Abstract
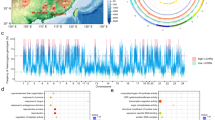
Forest trees are dominant components of terrestrial ecosystems that have global ecological and economic importance. Despite distributions that span wide environmental gradients, many tree populations are locally adapted, and mechanisms underlying this adaptation are poorly understood. Here we use a combination of whole-genome selection scans and association analyses of 544 Populus trichocarpa trees to reveal genomic bases of adaptive variation across a wide latitudinal range. Three hundred ninety-seven genomic regions showed evidence of recent positive and/or divergent selection and enrichment for associations with adaptive traits that also displayed patterns consistent with natural selection. These regions also provide unexpected insights into the evolutionary dynamics of duplicated genes and their roles in adaptive trait variation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
The 1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Cao, J. et al. Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nat. Genet. 43, 956–963 (2011).
Axelsson, E. et al. The genomic signature of dog domestication reveals adaptation to a starch-rich diet. Nature 495, 360–364 (2013).
Hufford, M.B. et al. Comparative population genomics of maize domestication and improvement. Nat. Genet. 44, 808–811 (2012).
Huang, X. et al. Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat. Genet. 44, 32–39 (2012).
Zhao, S. et al. Whole-genome sequencing of giant pandas provides insights into demographic history and local adaptation. Nat. Genet. 45, 67–71 (2013).
Miller, W. et al. Polar and brown bear genomes reveal ancient admixture and demographic footprints of past climate change. Proc. Natl. Acad. Sci. USA 109, E2382–E2390 (2012).
Voight, B.F., Kudaravalli, S., Wen, X. & Pritchard, J.K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Fournier-Level, A. et al. A map of local adaptation in Arabidopsis thaliana. Science 334, 86–89 (2011).
Tishkoff, S.A. et al. Convergent adaptation of human lactase persistence in Africa and Europe. Nat. Genet. 39, 31–40 (2007).
Jia, G. et al. A haplotype map of genomic variations and genome-wide association studies of agronomic traits in foxtail millet (Setaria italica). Nat. Genet. 45, 957–961 (2013).
Hancock, A.M. et al. Adaptation to climate across the Arabidopsis thaliana genome. Science 334, 83–86 (2011).
Grossman, S.R. et al. Identifying recent adaptations in large-scale genomic data. Cell 152, 703–713 (2013).
Savolainen, O., Pyhäjärvi, T. & Knürr, T. Gene flow and local adaptation in trees. Annu. Rev. Ecol. Evol. Syst. 38, 595–619 (2007).
Bonan, G.B. Forests and climate change: forcings, feedbacks, and the climate benefits of forests. Science 320, 1444–1449 (2008).
Ellison, A.M. et al. Loss of foundation species: consequences for the structure and dynamics of forested ecosystems. Front. Ecol. Environ 3, 479–486 (2005).
Whitham, T.G. et al. Extending genomics to natural communities and ecosystems. Science 320, 492–495 (2008).
Parmesan, C. Ecological and evolutionary responses to recent climate change. Annu. Rev. Ecol. Evol. Syst. 37, 637–669 (2006).
Ingvarsson, P.K., García, M.V., Hall, D., Luquez, V. & Jansson, S. Clinal variation in phyB2, a candidate gene for day-length-induced growth cessation and bud set, across a latitudinal gradient in European aspen (Populus tremula). Genetics 172, 1845–1853 (2006).
Neale, D.B. & Kremer, A. Forest tree genomics: growing resources and applications. Nat. Rev. Genet. 12, 111–122 (2011).
Jansson, S. & Douglas, C.J. Populus: a model system for plant biology. Annu. Rev. Plant Biol. 58, 435–458 (2007).
Slavov, G.T. et al. Genome resequencing reveals multiscale geographic structure and extensive linkage disequilibrium in the forest tree Populus trichocarpa. New Phytol. 196, 713–725 (2012).
