Abstract
Glioma, the most common central nervous system cancer in adults, has poor prognosis. Here we identify a new SNP associated with glioma risk, rs1920116 (near TERC), that reached genome-wide significance (Pcombined = 8.3 × 10−9) in a meta-analysis of genome-wide association studies (GWAS) of high-grade glioma and replication data (1,644 cases and 7,736 controls). This region has previously been associated with mean leukocyte telomere length (LTL). We therefore examined the relationship between LTL and both this new risk locus and other previously established risk loci for glioma using data from a recent GWAS of LTL (n = 37,684 individuals)1. Alleles associated with glioma risk near TERC and TERT were strongly associated with longer LTL (P = 5.5 × 10−20 and 4.4 × 10−19, respectively). In contrast, risk-associated alleles near RTEL1 were inconsistently associated with LTL, suggesting the presence of distinct causal alleles. No other risk loci for glioma were associated with LTL. The identification of risk alleles for glioma near TERC and TERT that also associate with telomere length implicates telomerase in gliomagenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Codd, V. et al. Identification of seven loci affecting mean telomere length and their association with disease. Nat. Genet. 45, 422–427 (2013).
Stupp, R. et al. Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol. 10, 459–466 (2009).
Sanson, M. et al. Chromosome 7p11.2 (EGFR) variation influences glioma risk. Hum. Mol. Genet. 20, 2897–2904 (2011).
Shete, S. et al. Genome-wide association study identifies five susceptibility loci for glioma. Nat. Genet. 41, 899–904 (2009).
Stacey, S.N. et al. A germline variant in the TP53 polyadenylation signal confers cancer susceptibility. Nat. Genet. 43, 1098–1103 (2011).
Wrensch, M. et al. Variants in the CDKN2B and RTEL1 regions are associated with high-grade glioma susceptibility. Nat. Genet. 41, 905–908 (2009).
Jenkins, R.B. et al. A low-frequency variant at 8q24.21 is strongly associated with risk of oligodendroglial tumors and astrocytomas with IDH1 or IDH2 mutation. Nat. Genet. 44, 1122–1125 (2012).
Killela, P.J. et al. TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc. Natl. Acad. Sci. USA 110, 6021–6026 (2013).
Horn, S. et al. TERT promoter mutations in familial and sporadic melanoma. Science 339, 959–961 (2013).
Huang, F.W. et al. Highly recurrent TERT promoter mutations in human melanoma. Science 339, 957–959 (2013).
Walsh, K.M. et al. Analysis of 60 reported glioma risk SNPs replicates published GWAS findings but fails to replicate associations from published candidate-gene studies. Genet. Epidemiol. 37, 222–228 (2013).
Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Walsh, K.M. et al. Genetic variants in telomerase-related genes are associated with an older age at diagnosis in glioma patients: evidence for distinct pathways of gliomagenesis. Neuro-oncol. 15, 1041–1047 (2013).
Melin, B.S., Nordfjall, K., Andersson, U. & Roos, G. hTERT cancer risk genotypes are associated with telomere length. Genet. Epidemiol. 36, 368–372 (2012).
Jones, A.M. et al. TERC polymorphisms are associated both with susceptibility to colorectal cancer and with longer telomeres. Gut 61, 248–254 (2012).
Bojesen, S.E. et al. Multiple independent variants at the TERT locus are associated with telomere length and risks of breast and ovarian cancer. Nat. Genet. 45, 371–384 (2013).
Vannier, J.B. et al. RTEL1 is a replisome-associated helicase that promotes telomere and genome-wide replication. Science 342, 239–242 (2013).
Chang, S., Khoo, C.M., Naylor, M.L., Maser, R.S. & DePinho, R.A. Telomere-based crisis: functional differences between telomerase activation and ALT in tumor progression. Genes Dev. 17, 88–100 (2003).
Vasa-Nicotera, M. et al. Mapping of a major locus that determines telomere length in humans. Am. J. Hum. Genet. 76, 147–151 (2005).
Fitzpatrick, A.L. et al. Leukocyte telomere length and cardiovascular disease in the Cardiovascular Health Study. Am. J. Epidemiol. 165, 14–21 (2007).
Codd, V. et al. Common variants near TERC are associated with mean telomere length. Nat. Genet. 42, 197–199 (2010).
Denham, J. et al. Longer leukocyte telomeres are associated with ultra-endurance exercise independent of cardiovascular risk factors. PLoS ONE 8, e69377 (2013).
Brouilette, S.W. et al. Telomere length, risk of coronary heart disease, and statin treatment in the West of Scotland Primary Prevention Study: a nested case-control study. Lancet 369, 107–114 (2007).
Ma, H. et al. Shortened telomere length is associated with increased risk of cancer: a meta-analysis. PLoS ONE 6, e20466 (2011).
Wentzensen, I.M., Mirabello, L., Pfeiffer, R.M. & Savage, S.A. The association of telomere length and cancer: a meta-analysis. Cancer Epidemiol. Biomarkers Prev. 20, 1238–1250 (2011).
