Abstract
Peripherally derived regulatory T (pTreg) cell generation requires T-cell receptor (TCR) signalling and the cytokines TGF-β1 and IL-2. Here we show that TCR signalling induces the microRNA miR-31, which negatively regulates pTreg-cell generation. miR-31 conditional deletion results in enhanced induction of pTreg cells, and decreased severity of experimental autoimmune encephalomyelitis (EAE). Unexpectedly, we identify Gprc5a as a direct target of miR-31. Gprc5a is known as retinoic acid-inducible protein 3, and its deficiency leads to impaired pTreg-cell induction and increased EAE severity. By generating miR-31 and Gprc5a double knockout mice, we show that miR-31 promotes the development of EAE through inhibiting Gprc5a. Thus, our data identify miR-31 and its target Gprc5a as critical regulators for pTreg-cell generation, suggesting a previously unrecognized epigenetic mechanism for dysfunctional Treg cells in autoimmune diseases.
Similar content being viewed by others
Introduction
T cells serve as a central cellular player in adaptive immunity, and their activation and differentiation are elicited by signals from T-cell receptor (TCR), co-stimulatory receptors and various cytokines1. Once activated by an antigen, naive CD4+ T cells proliferate and differentiate into various T helper (TH) cell subsets, including TH1, TH2, TH17 and regulatory T (Treg) cells, that release different cytokines and exhibit distinct effector functions2. Besides their critical role in driving immune responses against infections, TH1 and TH17 cells participate in the pathogenesis of autoimmune inflammatory diseases, such as experimental autoimmune encephalomyelitis (EAE)3. Moreover, naive T cells differentiate into Treg cells exhibiting immunosuppressive capacity, and the transcriptional factor FoxP3 controls their development and fucntion4,5. According to their origins, Treg cells are divided into thymus-derived Treg (tTreg) cells derived from the thymus, peripherally derived regulatory T (pTreg) cells generated out of the thymus under various inductive signals, and in vitro-induced regulatory T (iTreg) cells6,7. It is now clear that naive CD4+ T cells sorted as FoxP3− in the thymus possess full potential to differentiate into pTreg cells, thus are potential targets for therapeutic interventions for chronic inflammatory diseases7,8. Independent on thymus, pTreg cells differentiate in secondary lymphoid organs and tissues, and require TCR signalling and the cytokines TGF-β and IL-2 (ref. 7), and only a low antigen dose of a high-affinity TCR ligand is optimal to generate a persistent population of pTreg cells in vivo9.
So far, dysfunctional Treg cells are identified in several autoimmune disorders including mutiple sclerosis (MS)10,11,12. One of the failures of Treg-cell-mediated immunoregulation is inadequate numbers of Treg cells that may be due to defective induction of pTreg cells in the periphery13. Thus, understanding of molecular mechanisms underlying pTreg-cell generation might provide deeper insights into physiological and pathological immune responses in autoimmune inflammatory diseases.
MicroRNAs (miRNAs) are single-stranded, small noncoding RNAs located in introns or exons of protein-coding genes as well as in non-coding genes14. miRNAs have been implicated in maintaining immune homeostasis during stress, such as inflammation, by regulating gene expression at post-transcriptional level15. Several studies have reported that specific miRNA signatures were observed for specialized T-cell subsets, and these miRNAs are dynamically regulated during T-cell maturation16,17. Dicer and Drosha are two essential components for the generation of miRNA, and loss of these factors leads to defects in lymphocyte differentiation and autoimmune inflammation18,19. Recently, accumulating evidence has demonstrated that miRNAs are also crucial for Treg-cell development, function and stability17,18,19,20. Treg cells display a set of miRNAs that is distinct from conventional T cells21. However, intrinsic miRNAs involved in the polarization of Treg cells from naive T cells in vitro and in vivo settings are largely undetermined.