Pauley, S.S. & Perry, T.O. Ecotypic variation in the photoperiodic response in Populus. J. Arnold Arbor. 35, 167–188 (1954).
Howe, G.T. et al. From genotype to phenotype: unraveling the complexities of cold adaptation in forest trees. Can. J. Bot. 81, 1247–1266 (2003).
McKown, A.D. et al. Geographical and environmental gradients shape phenotypic trait variation and genetic structure in Populus trichocarpa. New Phytol. 201, 1263–1276 (2014).
Wegrzyn, J.L. et al. Association genetics of traits controlling lignin and cellulose biosynthesis in black cottonwood (Populus trichocarpa, Salicaceae) secondary xylem. New Phytol. 188, 515–532 (2010).
Porth, I. et al. Genome-wide association mapping for wood characteristics in Populus identifies an array of candidate single nucleotide polymorphisms. New Phytol. 200, 710–726 (2013).
Tang, H. et al. Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Res. 18, 1944–1954 (2008).
Tuskan, G.A. et al. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313, 1596–1604 (2006).
Rodgers-Melnick, E., Culp, M. & DiFazio, S.P. Predicting whole genome protein interaction networks from primary sequence data in model and non-model organisms using ENTS. BMC Genomics 14, 608 (2013).
Rodgers-Melnick, E. et al. Contrasting patterns of evolution following whole genome versus tandem duplication events in Populus. Genome Res. 22, 95–105 (2012).
Spitze, K. Population structure in Daphnia obtusa: quantitative genetic and allozymic variation. Genetics 135, 367–374 (1993).
Yang, W.-Y., Novembre, J., Eskin, E. & Halperin, E. A model-based approach for analysis of spatial structure in genetic data. Nat. Genet. 44, 725–731 (2012).
Günther, T. & Coop, G. Robust identification of local adaptation from allele frequencies. Genetics 195, 205–220 (2013).
Sun, J., Xie, D., Zhao, H. & Zou, D. Genome-wide identification of the class III aminotransferase gene family in rice and expression analysis under abiotic stress. Genes Genomics 35, 597–608 (2013).
Kang, H.M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Zhou, X. & Stephens, M. Efficient multivariate linear mixed model algorithms for genome-wide association studies. Nat. Methods 11, 407–409 (2014).
Ruttink, T. et al. A molecular timetable for apical bud formation and dormancy induction in poplar. Plant Cell 19, 2370–2390 (2007).
Werner, A.K. et al. The ureide-degrading reactions of purine ring catabolism employ three amidohydrolases and one aminohydrolase in Arabidopsis, soybean, and rice. Plant Physiol. 163, 672–681 (2013).
Hsu, C.-Y. et al. FLOWERING LOCUS T duplication coordinates reproductive and vegetative growth in perennial poplar. Proc. Natl. Acad. Sci. USA 108, 10756–10761 (2011).
Iñigo, S., Alvarez, M.J., Strasser, B., Califano, A. & Cerdán, P.D. PFT1, the MED25 subunit of the plant Mediator complex, promotes flowering through CONSTANS dependent and independent mechanisms in Arabidopsis. Plant J. 69, 601–612 (2012).
Rinne, P.L.H. et al. Chilling of dormant buds hyperinduces FLOWERING LOCUS T and recruits GA-inducible 1,3-β-glucanases to reopen signal conduits and release dormancy in Populus. Plant Cell 23, 130–146 (2011).
Hall, D. et al. Adaptive population differentiation in phenology across a latitudinal gradient in European aspen (Populus tremula, L.): a comparison of neutral markers, candidate genes and phenotypic traits. Evolution 61, 2849–2860 (2007).
Pritchard, J.K. & Di Rienzo, A. Adaptation—not by sweeps alone. Nat. Rev. Genet. 11, 665–667 (2010).
Platt, A., Vilhjálmsson, B.J. & Nordborg, M. Conditions under which genome-wide association studies will be positively misleading. Genetics 186, 1045–1052 (2010).
