Abstract
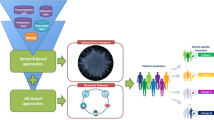
By using systems pathology, it might be possible to provide a predictive, personalized therapeutic recommendation for patients with prostate cancer. Systems pathology integrates quantitative data and information from many sources to generate a reliable prediction of the expected natural course of the disease and response to different therapeutic options. In other words, through the integration of relatively large data sets and the use of knowledge engineering, systems pathology aims at predicting the future behavior of tumors and their interaction with the host. In this Review, we introduce the methods used in systems pathology and summarize a recent study providing the first evidence of a concept for this strategy. The results show that systems pathology can provide a personalized prediction of the risk of recurrence after prostatectomy for cancer.
Key Points
-
Systems pathology integrates quantitative data from diverse sources to render a personalized predictive and prognostic report
-
Systems pathology combines conventional clinicopathologic information with quantitative morphology and molecular data
-
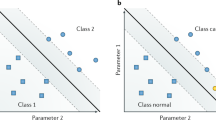
Systems pathology uses modern machine learning techniques to mine and interpret the data sets that produce a predictive personalized report
-
Prostate cancer has been chosen to provide the first proof that systems pathology can be used in patient care
-
Initial results show that it is possible to use systems pathology to provide a personalized prediction of the risk of recurrence after prostatectomy in men with prostate cancer
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Albertsen PC et al. (1998) Competing risk analysis of men aged 55 to74 years at diagnosis managed conservatively for clinically localized prostate cancer. JAMA 280: 975–980
Bianco FJ Jr et al. (2003) Ten-year survival after radical prostatectomy: specimen Gleason score is the predictor in organ-confined prostate cancer. Clin Prostate Cancer 1: 242–247
Allan RW et al. (2003) Correlation of minute (0.5 mm or less) focus of prostate adenocarcinoma on needle biopsy with radical prostatectomy specimen: role of prostate specific antigen density. J Urol 170: 370–372
Quinn DI et al. (2005) Molecular markers of prostate cancer outcome. European J Cancer 41: 858–887
Diamond J et al. (2004) The use of morphological characteristics and texture analysis in the identification of tissue composition in prostatic neoplasia. Hum Pathol 35: 1121–1131
Roula MA et al. (2002) A multispectral computer vision system for automatic grading of prostatic neoplasia. In Proceedings of the IEEE International Symposium on Biomedical Imaging, July 7–10 2002, Washington, DC, 193–196
Stotzka R et al. (1995) A hybrid neural and statistical classifier system for histopathologic grading of prostate lesions. Anal Quant Cytol Histol 17: 204–218
Smith Y et al. (1999) Similarity measurement method for the classification of architecturally differentiated images. Comp Biomed Res 32: 1–12
Wetzel AW et al. (1999) Evaluation of prostate tumor grades by content-based image retrieval. In 27th AIPR Workshop: Advances in Computer-Assisted Recognition, 14 October 1998, Washington, DC, 3584: 244–252 (Ed. Merickso RJ)
Jafari-Khouzani K and Soltanian-Zadeh H (2003) Multiwavelet grading of pathological images of prostate. IEEE Trans Biomed Eng 50: 697–704
Vapnik VN (1995) The Nature of Statistical Learning Theory. New York: Springer–Verlag
Su AI et al. (2001) Molecular classification of human carcinomas by use of gene expression signatures. Cancer Res 61: 7388–7393
Yeang CH et al. (2001) Molecular classification of multiple tumor types. Bioinformatics 17 (Suppl 1): S316–S322
Cristianini N and Shawe-Taylor J. (2000) An Introduction to Support Vector Machines and Other Kernel-Based Learning Methods. Cambridge, UK: Cambridge University Press
Ye QH et al. (2003) Predicting hepatitis B virus-positive metastatic hepatocellular carcinomas using gene expression profiling and supervised machine learning. Nat Med 9: 416–423
Brown MPS et al. (2000) Knowledgebased analysis of microarray gene expression data using support vector machines. Proc Natl Acad Sci USA 97: 262–267
Biganzoli E et al. (1998) Feed forward neural networks for the analysis of censored survival data: a partial logistic regression approach. Stat Med 17: 1169–1186
Ohno-Machado L and Musen MA. (1997) Modular neural networks for medical prognosis: quantifying the benefits of combining neural networks for survival prediction. Connect Sci 9: 71–86
Kattan MW et al. (1998) Experiments to determine whether recursive partitioning or an artifi-cial neural network overcomes theoretical limitation of cox proportional hazards regression. Comput Biomed Res 31: 363–373
Brown SF et al. (1997) On the use of artificial neural networks for the analysis of survival data. IEEE Trans Neural Netw 8: 1071–1077
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Oliver Saidi is a consultant at Aureon Laboratories Inc. and is listed as inventor of IP created at Aureon.
Carols Cordon-Cardo is on the Board of Directors of Aureon Laboratories Inc.
Jose Costa is on the Board of Directors of Aureon Laboratories Inc.
Rights and permissions
About this article
Cite this article
Saidi, O., Cordon-Cardo, C. & Costa, J. Technology Insight: will systems pathology replace the pathologist?. Nat Rev Urol 4, 39–45 (2007). https://doi.org/10.1038/ncpuro0669
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ncpuro0669
This article is cited by
-
Multiple immunofluorescence assay identifies upregulation of Active β-catenin in prostate cancer
BMC Research Notes (2019)
-
Systems pathology—or how to solve the complex problem of predictive pathology
Virchows Archiv (2008)