Abstract
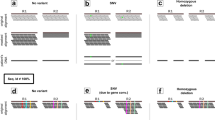
Although genetic lesions responsible for some mendelian disorders can be rapidly discovered through massively parallel sequencing of whole genomes or exomes, not all diseases readily yield to such efforts. We describe the illustrative case of the simple mendelian disorder medullary cystic kidney disease type 1 (MCKD1), mapped more than a decade ago to a 2-Mb region on chromosome 1. Ultimately, only by cloning, capillary sequencing and de novo assembly did we find that each of six families with MCKD1 harbors an equivalent but apparently independently arising mutation in sequence markedly under-represented in massively parallel sequencing data: the insertion of a single cytosine in one copy (but a different copy in each family) of the repeat unit comprising the extremely long (∼1.5–5 kb), GC-rich (>80%) coding variable-number tandem repeat (VNTR) sequence in the MUC1 gene encoding mucin 1. These results provide a cautionary tale about the challenges in identifying the genes responsible for mendelian, let alone more complex, disorders through massively parallel sequencing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bleyer, A.J., Hart, P.S. & Kmoch, S. Hereditary interstitial kidney disease. Semin. Nephrol. 30, 366–373 (2010).
Castro, A.F. & Coresh, J. CKD surveillance using laboratory data from the population-based National Health and Nutrition Examination Survey (NHANES). Am. J. Kidney Dis. 53, S46–S55 (2009).
Christodoulou, K. et al. Chromosome 1 localization of a gene for autosomal dominant medullary cystic kidney disease. Hum. Mol. Genet. 7, 905–911 (1998).
Wolf, M.T.F. et al. Medullary cystic kidney disease type 1: mutational analysis in 37 genes based on haplotype sharing. Hum. Genet. 119, 649–658 (2006).
Fuchshuber, A. et al. Refinement of the gene locus for autosomal dominant medullary cystic kidney disease type 1 (MCKD1) and construction of a physical and partial transcriptional map of the region. Genomics 72, 278–284 (2001).
Kiser, R.L. et al. Medullary cystic kidney disease type 1 in a large Native-American kindred. Am. J. Kidney Dis. 44, 611–617 (2004).
Wolf, M.T. et al. Telomeric refinement of the MCKD1 locus on chromosome 1q21. Kidney Int. 66, 580–585 (2004).
Choi, M. et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc. Natl. Acad. Sci. USA 106, 19096–19101 (2009).
Al-Romaih, K.I. et al. Genetic diagnosis in consanguineous families with kidney disease by homozygosity mapping coupled with whole-exome sequencing. Am. J. Kidney Dis. 58, 186–195 (2011).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Gemayel, R., Vinces, M.D., Legendre, M. & Verstrepen, K.J. Variable tandem repeats accelerate evolution of coding and regulatory sequences. Annu. Rev. Genet. 44, 445–477 (2010).
Legendre, M., Pochet, N., Pak, T. & Verstrepen, K.J. Sequence-based estimation of minisatellite and microsatellite repeat variability. Genome Res. 17, 1787–1796 (2007).
Horne, A.W. et al. MUC 1: a genetic susceptibility to infertility? Lancet 357, 1336–1337 (2001).
Fowler, J.C., Teixeira, A.S., Vinall, L.E. & Swallow, D.M. Hypervariability of the membrane-associated mucin and cancer marker MUC1. Hum. Genet. 113, 473–479 (2003).
Brayman, M., Thathiah, A. & Carson, D.D. MUC1: a multifunctional cell surface component of reproductive tissue epithelia. Reprod. Biol. Endocrinol. 2, 4 (2004).
Levitin, F. et al. The MUC1 SEA module is a self-cleaving domain. J. Biol. Chem. 280, 33374–33386 (2005).
Auranen, M., Ala-Mello, S., Turunen, J.A. & Järvelä, I. Further evidence for linkage of autosomal-dominant medullary cystic kidney disease on chromosome 1q21. Kidney Int. 60, 1225–1232 (2001).
Spicer, A.P., Rowse, G.J., Lidner, T.K. & Gendler, S.J. Delayed mammary tumor progression in Muc-1 null mice. J. Biol. Chem. 270, 30093–30101 (1995).
Lander, E. & Kruglyak, L. Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat. Genet. 11, 241–247 (1995).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Abecasis, G.R., Cherny, S.S., Cookson, W.O. & Cardonl, L.R. Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nat. Genet. 30, 97–101 (2002).
Korn, J.M. et al. Integrated genotype calling and association analysis of SNPs, common copy number polymorphisms and rare CNVs. Nat. Genet. 40, 1253–1260 (2008).