Atwell, S. et al. Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 465, 627–631 (2010).
Böhlenius, H. et al. CO/FT regulatory module controls timing of flowering and seasonal growth cessation in trees. Science 312, 1040–1043 (2006).
Mohamed, R. et al. Populus CEN/TFL1 regulates first onset of flowering, axillary meristem identity and dormancy release in Populus. Plant J. 62, 674–688 (2010).
Jaeger, K.E., Pullen, N., Lamzin, S., Morris, R.J. & Wigge, P.A. Interlocking feedback loops govern the dynamic behavior of the floral transition in Arabidopsis. Plant Cell 25, 820–833 (2013).
Freeling, M. Bias in plant gene content following different sorts of duplication: tandem, whole-genome, segmental, or by transposition. Annu. Rev. Plant Biol. 60, 433–453 (2009).
Birchler, J.A. & Veitia, R.A. The gene balance hypothesis: implications for gene regulation, quantitative traits and evolution. New Phytol. 186, 54–62 (2010).
Lynch, M. & Force, A. The probability of duplicate gene preservation by subfunctionalization. Genetics 154, 459–473 (2000).
Taylor, J.S. & Raes, J. Duplication and divergence: the evolution of new genes and old ideas. Annu. Rev. Genet. 38, 615–643 (2004).
Vatén, A. et al. Callose biosynthesis regulates symplastic trafficking during root development. Dev. Cell 21, 1144–1155 (2011).
Xie, B., Wang, X., Zhu, M., Zhang, Z. & Hong, Z. CalS7 encodes a callose synthase responsible for callose deposition in the phloem. Plant J. 65, 1–14 (2011).
Langlet, O. Two hundred years of genecology. Taxon 20, 653–722 (1971).
Wang, T., O'Neill, G.A. & Aitken, S. N. Integrating environmental and genetic effects to predict responses of tree populations to climate. Ecol. Appl. 20, 153–163 (2010).
Grattapaglia, D. & Resende, M.D.V. Genomic selection in forest tree breeding. Tree Genet. Genomes 7, 241–255 (2011).
Vanholme, B. et al. Breeding with rare defective alleles (BRDA): a natural Populus nigra HCT mutant with modified lignin as a case study. New Phytol. 198, 765–776 (2013).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly (Austin) 6, 80–92 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Geraldes, A. et al. A 34K SNP genotyping array for Populus trichocarpa: design, application to the study of natural populations and transferability to other Populus species. Mol. Ecol. Resour. 13, 306–323 (2013).
Huang, X. et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 42, 961–967 (2010).
Jiao, Y. et al. Genome-wide genetic changes during modern breeding of maize. Nat. Genet. 44, 812–815 (2012).
Eckenwalder, J.E. in Biology of Populus and its Implications for Management and Conservation (eds. Stettler, R.F. et al.) 7–32 (NRC Research Press, 1996).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Patterson, N., Price, A.L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Yang, J., Lee, S.H., Goddard, M.E. & Visscher, P.M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Goudet, J. & Büchi, L. The effects of dominance, regular inbreeding and sampling design on Q(ST), an estimator of population differentiation for quantitative traits. Genetics 172, 1337–1347 (2006).
Wang, T., Hamann, A., Spittlehouse, D.L. & Murdock, T.Q. ClimateWNA—high-resolution spatial climate data for western North America. J. Appl. Meteorol. Climatol. 51, 16–29 (2012).
Charlesworth, B. Measures of divergence between populations and the effect of forces that reduce variability. Mol. Biol. Evol. 15, 538–543 (1998).
Johnson, P.L.F. & Slatkin, M. Accounting for bias from sequencing error in population genetic estimates. Mol. Biol. Evol. 25, 199–206 (2008).
Sabeti, P.C., Reich, D. & Higgins, J. Detecting recent positive selection in the human genome from haplotype structure. Nature 419, 832–837 (2002).
