Abstract
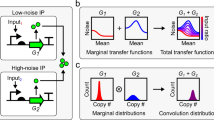
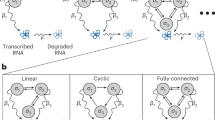
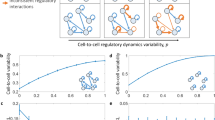
Gene regulatory interactions are context dependent, active in some cellular states but not in others. Stochastic fluctuations, or 'noise', in gene expression propagate through active, but not inactive, regulatory links1,2. Thus, correlations in gene expression noise could provide a noninvasive means to probe the activity states of regulatory links. However, global, 'extrinsic', noise sources generate correlations even without direct regulatory links. Here we show that single-cell time-lapse microscopy, by revealing time lags due to regulation, can discriminate between active regulatory connections and extrinsic noise. We demonstrate this principle mathematically, using stochastic modeling, and experimentally, using simple synthetic gene circuits. We then use this approach to analyze dynamic noise correlations in the galactose metabolism genes of Escherichia coli. We find that the CRP-GalS-GalE feed-forward loop is inactive in standard conditions but can become active in a GalR mutant. These results show how noise can help analyze the context dependence of regulatory interactions in endogenous gene circuits.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pedraza, J.M. & van Oudenaarden, A. Noise propagation in gene networks. Science 307, 1965–1969 (2005).
Rosenfeld, N. et al. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005).
Toledo, F. & Wahl, G.M. Regulating the p53 pathway: in vitro hypothesis, in vivo veritas. Nat. Rev. Cancer 6, 909–923 (2006).
Piggot, P.J. & Hilbert, D.W. Sporulation of Bacillus subtilis. Curr. Opin. Microbiol. 7, 579–586 (2004).
Suel, G.M. et al. An excitable gene regulatory circuit induces transient cellular differentiation. Nature 440, 545–550 (2006).
Elowitz, M.B. et al. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002).
Raser, J.M. & O'Shea, E.K. Noise in gene expression: origins, consequences, and control. Science 309, 2010–2013 (2005).
Kaern, M. et al. Stochasticity in gene expression: from theories to phenotypes. Nat. Rev. Genet. 6, 451–464 (2005).
Paulsson, J. Summing up the noise in gene networks. Nature 427, 415–418 (2004).
Sigal, A. et al. Variability and memory of protein levels in human cells. Nature 444, 643–646 (2006).
Rosenfeld, N., Elowitz, M.B. & Alon, U. Negative autoregulation speeds the response times of transcription networks. J. Mol. Biol. 323, 785–793 (2002).
Arkin, A.P. & Ross, J. Statistical construction of chemical-reaction mechanisms from measured time-series. J. Phys. Chem. 99, 970–979 (1995).
Arkin, A.P., Shen, P. & Ross, J. A test case of correlation metric construction of a reaction pathway from measurements. Science 277, 1275–1279 (1997).
Gillespie, D.T. Exact numerical simulation of the Ornstein-Uhlenbeck process and its integral. Phys. Rev. E Stat. Phys. Plasmas Fluids Relat. Interdiscip. Topics 54, 2084–2091 (1996).
Meyer, B.J., Maurer, R. & Ptashne, M. Gene regulation at the right operator (OR) of bacteriophage lambda. II. OR1, OR2, and OR3: their roles in mediating the effects of repressor and cro. J. Mol. Biol. 139, 163–194 (1980).
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1–I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997).
Shen-Orr, S.S., Milo, R., Mangan, S. & Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nat. Genet. 31, 64–68 (2002).
Mangan, S. et al. The incoherent feed-forward loop accelerates the response-time of the gal system of Escherichia coli. J. Mol. Biol. 356, 1073–1081 (2006).
Kaplan, S. et al. The incoherent feed-forward loop can generate non-monotonic input functions for genes. Mol. Syst. Biol. 4, 203 (2008).
Semsey, S. et al. Signal integration in the galactose network of Escherichia coli. Mol. Microbiol. 65, 465–476 (2007).
Cox, C.D. et al. Using noise to probe and characterize gene circuits. Proc. Natl. Acad. Sci. USA 105, 10809–10814 (2008).
Blake, W.J. et al. Phenotypic consequences of promoter-mediated transcriptional noise. Mol. Cell 24, 853–865 (2006).
Maamar, H., Raj, A. & Dubnau, D. Noise in gene expression determines cell fate in Bacillus subtilis. Science 317, 526–529 (2007).
Arkin, A., Ross, J. & McAdams, H.H. Stochastic kinetic analysis of developmental pathway bifurcation in phage lambda-infected Escherichia coli cells. Genetics 149, 1633–1648 (1998).
Tsang, J. & van Oudenaarden, A. Exciting fluctuations: monitoring competence induction dynamics at the single-cell level. Mol. Syst. Biol. 2, 2006.0025 (2006).
Megason, S.G. & Fraser, S.E. Imaging in systems biology. Cell 130, 784–795 (2007).
Yu, D. et al. An efficient recombination system for chromosome engineering in Escherichia coli. Proc. Natl. Acad. Sci. USA 97, 5978–5983 (2000).
Zaslaver, A. et al. A comprehensive library of fluorescent transcriptional reporters for Escherichia coli. Nat. Methods 3, 623–628 (2006).
Datsenko, K.A. & Wanner, B.L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640–6645 (2000).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Acknowledgements
We thank M. Fontes, F. Tan, L. Cai, E. Franco, and all members of the Elowitz and Murray groups for their feedback and suggestions. H. Garcia provided advice on the chromosomal integration and gene knockout experiments. We thank J. Garcia-Ojalvo, U. Alon, R. Kishony, N. Rosenfeld and B. Shraiman for discussions. M.J.D. and R.M.M. are supported by the Institute for Collaborative Biotechnologies through grant DAAD19-03-D-0004 from the US Army Research Office. M.J.D. was additionally supported by a Department of Energy Computational Science Graduate Fellowship. This research was supported by US National Institutes of Health grants R01GM079771, P50 GM068763, National Science Foundation CAREER Award 0644463 and the Packard Foundation.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1 and Supplementary Note (PDF 157 kb)
Supplementary Movie 1
Movie of YFP (false colored in green) and RFP (red) for the chromosomally integrated synthetic circuit. Frames are spaced at a 10 minute interval. Selected frames from this movie are shown in Fig. 3B of the text. (AVI 264 kb)
Supplementary Movie 2
Movie of YFP and CFP for the chromosomally integrated synthetic circuit. (AVI 265 kb)
Rights and permissions
About this article
Cite this article
Dunlop, M., Cox, R., Levine, J. et al. Regulatory activity revealed by dynamic correlations in gene expression noise. Nat Genet 40, 1493–1498 (2008). https://doi.org/10.1038/ng.281
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.281
This article is cited by
-
Dynamic fluctuations in a bacterial metabolic network
Nature Communications (2023)
-
Role of integrated noise in pathway-specific signal propagation in feed-forward loops
Theory in Biosciences (2021)
-
Research progress and the biotechnological applications of multienzyme complex
Applied Microbiology and Biotechnology (2021)
-
Corruption of the Pearson correlation coefficient by measurement error and its estimation, bias, and correction under different error models
Scientific Reports (2020)
-
DynamicME: dynamic simulation and refinement of integrated models of metabolism and protein expression
BMC Systems Biology (2019)