Abstract
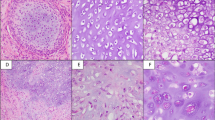
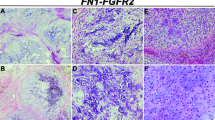
Chondrosarcoma is a heterogeneous collection of malignant bone tumors and is the second most common primary malignancy of bone after osteosarcoma. Recent work has identified frequent, recurrent mutations in IDH1 or IDH2 in nearly half of central chondrosarcomas. However, there has been little systematic genomic analysis of this tumor type, and, thus, the contribution of other genes is unclear. Here we report comprehensive genomic analyses of 49 individuals with chondrosarcoma (cases). We identified hypermutability of the major cartilage collagen gene COL2A1, with insertions, deletions and rearrangements identified in 37% of cases. The patterns of mutation were consistent with selection for variants likely to impair normal collagen biosynthesis. In addition, we identified mutations in IDH1 or IDH2 (59%), TP53 (20%), the RB1 pathway (33%) and Hedgehog signaling (18%).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Stephens, P.J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–404 (2012).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl. Acad. Sci. USA 107, 16910–16915 (2010).
Kim, N. & Jinks-Robertson, S. Transcription as a source of genome instability. Nat. Rev. Genet. 13, 204–214 (2012).
Barbieri, O. et al. Depletion of cartilage collagen fibrils in mice carrying a dominant negative Col2a1 transgene affects chondrocyte differentiation. Am. J. Physiol. Cell Physiol. 285, C1504–C1512 (2003).
Yang, G. et al. PTEN deficiency causes dyschondroplasia in mice by enhanced hypoxia-inducible factor 1α signaling and endoplasmic reticulum stress. Development 135, 3587–3597 (2008).
Chiu, L.H. et al. Differential effect of ECM molecules on re-expression of cartilaginous markers in near quiescent human chondrocytes. J. Cell Physiol. 226, 1981–1988 (2011).
Amary, M.F. et al. IDH1 and IDH2 mutations are frequent events in central chondrosarcoma and central and periosteal chondromas but not in other mesenchymal tumours. J. Pathol. 224, 334–343 (2011).
Popovici-Muller, J. et al. Discovery of the first potent inhibitors of mutant IDH1 that lower tumor 2-HG in vivo. ACS Med. Chem. Lett. 3, 850–855 (2012).
Fathi, A.T. et al. Prospective serial evaluation of 2-hydroxyglutarate, during treatment of newly diagnosed acute myeloid leukemia, to assess disease activity and therapeutic response. Blood 120, 4649–4652 (2012).
Hallor, K.H. et al. Genomic profiling of chondrosarcoma: chromosomal patterns in central and peripheral tumors. Clin. Cancer Res. 15, 2685–2694 (2009).
Schrage, Y.M. et al. Central chondrosarcoma progression is associated with pRb pathway alterations: CDK4 down-regulation and p16 overexpression inhibit cell growth in vitro. J. Cell Mol. Med. 13, 2843–2852 (2009).
Aceto, G. et al. Mutations of APC and MYH in unrelated Italian patients with adenomatous polyposis coli. Hum. Mutat. 26, 394 (2005).
Ahn, J. et al. Cloning of the putative tumour suppressor gene for hereditary multiple exostoses (EXT1). Nat. Genet. 11, 137–143 (1995).
Hopyan, S. et al. A mutant PTH/PTHrP type I receptor in enchondromatosis. Nat. Genet. 30, 306–310 (2002).
Stickens, D. et al. The EXT2 multiple exostoses gene defines a family of putative tumour suppressor genes. Nat. Genet. 14, 25–32 (1996).
Bruce, S.J. et al. Inactivation of Patched1 in the mouse limb has novel inhibitory effects on the chondrogenic program. J. Biol. Chem. 285, 27967–27981 (2010).
Wuelling, M. & Vortkamp, A. Transcriptional networks controlling chondrocyte proliferation and differentiation during endochondral ossification. Pediatr. Nephrol. 25, 625–631 (2010).
Tang, J.Y. et al. Inhibiting the hedgehog pathway in patients with the basal-cell nevus syndrome. N. Engl. J. Med. 366, 2180–2188 (2012).
Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, 539–542 (2011).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Stephens, P.J. et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 144, 27–40 (2011).
Ye, K., Schulz, M.H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Shalaby, A. et al. The role of epidermal growth factor receptor in chordoma pathogenesis: a potential therapeutic target. J. Pathol. 223, 336–346 (2011).
McLaren, W. et al. Deriving the consequences of genomic variants with the Ensembl API and SNP Effect Predictor. Bioinformatics 26, 2069–2070 (2010).
Acknowledgements
We are grateful to the patients for participating in the research and to the clinicians and support staff of the London Sarcoma Service involved in their care. This work was supported by funding from the Wellcome Trust (grant 077012/Z/05/Z) and the Skeletal Cancer Action Trust (SCAT), UK. Material was obtained from the Royal National Orthopaedic Hospital Musculoskeletal Research Program and Biobank. Support was provided to A.M.F. by the National Institute for Health Research, the University College London Hospital Biomedical Research Centre and the UCL Experimental Cancer Centre. P.J.C. is personally funded through a Wellcome Trust Senior Clinical Research Fellowship (grant WT088340MA). P.V.L. is a postdoctoral researcher of the Research Foundation–Flanders (FWO). S.B. is funded through the Wellcome Trust PhD Programme for Clinicians.
Author information
Authors and Affiliations
Contributions
P.S.T. and S.B. analyzed the sequence data. S.L.C. and J.M.C.T. performed rearrangement analysis. P.V.L. analyzed the SNP6 array data. D.C.W. performed statistical analyses. S. McLaren, D.H. and S.O. coordinated sample processing and technical investigations. S. Martin coordinated sample acquisition and processing. C.H., C.L., L.M. and M.M. performed technical investigations. A.B., J.G., J.H., M.M.J., A.J., D.J., A.M., J.M., K.R., J.W.T., H.D., S.N.-Z. and D.B. performed informatic investigations. F.A., R.T., N.P. and A.M.F. provided samples and clinical data and performed immunohistochemistry. P.J.C., M.R.S., A.M.F. and P.A.F. directed the research and contributed to the manuscript. P.A.F. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1 and 3–6 (PDF 11884 kb)
Supplementary Table 2
Variants identified in the primary whole exome screen and in the extension series (Excel file) (XLSX 156 kb)
Supplementary Table 7
Copy number changes identified in 49 chondrosarcomas (Excel file) (XLSX 30 kb)
Rights and permissions
About this article
Cite this article
Tarpey, P., Behjati, S., Cooke, S. et al. Frequent mutation of the major cartilage collagen gene COL2A1 in chondrosarcoma. Nat Genet 45, 923–926 (2013). https://doi.org/10.1038/ng.2668
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2668
This article is cited by
-
IDH mutations in G2-3 conventional central bone chondrosarcoma: a mono institutional experience
BMC Cancer (2023)
-
Identification of gene profiles related to the development of oral cancer using a deep learning technique
BMC Medical Genomics (2023)
-
A genetic model for central chondrosarcoma evolution correlates with patient outcome
Genome Medicine (2022)
-
Mutation in XPO5 causes adult-onset autosomal dominant familial focal segmental glomerulosclerosis
Human Genomics (2022)
-
Somatic mutations in collagens are associated with a distinct tumor environment and overall survival in gastric cancer
BMC Cancer (2022)