Abstract
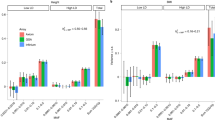
Hundreds of genes reside in structurally complex, poorly understood regions of the human genome1,2,3. One such region contains the three amylase genes (AMY2B, AMY2A and AMY1) responsible for digesting starch into sugar. Copy number of AMY1 is reported to be the largest genomic influence on obesity4, although genome-wide association studies for obesity have found this locus unremarkable. Using whole-genome sequence analysis3,5, droplet digital PCR6 and genome mapping7, we identified eight common structural haplotypes of the amylase locus that suggest its mutational history. We found that the AMY1 copy number in an individual's genome is generally even (rather than odd) and partially correlates with nearby SNPs, which do not associate with body mass index (BMI). We measured amylase gene copy number in 1,000 obese or lean Estonians and in 2 other cohorts totaling ∼3,500 individuals. We had 99% power to detect the lower bound of the reported effects on BMI4, yet found no association.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Conrad, D.F. et al. Origins and functional impact of copy number variation in the human genome. Nature 464, 704–712 (2010).
Sudmant, P.H. et al. Diversity of human copy number variation and multicopy genes. Science 330, 641–646 (2010).
Handsaker, R.E. et al. Large multiallelic copy number variations in humans. Nat. Genet. 47, 296–303 (2015).
Falchi, M. et al. Low copy number of the salivary amylase gene predisposes to obesity. Nat. Genet. 46, 492–497 (2014).
Handsaker, R.E., Korn, J.M., Nemesh, J. & McCarroll, S.A. Discovery and genotyping of genome structural polymorphism by sequencing on a population scale. Nat. Genet. 43, 269–276 (2011).
Hindson, B.J. et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal. Chem. 83, 8604–8610 (2011).
Hastie, A.R. et al. Rapid genome mapping in nanochannel arrays for highly complete and accurate de novo sequence assembly of the complex Aegilops tauschii genome. PLoS ONE 8, e55864 (2013).
Groot, P.C. et al. The human α-amylase multigene family consists of haplotypes with variable numbers of genes. Genomics 5, 29–42 (1989).
Perry, G.H. et al. Diet and the evolution of human amylase gene copy number variation. Nat. Genet. 39, 1256–1260 (2007).
Groot, P.C., Mager, W.H. & Frants, R.R. Interpretation of polymorphic DNA patterns in the human α-amylase multigene family. Genomics 10, 779–785 (1991).
Locke, A.E. et al. Genetic studies of body mass index yield new insights for obesity biology. Nature 518, 197–206 (2015).
Cantsilieris, S. & White, S.J. Correlating multiallelic copy number polymorphisms with disease susceptibility. Hum. Mutat. 34, 1–13 (2013).
Barnes, C. et al. A robust statistical method for case-control association testing with copy number variation. Nat. Genet. 40, 1245–1252 (2008).
Clayton, D.G. et al. Population structure, differential bias and genomic control in a large-scale, case-control association study. Nat. Genet. 37, 1243–1246 (2005).
Zanda, M. et al. A genome-wide assessment of the role of untagged copy number variants in type 1 diabetes. PLoS Genet. 10, e1004367 (2014).
1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
International HapMap Consortium. The International HapMap Project. Nature 426, 789–796 (2003).
Carpenter, D. et al. Obesity, starch digestion and amylase: association between copy number variants at human salivary (AMY1) and pancreatic (AMY2) amylase genes. Hum. Mol. Genet. 24, 3472–3480 (2015).
Groot, P.C. et al. Evolution of the human α-amylase multigene family through unequal, homologous, and inter- and intrachromosomal crossovers. Genomics 8, 97–105 (1990).
Boettger, L.M., Handsaker, R.E., Zody, M.C. & McCarroll, S.A. Structural haplotypes and recent evolution of the human 17q21.31 region. Nat. Genet. 44, 881–885 (2012).
Steinberg, K.M. et al. Structural diversity and African origin of the 17q21.31 inversion polymorphism. Nat. Genet. 44, 872–880 (2012).
Lupski, J.R. & Stankiewicz, P. Genomic disorders: molecular mechanisms for rearrangements and conveyed phenotypes. PLoS Genet. 1, e49 (2005).
Leitsalu, L. et al. Cohort Profile: Estonian Biobank of the Estonian Genome Center, University of Tartu. Int. J. Epidemiol. doi:10.1093/ije/dyt268 (2014).
Speliotes, E.K. et al. Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat. Genet. 42, 937–948 (2010).
Ferrucci, L. et al. Subsystems contributing to the decline in ability to walk: bridging the gap between epidemiology and geriatric practice in the InCHIANTI study. J. Am. Geriatr. Soc. 48, 1618–1625 (2000).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Wellcome Trust Case Control Consortium. Genome-wide association study of CNVs in 16,000 cases of eight common diseases and 3,000 shared controls. Nature 464, 713–720 (2010).
Tognon, G. et al. Mediterranean diet, overweight and body composition in children from eight European countries: cross-sectional and prospective results from the IDEFICS study. Nutr. Metab. Cardiovasc. Dis. 24, 205–213 (2014).
Mottus, R. et al. Personality traits and eating habits in a large sample of Estonians. Health Psychol. 31, 806–814 (2012).
Berndt, S.I. et al. Genome-wide meta-analysis identifies 11 new loci for anthropometric traits and provides insights into genetic architecture. Nat. Genet. 45, 501–512 (2013).
Mejía-Benítez, M.A. et al. Beneficial effect of a high number of copies of salivary amylase AMY1 gene on obesity risk in Mexican children. Diabetologia 58, 290–294 (2015).
