Abstract
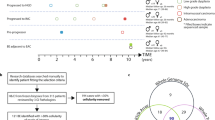
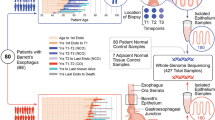
Cancer genome sequencing studies have identified numerous driver genes, but the relative timing of mutations in carcinogenesis remains unclear. The gradual progression from premalignant Barrett's esophagus to esophageal adenocarcinoma (EAC) provides an ideal model to study the ordering of somatic mutations. We identified recurrently mutated genes and assessed clonal structure using whole-genome sequencing and amplicon resequencing of 112 EACs. We next screened a cohort of 109 biopsies from 2 key transition points in the development of malignancy: benign metaplastic never-dysplastic Barrett's esophagus (NDBE; n = 66) and high-grade dysplasia (HGD; n = 43). Unexpectedly, the majority of recurrently mutated genes in EAC were also mutated in NDBE. Only TP53 and SMAD4 mutations occurred in a stage-specific manner, confined to HGD and EAC, respectively. Finally, we applied this knowledge to identify high-risk Barrett's esophagus in a new non-endoscopic test. In conclusion, mutations in EAC driver genes generally occur exceptionally early in disease development with profound implications for diagnostic and therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chin, L., Andersen, J.N. & Futreal, P.A. Cancer genomics: from discovery science to personalized medicine. Nat. Med. 17, 297–303 (2011).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Jones, S. et al. Comparative lesion sequencing provides insights into tumor evolution. Proc. Natl. Acad. Sci. USA 105, 4283–4288 (2008).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Vogelstein, B. et al. Genetic alterations during colorectal-tumor development. N. Engl. J. Med. 319, 525–532 (1988).
Goh, X.Y. et al. Integrative analysis of array–comparative genomic hybridisation and matched gene expression profiling data reveals novel genes with prognostic significance in oesophageal adenocarcinoma. Gut 60, 1317–1326 (2011).
Quante, M. et al. Bile acid and inflammation activate gastric cardia stem cells in a mouse model of Barrett-like metaplasia. Cancer Cell 21, 36–51 (2012).
Greaves, M. & Maley, C.C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Varghese, S., Lao-Sirieix, P. & Fitzgerald, R.C. Identification and clinical implementation of biomarkers for Barrett's esophagus. Gastroenterology 142, 435–441 (2012).
Dulak, A.M. et al. Gastrointestinal adenocarcinomas of the esophagus, stomach, and colon exhibit distinct patterns of genome instability and oncogenesis. Cancer Res. 72, 4383–4393 (2012).
Dulak, A.M. et al. Exome and whole-genome sequencing of esophageal adenocarcinoma identifies recurrent driver events and mutational complexity. Nat. Genet. 45, 478–486 (2013).
Agrawal, N. et al. Comparative genomic analysis of esophageal adenocarcinoma and squamous cell carcinoma. Cancer Discov. 2, 899–905 (2012).
Corley, D.A. et al. Impact of endoscopic surveillance on mortality from Barrett's esophagus–associated esophageal adenocarcinomas. Gastroenterology 145, 312–319 (2013).
Shaheen, N.J. & Hur, C. Garlic, silver bullets, and surveillance upper endoscopy for Barrett's esophagus. Gastroenterology 145, 273–276 (2013).
Hayes, D.F. et al. Breaking a vicious cycle. Sci. Transl. Med. 5, 196cm6 (2013).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Fujimoto, A. et al. Whole-genome sequencing of liver cancers identifies etiological influences on mutation patterns and recurrent mutations in chromatin regulators. Nat. Genet. 44, 760–764 (2012).
Bass, A.J. et al. Genomic sequencing of colorectal adenocarcinomas identifies a recurrent VTI1A-TCF7L2 fusion. Nat. Genet. 43, 964–968 (2011).
Streppel, M.M. et al. Next-generation sequencing of endoscopic biopsies identifies ARID1A as a tumor-suppressor gene in Barrett's esophagus. Oncogene 33, 347–357 (2014).
Curvers, W.L. et al. Low-grade dysplasia in Barrett's esophagus: overdiagnosed and underestimated. Am. J. Gastroenterol. 105, 1523–1530 (2010).
Wang, K. et al. Exome sequencing identifies frequent mutation of ARID1A in molecular subtypes of gastric cancer. Nat. Genet. 43, 1219–1223 (2011).
Jones, S. et al. Frequent mutations of chromatin remodeling gene ARID1A in ovarian clear cell carcinoma. Science 330, 228–231 (2010).
Reid, B.J., Li, X., Galipeau, P.C. & Vaughan, T.L. Barrett's oesophagus and oesophageal adenocarcinoma: time for a new synthesis. Nat. Rev. Cancer 10, 87–101 (2010).
Kadri, S.R. et al. Acceptability and accuracy of a non-endoscopic screening test for Barrett's oesophagus in primary care: cohort study. Br. Med. J. 341, c4372 (2010).
Lao-Sirieix, P. et al. Non-endoscopic screening biomarkers for Barrett's oesophagus: from microarray analysis to the clinic. Gut 58, 1451–1459 (2009).
Forshew, T. et al. Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci. Transl. Med. 4, 136ra68 (2012).
Dawson, S.J. et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N. Engl. J. Med. 368, 1199–1209 (2013).
Theisen, J. et al. Preoperative chemotherapy unmasks underlying Barrett's mucosa in patients with adenocarcinoma of the distal esophagus. Surg. Endosc. 16, 671–673 (2002).