Delaneau, O., Zagury, J.-F. & Marchini, J. Improved whole-chromosome phasing for disease and population genetic studies. Nat. Methods 10, 5–6 (2013).
Browning, S.R. & Browning, B.L. Haplotype phasing: existing methods and new developments. Nat. Rev. Genet. 12, 703–714 (2011).
Browning, B.L. & Browning, S.R. A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am. J. Hum. Genet. 84, 210–223 (2009).
Pyhäjärvi, T., Hufford, M.B., Mezmouk, S. & Ross-Ibarra, J. Complex patterns of local adaptation in teosinte. Genome Biol. Evol. 5, 1594–1609 (2013).
Zhang, H. et al. PlantTFDB 2.0: update and improvement of the comprehensive plant transcription factor database. Nucleic Acids Res. 39, D1114–D1117 (2011).
Price, A.L., Zaitlen, N.A., Reich, D. & Patterson, N. New approaches to population stratification in genome-wide association studies. Nat. Rev. Genet. 11, 459–463 (2010).
Dudbridge, F. & Gusnanto, A. Estimation of significance thresholds for genomewide association scans. Genet. Epidemiol. 32, 227–234 (2008).
Acknowledgements
We thank the members of BioEnergy Science Center for their varied contributions to this work, and especially those involved in the collection, propagation and maintenance of the common gardens, including G. Howe, A. Groover, R. Stettler, J. Johnson and the staff at Mt. Jefferson Farms and Greenwood Resources. We thank the West Virginia University High Performance Computing facility, in particular N. Gregg and M. Carlise. P. balsamifera transcriptomes were provided by M. Olson (Texas Tech University). This work was supported by funding from the BioEnergy Science Center, a US Department of Energy (DOE) Bioenergy Research Center supported by the Office of Biological and Environmental Research in the DOE Office of Science. A.M.B. acknowledges support from the Virginia Agricultural Experiment Station and the McIntire Stennis Program of the National Institute of Food and Agriculture, US Department of Agriculture.
Author information
Authors and Affiliations
Contributions
G.A.T., S.P.D., G.T.S. and L.M.E. conceived and designed the study. All authors performed measurements. L.G., J.M. and W.S. performed sequencing. L.M.E., S.P.D., G.T.S., E. R.-M., J.M., P.R., W.M. and W.S. performed analyses. L.M.E., S.P.D. and A.M.B. drafted the manuscript. All authors read, revised, and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–18 and Supplementary Note. (PDF 12507 kb)
Supplementary Table 13
List of SNPs and their associated P values for phenotypic associations as determined by the emmax mixed model analysis. The nearest predicted gene is listed, as well as annotations of predicted SNP effects for those that fall within genes, as determined by the SNPeff program. When a SNP lies within two or more genes, additional distance to gene and gene model columns are added before the snpEFF annotation column. (XLSX 5856 kb)
Rights and permissions
About this article
Cite this article
Evans, L., Slavov, G., Rodgers-Melnick, E. et al. Population genomics of Populus trichocarpa identifies signatures of selection and adaptive trait associations. Nat Genet 46, 1089–1096 (2014). https://doi.org/10.1038/ng.3075
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3075
This article is cited by
-
Signatures of local adaptation to current and future climate in phenology-related genes in natural populations of Quercus robur
BMC Genomics (2024)
-
Genotyping SNPs in lignin biosynthesis gene (CAD1) and transcription factors (MYB1 and MYB2) exhibits association with wood density in teak (Tectona grandis L.f.)
Molecular Biology Reports (2024)
-
An integrated metagenomic, metabolomic and transcriptomic survey of Populus across genotypes and environments
Scientific Data (2024)
-
Rapid screening of secondary aromatic metabolites in Populus trichocarpa leaves
Biotechnology for Biofuels and Bioproducts (2023)
-
Molecular mechanisms of adaptive evolution in wild animals and plants
Science China Life Sciences (2023)