Abstract
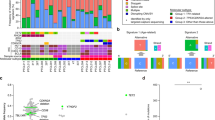
Peripheral T cell lymphomas (PTCLs) are a heterogeneous and poorly understood group of non-Hodgkin lymphomas1,2. Here we combined whole-exome sequencing of 12 tumor-normal DNA pairs, RNA sequencing analysis and targeted deep sequencing to identify new genetic alterations in PTCL transformation. These analyses identified highly recurrent epigenetic factor mutations in TET2, DNMT3A and IDH2 as well as a new highly prevalent RHOA mutation encoding a p.Gly17Val alteration present in 22 of 35 (67%) angioimmunoblastic T cell lymphoma (AITL) samples and in 8 of 44 (18%) PTCL, not otherwise specified (PTCL-NOS) samples. Mechanistically, the RHOA Gly17Val protein interferes with RHOA signaling in biochemical and cellular assays, an effect potentially mediated by the sequestration of activated guanine-exchange factor (GEF) proteins. In addition, we describe new and recurrent, albeit less frequent, genetic defects including mutations in FYN, ATM, B2M and CD58 implicating SRC signaling, impaired DNA damage response and escape from immune surveillance mechanisms in the pathogenesis of PTCL.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Armitage, J.O. The aggressive peripheral T-cell lymphomas: 2012 update on diagnosis, risk stratification, and management. Am. J. Hematol. 87, 511–519 (2012).
Rüdiger, T. et al. Peripheral T-cell lymphoma (excluding anaplastic large-cell lymphoma): results from the Non-Hodgkin's Lymphoma Classification Project. Ann. Oncol. 13, 140–149 (2002).
Schiller, M.R. Coupling receptor tyrosine kinases to Rho GTPases–GEFs what's the link. Cell. Signal. 18, 1834–1843 (2006).
Bar-Sagi, D. & Hall, A. Ras and Rho GTPases: a family reunion. Cell 103, 227–238 (2000).
Vega, F.M. & Ridley, A.J. Rho GTPases in cancer cell biology. FEBS Lett. 582, 2093–2101 (2008).
Hanna, S. & El-Sibai, M. Signaling networks of Rho GTPases in cell motility. Cell. Signal. 25, 1955–1961 (2013).
Hall, A. Rho family GTPases. Biochem. Soc. Trans. 40, 1378–1382 (2012).
Longenecker, K. et al. Structure of a constitutively activated RhoA mutant (Q63L) at 1.55 Å resolution. Acta Crystallogr. D Biol. Crystallogr. 59, 876–880 (2003).
Mayer, T., Meyer, M., Janning, A., Schiedel, A.C. & Barnekow, A. A mutant form of the rho protein can restore stress fibers and adhesion plaques in v-src transformed fibroblasts. Oncogene 18, 2117–2128 (1999).
Zhang, S. et al. Rho family GTPases regulate p38 mitogen-activated protein kinase through the downstream mediator Pak1. J. Biol. Chem. 270, 23934–23936 (1995).
Ghosh, P.M. et al. Role of RhoA activation in the growth and morphology of a murine prostate tumor cell line. Oncogene 18, 4120–4130 (1999).
Pan, Z.K. et al. Role of the Rho GTPase in bradykinin-stimulated nuclear factor–κB activation and IL-1β gene expression in cultured human epithelial cells. J. Immunol. 160, 3038–3045 (1998).
Reid, T. et al. Rhotekin, a new putative target for Rho bearing homology to a serine/threonine kinase, PKN, and rhophilin in the rho-binding domain. J. Biol. Chem. 271, 13556–13560 (1996).
García-Mata, R. et al. Analysis of activated GAPs and GEFs in cell lysates. Methods Enzymol. 406, 425–437 (2006).
Couronné, L., Bastard, C. & Bernard, O.A. TET2 and DNMT3A mutations in human T-cell lymphoma. N. Engl. J. Med. 366, 95–96 (2012).
Quivoron, C. et al. TET2 inactivation results in pleiotropic hematopoietic abnormalities in mouse and is a recurrent event during human lymphomagenesis. Cancer Cell 20, 25–38 (2011).
Cairns, R.A. et al. IDH2 mutations are frequent in angioimmunoblastic T-cell lymphoma. Blood 119, 1901–1903 (2012).
Palacios, E.H. & Weiss, A. Function of the Src-family kinases, Lck and Fyn, in T-cell development and activation. Oncogene 23, 7990–8000 (2004).
McCormack, P.L. & Keam, S.J. Dasatinib: a review of its use in the treatment of chronic myeloid leukaemia and Philadelphia chromosome–positive acute lymphoblastic leukaemia. Drugs 71, 1771–1795 (2011).
Harr, M.W. et al. Inhibition of Lck enhances glucocorticoid sensitivity and apoptosis in lymphoid cell lines and in chronic lymphocytic leukemia. Cell Death Differ. 17, 1381–1391 (2010).
Widmann, C., Gerwins, P., Johnson, N.L., Jarpe, M.B. & Johnson, G.L. MEK kinase 1, a substrate for DEVD-directed caspases, is involved in genotoxin-induced apoptosis. Mol. Cell. Biol. 18, 2416–2429 (1998).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Schmitz, R. et al. Burkitt lymphoma pathogenesis and therapeutic targets from structural and functional genomics. Nature 490, 116–120 (2012).