Bhat, S. et al. Risk of malignant progression in Barrett's esophagus patients: results from a large population-based study. J. Natl. Cancer Inst. 103, 1049–1057 (2011).
Nowell, P.C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Mutter, G.L. et al. Molecular identification of latent precancers in histologically normal endometrium. Cancer Res. 61, 4311–4314 (2001).
Kinde, I. et al. Evaluation of DNA from the Papanicolaou test to detect ovarian and endometrial cancers. Sci. Transl. Med. 5, 167ra4 (2013).
Maley, C.C. et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat. Genet. 38, 468–473 (2006).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Larson, D.E. et al. SomaticSniper: identification of somatic point mutations in whole genome sequencing data. Bioinformatics 28, 311–317 (2012).
Koboldt, D.C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Ye, K., Schulz, M.H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Shah, S.P. et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486, 395–399 (2012).
Acknowledgements
Whole-genome sequencing of EAC is part of the International Cancer Genome Consortium (ICGC) through the Esophageal Cancer Clinical and Molecular Stratification (OCCAMS) Consortium and is funded by Cancer Research UK. We thank the ICGC members for their input on verification standards as part of the benchmarking exercise. Cytosponge samples were collected as part of the Cancer Research UK–funded BEST2 trial. We thank M. Griffin, L. Lovat and K. Ragunath for their contribution to Cytosponge collection. The MRC developed the Cytosponge and also funded laboratory work through a program grant to R.C.F. J.M.J.W. was funded by a Wellcome Trust Translational Medicine and Therapeutics grant. R.C.F. and C.C. are supported by additional clinical research infrastructure funding from the NHS National Institute for Health Research (NIHR), the Experimental Cancer Medicine Centre Network and the NIHR Cambridge Biomedical Research Centre. Bioinformatics work was also supported by a Cancer Research UK program grant to S.T.
We thank the Genomics Core at the Cancer Research UK Cambridge Institute for their help with processing some of the Access Array experiments as well as for running the targeted resequencing experiments. We thank the IT department at the Cancer Research UK Cambridge Institute for their support. We thank F. Marass for assistance with data analysis. We thank the Human Research Tissue Bank, supported by the NIHR Cambridge Biomedical Research Centre, from Addenbrooke's Hospital as well as the University Hospital of Southampton Trust and the Southampton Experimental Cancer Medicine Centre. We are grateful to all patients who provided written consent for participation in this study, and the staff at Addenbrooke's and the University of Southampton Tissue Bank.
Author information
Authors and Affiliations
Consortia
Contributions
R.C.F. obtained funding and conceived and supervised the study. M.D.E. undertook and supervised the development of the whole-genome analysis pipeline. M.J.D., N.S., A.G.L., M.L.S., B.C. and C.L.A. undertook development of the whole-genome analysis pipeline. S.T., P.A.W.E., N.R., M.D.E., J.M.J.W., C.S.R.-I., N.S. and A.G.L. designed various aspects of the study. J.M.J.W., C.S.R.-I., T.F., M.B. and P.L.-S. extracted the samples and performed the molecular analyses. M.O. performed histopathological diagnosis. T.J.U., N.G., R.H., C.-A.J.O. and L.S. identified and collected samples. J.M.J.W., C.S.R.-I., N.S., A.G.L., M.M., M.J.D., M.L.S., C.L.A., B.C. and M.D.E. analyzed the data. J.M.J.W. performed the analysis of mutational context. A.P.M. designed the Fluidigm primers. J.D. developed the clinical database. C.C., A.O. and S.A. developed the strategy for and performed the verification experiments. J.M.J.W., C.S.R.-I., N.S., A.G.L. and R.C.F. wrote the manuscript. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.C.F. developed the Cytosponge technology with MRC-Technology, through whom devices were provided for research. The technology has recently been licensed to Covidien; R.C.F. has no direct financial relationship with Covidien.
Integrated supplementary information
Supplementary Figure 1 Mutational patterns in the 22 discovery cohort EACs.
The bar graph represents the total number of each possible mutation class change. The heat map displays the enrichment of each mutation class at any given trinucleotide context. For example, the strong red strip running down the center of the figure represents enrichment for C:G>T:A mutations at the XpCpG trinucleotide. Enrichment at the CTT trinucleotide can be seen for all three classes of thymidine mutation in the majority of samples.
Supplementary Figure 2 ARID1A DNA mutations and protein expression.
(a) Schematic showing ARID1A mutations identified in the discovery and validation cohort (112 EAC cases). (b) Examples of two EAC tumors, one staining positively for ARID1A protein expression and one staining negatively for ARID1A protein expression.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–4 and Supplementary Tables 1–17 (PDF 2931 kb)
Rights and permissions
About this article
Cite this article
Weaver, J., Ross-Innes, C., Shannon, N. et al. Ordering of mutations in preinvasive disease stages of esophageal carcinogenesis. Nat Genet 46, 837–843 (2014). https://doi.org/10.1038/ng.3013
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3013
This article is cited by
-
Mutational signature dynamics shaping the evolution of oesophageal adenocarcinoma
Nature Communications (2023)
-
Targeted genetic and epigenetic profiling of esophageal adenocarcinomas and non-dysplastic Barrett’s esophagus
Clinical Epigenetics (2022)
-
Comprehensive RNA dataset of tissue and plasma from patients with esophageal cancer or precursor lesions
Scientific Data (2022)
-
Spatiotemporal dynamics of clonal selection and diversification in normal endometrial epithelium
Nature Communications (2022)
-
Genetic and epigenetic associations of ANRIL with coronary artery disease and risk factors
BMC Medical Genomics (2021)