et al. Genomics and drug profiling of fatal TCF3-HLF−positive acute lymphoblastic leukemia identifies recurrent mutation patterns and therapeutic options. Nat Genet 47, 1020–1029 (2015). https://doi.org/10.1038/ng.3362
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3362
This article is cited by
-
Novel Biomarkers and Molecular Targets in ALL
Current Hematologic Malignancy Reports (2024)
-
International Consensus Classification of acute lymphoblastic leukemia/lymphoma
Virchows Archiv (2023)
-
Pharmacotypes across the genomic landscape of pediatric acute lymphoblastic leukemia and impact on treatment response
Nature Medicine (2023)
-
An alternative CYB5A transcript is expressed in aneuploid ALL and enriched in relapse
BMC Genomic Data (2022)
-
Potent, p53-independent induction of NOXA sensitizes MLL-rearranged B-cell acute lymphoblastic leukemia cells to venetoclax
Oncogene (2022)