Maher, C.A. et al. Chimeric transcript discovery by paired-end transcriptome sequencing. Proc. Natl. Acad. Sci. USA 106, 12353–12358 (2009).
McPherson, A. et al. deFuse: an algorithm for gene fusion discovery in tumor RNA-Seq data. PLoS Comput. Biol. 7, e1001138 (2011).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Roy, A., Kucukural, A. & Zhang, Y. I-TASSER: a unified platform for automated protein structure and function prediction. Nat. Protoc. 5, 725–738 (2010).
Worth, C.L., Preissner, R. & Blundell, T.L. SDM—a server for predicting effects of mutations on protein stability and malfunction. Nucleic Acids Res. 39, W215–W222 (2011).
Subauste, M.C. et al. Rho family proteins modulate rapid apoptosis induced by cytotoxic T lymphocytes and Fas. J. Biol. Chem. 275, 9725–9733 (2000).
Mariotti, A. et al. EGF-R signaling through Fyn kinase disrupts the function of integrin α6β4 at hemidesmosomes: role in epithelial cell migration and carcinoma invasion. J. Cell Biol. 155, 447–458 (2001).
Kamanova, J. et al. Adenylate cyclase toxin subverts phagocyte function by RhoA inhibition and unproductive ruffling. J. Immunol. 181, 5587–5597 (2008).
Pallotta, M.T. et al. Indoleamine 2,3-dioxygenase is a signaling protein in long-term tolerance by dendritic cells. Nat. Immunol. 12, 870–878 (2011).
Schenk, S. et al. Sirt1 enhances skeletal muscle insulin sensitivity in mice during caloric restriction. J. Clin. Invest. 121, 4281–4288 (2011).
Wang, Q. et al. Thrombin and lysophosphatidic acid receptors utilize distinct rhoGEFs in prostate cancer cells. J. Biol. Chem. 279, 28831–28834 (2004).
Acknowledgements
We would like to thank V. Ribrag (Institut Gustave Roussy) for providing a PTCL sample and R. Parsons (Mount Sinai School of Medicine) for assistance in the setup of RHOA GTP-loading assays. This work was supported by a Leukemia and Lymphoma Society Translational Research Grant (A.A.F.), a Herbert Irving Comprehensive Cancer Center interprogrammatic pilot project grant (A.A.F. and R.R.), a US National Institutes of Health Ruth L. Kirschstein National Research Service Award (1F30CA174099 to J.E.H.), a grant from the Ligue Nationale Contre le Cancer (O.A.B.) and an Institut National du Cancer (INCa)/–Direction de l'Hospitalisation et de l'Organisation des soins (DHOS) translational research grant (O.A.B.). L.C. is funded by ITMO (Institut Multi-Organismes Cancer) and INCa. M.-Y.K. is funded by the Leukemia and Lymphoma Society. E.C. is supported by the Spanish Ministry of Science and Innovation (SAF2012-38432) and Institucio Catalana de Recerca i Estudis Avancats.
Author information
Authors and Affiliations
Contributions
T.P. directed and supervised mutation analysis, performed functional assays and wrote the manuscript. L.C. performed validation and recurrence mutation analysis and functional assays. H.K. performed exome and RNA-seq mutation analysis. M.-Y.K. performed peptide binding assays. A.A.-I. analyzed Illumina deep sequencing amplicon sequencing data. A.P.-G. performed functional assays. Z.C. performed structure-function analysis and analyzed Illumina sequence data. F.A. performed RNA-seq mutation analysis and fusion oncogene identification. M.A. performed validation analysis of Illumina sequencing results. J.E.H. performed functional assays. X.J. performed functional assays. I.S.L. contributed clinical samples and supervised functional assays. C.N., M.B., C.B., G.B., M.A.P. and E.C. contributed clinical samples. O.A.B. designed the study, contributed samples and supervised research. R.R. directed and supervised the analysis of Illumina sequencing data. A.A.F. designed the study, directed and supervised research and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 and Supplementary Tables 1, 2, 4 and 6–8 (PDF 918 kb)
Supplementary Tables 3, 5 and 9
Supplementary Tables 3, 5 and 9 (XLSX 69 kb)
Rights and permissions
About this article
Cite this article
Palomero, T., Couronné, L., Khiabanian, H. et al. Recurrent mutations in epigenetic regulators, RHOA and FYN kinase in peripheral T cell lymphomas. Nat Genet 46, 166–170 (2014). https://doi.org/10.1038/ng.2873
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2873
This article is cited by
-
Therapeutic challenges in peripheral T-cell lymphoma
Molecular Cancer (2024)
-
DNMT3AR882H accelerates angioimmunoblastic T-cell lymphoma in mice
Oncogene (2023)
-
Updates in the Classification of T-cell Lymphomas and Lymphoproliferative Disorders
Current Hematologic Malignancy Reports (2023)
-
Advances in the pathogenesis and therapeutic strategies of angioimmunoblastic T-cell lymphoma
Clinical and Experimental Medicine (2023)
-
Follicular helper T-cell lymphomas: disease spectrum, relationship with clonal hematopoiesis, and mimics. A report of the 2022 EA4HP/SH lymphoma workshop
Virchows Archiv (2023)