Iskow, R.C., Gokcumen, O. & Lee, C. Exploring the role of copy number variants in human adaptation. Trends Genet. 28, 245–257 (2012).
Stankiewicz, P. & Lupski, J.R. Structural variation in the human genome and its role in disease. Annu. Rev. Med. 61, 437–455 (2010).
Zhang, F., Gu, W., Hurles, M.E. & Lupski, J.R. Copy number variation in human health, disease, and evolution. Annu. Rev. Genomics Hum. Genet. 10, 451–481 (2009).
Zeggini, E. et al. Replication of genome-wide association signals in UK samples reveals risk loci for type 2 diabetes. Science 316, 1336–1341 (2007).
Scott, L.J. et al. A genome-wide association study of type 2 diabetes in Finns detects multiple susceptibility variants. Science 316, 1341–1345 (2007).
Diabetes Genetics Initiative of Broad Institute of Harvard and MIT. Genome-wide association analysis identifies loci for type 2 diabetes and triglyceride levels. Science 316, 1331–1336 (2007).
Heid, I.M. et al. Genetic architecture of the APM1 gene and its influence on adiponectin plasma levels and parameters of the metabolic syndrome in 1,727 healthy Caucasians. Diabetes 55, 375–384 (2006).
Melzer, D. et al. A genome-wide association study identifies protein quantitative trait loci (pQTLs). PLoS Genet. 4, e1000072 (2008).
Church, D.M. et al. Modernizing reference genome assemblies. PLoS Biol. 9, e1001091 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Koren, A. et al. Differential relationship of DNA replication timing to different forms of human mutation and variation. Am. J. Hum. Genet. 91, 1033–1040 (2012).
Cao, H. et al. Rapid detection of structural variation in a human genome using nanochannel-based genome mapping technology. Gigascience 3, 34 (2014).
Teague, B. et al. High-resolution human genome structure by single-molecule analysis. Proc. Natl. Acad. Sci. USA 107, 10848–10853 (2010).
Hach, F. et al. mrsFAST: a cache-oblivious algorithm for short-read mapping. Nat. Methods 7, 576–577 (2010).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Alkan, C. et al. Personalized copy number and segmental duplication maps using next-generation sequencing. Nat. Genet. 41, 1061–1067 (2009).
Browning, S.R. & Browning, B.L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Purcell, S., Cherny, S.S. & Sham, P.C. Genetic Power Calculator: design of linkage and association genetic mapping studies of complex traits. Bioinformatics 19, 149–150 (2003).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2015).
Acknowledgements
This work was supported by a grant from the National Human Genome Research Institute (R01 HG006855) to S.A.M. to support C.L.U., R.E.H. and S.A.M. Work by T.E. and A.M. was supported through the Estonian Genome Center of the University of Tartu (EGCUT) by Targeted Financing from the Estonian Ministry of Science and Education (SF0180142s08), the Development Fund of the University of Tartu (SP1GVARENG) and the European Regional Development Fund to the Centre of Excellence in Genomics (3.2.0304.11-0312) and through Framework Programme 7 grant 313010. T.E., A.M. and J.N.H. were further supported by the US National Institutes of Health (R01 DK075787). T.M.F. is supported by European Research Council funding (Framework Programme 7, SZ-50371-GLUCOSEGENES), M.A.T. and M.N.W. are supported by the Wellcome Trust Institutional Strategic Support Award (WT097835MF), and M.B. is supported by US National Institutes of Health grant DK062370.
Author information
Authors and Affiliations
Contributions
C.L.U., J.N.H. and S.A.M. conceived the project. C.L.U. pursued molecular (ddPCR) and statistical analyses of amylase locus structural variation. R.E.H. contributed analyses of whole-genome sequence data. T.E., A.M., C.L.U., J.E.M. and J.N.H. analyzed the Estonian cohort. M.A.T., M.N.W., T.M.F., R.E.H. and S.K. analyzed the InCHIANTI cohort. M.I.M., M.B., D.M.A., R.E.H., C.L.U. and C.F. analyzed the GoT2D cohort. C.N.P., M.T.P., C.L.U. and R.E.H. analyzed the GPC cohort. A.R.H. and H.C. performed the NanoChannel-based genome mapping. C.L.U., J.N.H. and S.A.M. wrote the manuscript, with contributions from D.M.A., T.M.F., M.B., M.I.M. and T.E.
Corresponding authors
Ethics declarations
Competing interests
A.R.H. and H.C. are employees at BioNano Genomics, Inc., and own company stock options.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–17. (PDF 7125 kb)
Supplementary Tables 1–14
Supplementary Tables 1–14. (XLSX 1504 kb)
Rights and permissions
About this article
Cite this article
Usher, C., Handsaker, R., Esko, T. et al. Structural forms of the human amylase locus and their relationships to SNPs, haplotypes and obesity. Nat Genet 47, 921–925 (2015). https://doi.org/10.1038/ng.3340
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3340
This article is cited by
-
A genomics perspective of personalized prevention and management of obesity
Human Genomics (2024)
-
Starch intake, amylase gene copy number variation, plasma proteins, and risk of cardiovascular disease and mortality
BMC Medicine (2023)
-
Structural variation of the coding and non-coding human pharmacogenome
npj Genomic Medicine (2023)
-
Genetic factors associated with serum amylase in a Japanese population: combined analysis of copy-number and single-nucleotide variants
Journal of Human Genetics (2023)
-
GeneToCN: an alignment-free method for gene copy number estimation directly from next-generation sequencing reads
Scientific Reports (2023)