Hou, L., Zhang, X., Gawron, A.J. & Liu, J. Surrogate tissue telomere length and cancer risk: shorter or longer? Cancer Lett. 319, 130–135 (2012).
Wilson, W.R. et al. Blood leucocyte telomere DNA content predicts vascular telomere DNA content in humans with and without vascular disease. Eur. Heart J. 29, 2689–2694 (2008).
Okuda, K. et al. Telomere length in the newborn. Pediatr. Res. 52, 377–381 (2002).
Romano, G.H. et al. Environmental stresses disrupt telomere length homeostasis. PLoS Genet. 9, e1003721 (2013).
Houlston, R.S. et al. Meta-analysis of three genome-wide association studies identifies susceptibility loci for colorectal cancer at 1q41, 3q26.2, 12q13.13 and 20q13.33. Nat. Genet. 42, 973–977 (2010).
Fingerlin, T.E. et al. Genome-wide association study identifies multiple susceptibility loci for pulmonary fibrosis. Nat. Genet. 45, 613–620 (2013).
Samani, N.J. & van der Harst, P. Biological ageing and cardiovascular disease. Heart 94, 537–539 (2008).
Cawthon, R.M. Telomere length measurement by a novel monochrome multiplex quantitative PCR method. Nucleic Acids Res. 37, e21 (2009).
Cawthon, R.M. Telomere measurement by quantitative PCR. Nucleic Acids Res. 30, e47 (2002).
Hansen, H.M., Wiemels, J.L., Wrensch, M. & Wiencke, J.K. DNA quantification of whole genome amplified samples for genotyping on a multiplexed bead array platform. Cancer Epidemiol. Biomarkers Prev. 16, 1686–1690 (2007).
Howie, B.N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Liu, J.Z. et al. Meta-analysis and imputation refines the association of 15q25 with smoking quantity. Nat. Genet. 42, 436–440 (2010).
Higgins, J.P., Thompson, S.G., Deeks, J.J. & Altman, D.G. Measuring inconsistency in meta-analyses. Br. Med. J. 327, 557–560 (2003).
Pe'er, I., Yelensky, R., Altshuler, D. & Daly, M.J. Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet. Epidemiol. 32, 381–385 (2008).
Ward, L.D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2012).
Boyle, A.P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22, 1790–1797 (2012).
Cooper, G.M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005).
Stranger, B.E. et al. Patterns of cis regulatory variation in diverse human populations. PLoS Genet. 8, e1002639 (2012).
Yang, T.P. et al. Genevar: a database and Java application for the analysis and visualization of SNP-gene associations in eQTL studies. Bioinformatics 26, 2474–2476 (2010).
Acknowledgements
Work at UCSF was supported by the US National Institutes of Health (grants R25CA112355, R01CA52689, P50CA097257, R01CA126831 and R01CA139020), as well as by the National Brain Tumor Foundation, the UCSF Lewis Chair in Brain Tumor Research, the UCSF Robert Magnin Newman chair in Neuro-Oncology and donations from the families and friends of John Berardi, Helen Glaser, Elvera Olsen, Raymond E. Cooper and William Martinusen. Work at the Mayo Clinic was supported by the US National Institutes of Health (grants P50CA108961 and P30CA15083), the National Institute of Neurological Disorders and Stroke (grant RC1NS068222Z), the Bernie and Edith Waterman Foundation, and the Ting Tsung and Wei Fong Chao Family Foundation. Work at the University of Leicester was undertaken under the European Union Framework Programme 7 ENGAGE Project (HEALTH-F4-2007-201413). V.C. and N.J.S. are supported by the British Heart Foundation.
This project was supported by the National Center for Research Resources and the National Center for Advancing Translational Sciences, US National Institutes of Health, through UCSF Clinical and Translational Science Institute grant UL1RR024131. Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the US National Institutes of Health.
The collection of cancer incidence data used in this study was supported by the California Department of Public Health as part of the statewide cancer reporting program mandated by California Health and Safety Code Section 103885; by the National Cancer Institute's Surveillance, Epidemiology and End Results Program under contract HHSN261201000140C awarded to the Cancer Prevention Institute of California, contract HHSN261201000035C awarded to the University of Southern California and contract HHSN261201000034C awarded to the Public Health Institute; and by the Centers for Disease Control and Prevention National Program of Cancer Registries, under agreement U58DP003862-01 awarded to the California Department of Public Health. The ideas and opinions expressed herein are those of the authors, and endorsement by the State of California Department of Public Health, the National Cancer Institute and the Centers for Disease Control and Prevention or their contractors and subcontractors is not intended nor should be inferred.
The results published here are in part based on data generated by TCGA managed by the National Cancer Institute and the National Human Genome Research Institute. Information about TCGA can be found at http://cancergenome.nih.gov/. This study makes use of data generated by WTCCC. A full list of the investigators who contributed to the generation of the data is available from http://www.wtccc.org.uk/. Funding for the project was provided by the Wellcome Trust under awards 076113 and 085475.
