Abstract
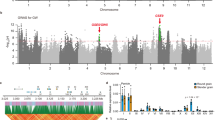
The transition from the prostrate growth of ancestral wild rice (O. rufipogon Griff.) to the erect growth of Oryza sativa cultivars was one of the most critical events in rice domestication. This evolutionary step importantly improved plant architecture and increased grain yield. Here we find that prostrate growth of wild rice from Yuanjiang County in China is controlled by a semi-dominant gene, PROG1 (PROSTRATE GROWTH 1), on chromosome 7 that encodes a single Cys2-His2 zinc-finger protein. prog1 variants identified in O. sativa disrupt the prog1 function and inactivate prog1 expression, leading to erect growth, greater grain number and higher grain yield in cultivated rice. Sequence comparison shows that 182 varieties of cultivated rice, including 87 indica and 95 japonica cultivars from 17 countries, carry identical mutations in the prog1 coding region that may have become fixed during rice domestication.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Doebley, J., Gaut, B. & Smith, B. The molecular genetics of crop domestication. Cell 127, 1309–1321 (2006).
Khush, G. Origin, dispersal, cultivation and variation of rice. Plant Mol. Biol. 35, 25–34 (1997).
Kovach, M., Sweeney, M. & McCouch, S. New insights into the history of rice domestication. Trends Genet. 23, 578–587 (2007).
Li, C., Zhou, A. & Sang, T. Rice domestication by reducing shattering. Science 311, 1936–1939 (2006).
Lin, Z. et al. Origin of seed shattering in rice (O. sativa L.). Planta 226, 11–20 (2007).
Konishi, S. et al. An SNP caused loss of seed shattering during rice domestication. Science 312, 1392–1396 (2006).
Sweeney, M. et al. Caught red-handed: Rc encodes a basic helix-loop-helix protein conditioning red pericarp in rice. Plant Cell 18, 283–294 (2006).
Tan, L. et al. Development of Oryza rufipogon and Oryza sativa introgression lines and assessment for yield-related quantitative trait loci. J. Integr. Plant Biol. 49, 871–884 (2007).
Yuan, Q. et al. The TIGR rice genome annotation resource: annotating the rice genome and creating resources for plant biologists. Nucleic Acids Res. 31, 229–233 (2003).
Takatsuji, H. Zinc-finger proteins: the classical zinc finger emerges in contemporary plant science. Plant Mol. Biol. 39, 1073–1078 (1999).
Hiratsu, K., Ohta, M., Matsui, K. & Ohme-Takagi, M. The SUPPERMAN protein is an active repressor whose carboxy-terminal repression domain is required for the development of normal flowers. FEBS Lett. 514, 351–354 (2002).
Yoshihara, T. & Iino, M. Identification of the gravitropism-related rice gene LAZY1 and elucidation of LAZY1-dependent and –independent gravity signaling pathways. Plant Cell Physiol. 48, 678–688 (2007).
Li, P. et al. LAZY1 controls rice shoot gravitropism through regulating poplar auxin transport. Cell Res. 17, 402–410 (2007).
Yu, B. et al. TAC1, a major quantitative trait locus controlling tiller angle in rice. Plant J. 52, 891–898 (2007).
Waisel, Y. Evolution of erect growth forms in domesticated wheats: possible effects of grazing. Oecologia 73, 630–632 (1987).
Doebley, J., Stec, A. & Hubbard, L. The evolution of apical dominance in maize. Nature 386, 485–488 (1997).
Clark, R., Wagler, T., Quijada, P. & Doebley, J. A distant upstream enhancer at the maize domestication gene tb1 has pleiotropic effects on plant and inflorescent architecture. Nat. Genet. 38, 594–597 (2006).
Li, X. et al. Construction of a bacterial artificial chromosome (BAC) library of common wild rice (Oryza rufipogon Griff.) for map-based cloning of genes selected during the domestication of rice. Biotechnol. Lett. 30, 555–561 (2008).
Scarpella, E., Rueb, S. & Meijer, A.H. The RADICLELESS1 gene is a required for vascular pattern formation in rice. Development 130, 645–658 (2003).
Thompson, J., Gibson, T., Plewniak, F., Jeanmougin, F. & Higgins, D. The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882 (1997).
Crooks, G., Hon, G., Chandonia, J. & Brenner, S. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Acknowledgements
We thank N. Harberd for comments on the manuscript, Y. Wang for assisting in cross sections, and the International Rice Research Institute, Chinese Rice Research Institute, Institute of Crop Sciences of Chinese Academy of Agricultural Sciences, Guangxi Academy of Agricultural Sciences and Guangdong Academy of Agricultural Sciences for providing the wild rice and cultivated rice samples. This research was supported by the National Natural Science Foundation, the Ministry of Agriculture and the Ministry of Science and Technology of China.
Author information
Authors and Affiliations
Contributions
L.T. conducted characterization of introgression line, genetic mapping, gene cloning and gene expression analysis; X.L. constructed the genomic BAC library of YJCWR and screened the positive clone; F.L. helped L.T. with genetic mapping and gene expression analysis; X.S. maintained plant materials and performed the field management; C.L. performed the genetic transformation; Z.Z., Y.F., H.C. and X.W. conducted the collection of rice garmplasm and phenotypic data; C.S and D.X. designed and supervised this study; C.S., D.X. and L.T. analyzed the data and wrote the paper.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–4, Supplementary Figures 1–4 (PDF 551 kb)
Rights and permissions
About this article
Cite this article
Tan, L., Li, X., Liu, F. et al. Control of a key transition from prostrate to erect growth in rice domestication. Nat Genet 40, 1360–1364 (2008). https://doi.org/10.1038/ng.197
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.197
This article is cited by
-
Evolutionary and synteny analysis of HIS1, BADH2, GBSS1, and GBSS2 in rice: insights for effective introgression breeding strategies
Scientific Reports (2024)
-
A mediator of OsbZIP46 deactivation and degradation negatively regulates seed dormancy in rice
Nature Communications (2024)
-
Different contributions of PROG1 and TAC1 to the angular kinematics of the main culm and tillers of wild rice (Oryza rufipogon)
Planta (2024)
-
Substitution Mapping and Allelic Variations of the Domestication Genes from O. rufipogon and O. nivara
Rice (2023)
-
Can the Wild Perennial, Rhizomatous Rice Species Oryza longistaminata be a Candidate for De Novo Domestication?
Rice (2023)