Handsaker, R.E., Korn, J.M., Nemesh, J. & McCarroll, S.A. Discovery and genotyping of genome structural polymorphism by sequencing on a population scale. Nat. Genet. 43, 269–276 (2011).
Acknowledgements
We thank T.L. Hatte for reagent use. We thank D. Altshuler, T. Carter and J. Schlondorff for useful discussions and M. Cortes, M. Ilzarbe and M. Betancourt for helpful project management. We also thank F. Letendre, M. Coole, R.P. Frere, C. Bonnet, L. Mulrain, N. Norbui and H. Arachchi for Sanger sequencing. This work was conducted as part of the Slim Initiative for Genomic Medicine, a joint United States–Mexico project funded by the Carlos Slim Health Institute. This research was supported in part by the Intramural Research Program of the US NIH, National Human Genome Research Institute (NHGRI). S.K., H.H., J.S. and V.B. were funded by Charles University programs PRVOUK-P24/LF1/3 and UNCE 204011, and their work was supported by grants LH12015 and NT13116-4/2012 from the Ministry of Education and the Ministry of Health of the Czech Republic. S.L.A. was supported by US NIH grant DK34854 (The Harvard Digestive Diseases Center). N.P. is a Broad Fellow of the Broad Institute and a postdoctoral research fellow of the Fund for Scientific Research–Flanders (FWO Vlaanderen, Belgium). I.G.-V. was supported by the Human Frontier Science Program, Alon, the Israeli Centers of Research Excellence (I-CORE) and the Edmond J. Safra Center for Bioinformatics at Tel Aviv University.
Author information
Authors and Affiliations
Contributions
A.J.B., E.S.L. and M.J.D. jointly supervised the research. R.J.X., M.R.P. and S.L.A. provided study design and interpretation advice. C.A., S.J.S., P.S.H. and A.J.B. performed sample collection. C. Stevens managed the project. C. Sougnez and K.C. provided early genotyping and sequencing support. Linkage analysis was performed by A.K. on the basis of previous work by P.S.H. A.K. and M.J.D. developed variation discovery and analysis methods. A.K., J.T.R. and R.E.H. analyzed structural variation. T.G. performed CNV analysis. S.G. supervised the sequencing. S. Sigurdsson and K.L.-T. designed the custom capture array. M.P. performed direct PCR of the polymorphic VNTR candidates selected by N.P. A.G. and D.A. performed Southern blot and long-range PCR of the MUC1 VNTR. C.N. supervised the MUC1 VNTR sequencing approach. A.G. performed VNTR allele cloning and generation of sequencing libraries. E.K., R.D., D.P. and S. Steelman performed Sanger sequencing. D.B.J. assembled and analyzed VNTR Sanger sequencing. M.G. provided RNA-seq support. S.K. supervised the immunohistochemistry and immunofluorescence work performed by V.B., H.H., J.S. and P.V. A.K., B.B. and M.D. developed the C-insertion genotype assay. M.C.Z. provided informatic and sequencing consultation. A.R. provided informatic and analysis consultation. C.Y., J.T.R., M.N.C., I.G.-V., R.E.H. and E.R. provided informatic support. The manuscript was written primarily by A.K., A.G., A.J.B., E.S.L. and M.J.D. The supplementary information was prepared mainly by A.K., A.G., D.B.J., B.B., R.E.H., S. Sigurdsson, S.K. and A.J.B.
Corresponding authors
Ethics declarations
Competing interests
A.K., A.G., B.B. and M.D. are listed as inventors on the C-insertion genotyping assay under patent review, filed by the Broad Institute. The other authors declare no competing interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–5 and Supplementary Tables 1–3 (PDF 306 kb)
Rights and permissions
About this article
Cite this article
Kirby, A., Gnirke, A., Jaffe, D. et al. Mutations causing medullary cystic kidney disease type 1 lie in a large VNTR in MUC1 missed by massively parallel sequencing. Nat Genet 45, 299–303 (2013). https://doi.org/10.1038/ng.2543
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2543
This article is cited by
-
Var∣Decrypt: a novel and user-friendly tool to explore and prioritize variants in whole-exome sequencing data
Epigenetics & Chromatin (2023)
-
Prevalence of hereditary tubulointerstitial kidney diseases in the German Chronic Kidney Disease study
European Journal of Human Genetics (2022)
-
Autosomal dominant tubulointerstitial kidney disease: more than just HNF1β
Pediatric Nephrology (2022)
-
The causes and consequences of paediatric kidney disease on adult nephrology care
Pediatric Nephrology (2022)
-
Translating Organoids into Artificial Kidneys
Current Transplantation Reports (2022)