Author information
Authors and Affiliations
Consortia
Contributions
K.M.W., M.R.W. and J.K.W. led the study at UCSF, R.B.J. led the study at the Mayo Clinic, and N.J.S. led the study at the University of Leicester. K.M.W., V.C., R.B.J., M.R.W., M.P. and T.R. contributed to manuscript preparation. Study coordination was the responsibility of T.K. at the Mayo Clinic and T.R. and L.S.M. at UCSF. K.M.W. and V.C. codirected and conducted biostatistics and bioinformatics analyses with additional support from P.A.D., J.E.E.-P., M.L.K., A.M.M., P.M.B., T.R., H.S., A.R.P., I.V.S., P.v.d.H. and the ENGAGE Consortium Telomere Group. Laboratory work was performed by T.K. under the direction of R.B.J. at the Mayo Clinic and by H.M.H., S.Z. and B.S.C. under the direction of J.K.W. and J.L.W. at UCSF. Pathology support was provided by T.T. Subject enrollment or clinical record review was performed or facilitated by M.D.P., S.M.C., M.S.B., B.P.O., D.H.L. and P.v.d.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
A full list of members and affiliations appears in the Supplementary Note.
Integrated supplementary information
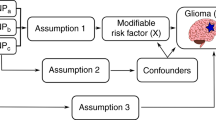
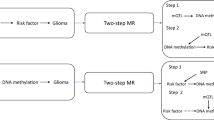
Supplementary Figure 1 Summary of study design and analysis.
The flowchart details the flow of analyses through the four stages of the study.
Supplementary Figure 2 Cumulative effect of three telomere-associated glioma risk SNPs (rs1920116, TERC; rs2736100, TERT; rs6010620, RTEL1).
(a) Plot of the increasing ORs for high-grade glioma with increasing numbers of risk alleles (range of 0–6). The ORs are relative to the median number of 4 risk alleles. Vertical bars correspond to 95% confidence intervals. Odds ratios increase in a montonic fashion. (b) Distribution of telomere-associated glioma risk alleles in high-grade glioma cases (dark gray bars) and controls (light gray bars).
Supplementary Figure 3 Changes in the magnitude of high-grade glioma risk associated with telomere-associated SNPs across subject age strata in analyses of UCSF AGS cases and controls.
Odds ratios for glioma were calculated in case-control analyses, adjusted for sex. The y axis is represented on a log scale (base 2), ranging from 0.50 to 2.0. Vertical bars correspond to 95% confidence intervals.
Supplementary Figure 4 SNP association plots for high-grade glioma risk (top) and mean leukocyte telomere length (bottom) at 3q26.2, 5p15.33 and 20q13.33.
The strength of linkage disequilibrium between each SNP and the high-grade glioma top hit (purple circle) is indicated by color. Recombination rates, plotted in light blue, are based on 1000 Genomes CEU samples. Black vertical bars in Supplementary Figure 4c mark the location of the RTEL1-PCNA interaction motif (PIP box).
Supplementary Figure 5 Odds ratios near TERC for high-grade glioma risk, plotted with –log10 (P values) for glioma risk and mean leukocyte telomere length.
Glioma odds ratios for SNPs with P values of <0.05 are plotted in black and correspond to the risk associated with each additional copy of the glioma risk allele. Odds ratios range from 1.11 to 1.26. –log10 (P values) for SNPs associated with high-grade glioma in the discovery stage (P < 0.05) appear in green, and –log10 (P values) for SNPs associated with LTL appear in pink. A horizontal black line indicates a –log10 (P value) of 2, corresponding to a P value of 0.01.
Supplementary Figure 6 SNP association plots for high-grade glioma risk at 3q26.2, conditioned on lead glioma SNP rs1920116 (top) and lead LTL SNP rs10936599 (bottom).
The strength of linkage disequilibrium between each SNP and rs1920116 is indicated by color. Recombination rates, plotted in light blue, are based on 1000 Genomes CEU samples.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–4 and Supplementary Figures 1–6 (PDF 3029 kb)
Source data
Rights and permissions
About this article
Cite this article
Walsh, K., Codd, V., Smirnov, I. et al. Variants near TERT and TERC influencing telomere length are associated with high-grade glioma risk. Nat Genet 46, 731–735 (2014). https://doi.org/10.1038/ng.3004
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3004
This article is cited by
-
Identification of h-TERT Promoter Mutations in Germline DNA from North Indian Lung Carcinoma Patients
Indian Journal of Clinical Biochemistry (2023)
-
Implementation of individualised polygenic risk score analysis: a test case of a family of four
BMC Medical Genomics (2022)
-
Mesenchymal stem cell-derived extracellular vesicles prevent glioma by blocking M2 polarization of macrophages through a miR-744-5p/TGFB1-dependent mechanism
Cell Biology and Toxicology (2022)
-
Telomere length is not a main factor for the development of islet autoimmunity and type 1 diabetes in the TEDDY study
Scientific Reports (2022)
-
Glioma exosomal microRNA-148a-3p promotes tumor angiogenesis through activating the EGFR/MAPK signaling pathway via inhibiting ERRFI1
Cancer Cell International (2020)