In this study, we showed that miR-31 expression was triggered by TCR signalling, and downregulated by TGF-β1-induced FoxP3. The conditional deletion of miR-31 in CD4+ T cells led to enhanced induction of pTreg cells in the periphery, and decreased severity of EAE. Retinoic acid (RA) regulates the expression of genes required for cell proliferation, differentiation and survival by binding its nuclear retinoic acid receptors (RARs) and retinoid X receptors (RXRs)22. Although RA has been shown to enforce pTreg-cell generation23, the mechanism by which RA promotes pTreg-cell induction is ill-defined. Unexpectedly, we here identified Gprc5a as a direct target of miR-31. Gprc5a is also known as retinoic acid-inducible protein 3 harbouring the functional RAR/RXR binding sites of RA in its core promoter24. Gprc5a was targeted by miR-31 through direct binding to its 3′-untranslated regions (3′-UTR), and its deficiency resulted in the impairment of pTreg-cell induction and increased EAE severity. Thus, our findings demonstrated that miR-31 negatively regulated pTreg-cell generation by targeting Gprc5a, suggesting a novel epigenetic mechanism for impaired pTreg-cell induction in autoimmunity.
Results
miR-31 expression is triggered by TCR signalling
Report of FoxP3 mRNA harbouring the target sequence of miR-31 promoted us to investigate its role in the induction and/or function of Treg cells which are vital for preventing autoimmune disease21. We induced EAE, an animal model of MS, with myelin oligodendrocyte glycoprotein peptide (MOG35–55) in mice to investigate expression pattern of miR-31 in pathogenic T cells in the tissue-specific autoimmune inflammation. miR-31 expression was assessed in splenocytes and sorted CD4+ T cells at day 10 post immunization. We found that the expression of miR-31 was significantly increased in both splenocytes and pathogenic CD4+ T cells in EAE mice compared with healthy controls (Fig. 1a). We next stimulated the TCR of naive T (CD4+CD25−CD62Lhigh) cells with plate-coated anti-CD3- and soluble anti-CD28-specific antibodies, and we detected that the miR-31 expression was increased ∼125-fold in activated CD4+ T cells compared with untreated naive T cells (Fig. 1b). Together, these data suggest that TCR signalling induces miR-31 expression in CD4+ T cells.
(a) qPCR analysis of miR-31 expression in total splenocytes and sorted CD4+ T cells from healthy controls (Ctr) or EAE mice (n=9–11) 10 days post immunization. (b) miR-31 expression in naive T cells alone or cultured with anti-CD3, anti-CD28 mAb for 3 days (n=4). (c) qPCR analysis of miR-31 expression in naive T cells and polarized TH1, TH17 and iTreg cells (n=4 per group); results are presented relative to miR-31 expression in isolated cells of control mice as in a or in naive T cells as in b,c. (d) Representative flow cytometry of FoxP3gfp expression in naive T cells cultured with anti-CD3, anti-CD28 mAb and rmIL-2 in the absence (left panel, FoxP3− T cells) or presence (right panel, FoxP3+ T cells) of TGF-β1 for 3 days. Numbers adjacent to outlined areas indicate per cent cells in each. (e) qPCR analysis of miR-31 expression in sorted FoxP3− and FoxP3+ T cells (n=4 per group). (f) The time course of miR-31 and FoxP3 expression in naive T cells activated by anti-CD3, anti-CD28 mAb and rmIL-2 in the absence or presence of TGF-β1 for 96 h (n=3 per group; results presented as in b). (g) FoxP3 was immunoprecipitated from iTreg cells. Immunoprecipitates were assayed for the expression levels of miR-31 promoter. **P<0.01, ***P<0.001, two-tailed Student’s t-test. Data are from one experiment representative of three (a–e) or two (f,g) independent experiments (mean±s.e.m.).
Because the TCR signal coordinating with lineage-specific cytokines triggers naive T cells to differentiate into specialized effector cells, we sought to examine miR-31 expression in different T-cell subsets. We differentiated naive T cells in vitro under polarizing conditions for the generation of TH1, TH17 and iTreg cells in cultures as these T-cell subsets are critical in the pathology of EAE25,26,27. At 4 days after activation, miR-31 expression was 29.5-fold higher in TH1 cells, 47.4-fold higher in TH17 cells, but there was 5.6-fold reduction in iTreg cells than that of naive T cells (Fig. 1c), which suggested a possible regulatory role for miR-31 in CD4+ T-cell lineage differentiation. Because miR-31 has been implicated to negatively regulate FoxP3 expression in human Treg cells21, we sought to investigate whether upregulation of miR-31 coincides with downregulation of FoxP3 during iTreg-cell induction. We polarized naive T cells derived from FoxP3gfp reporter mice into iTreg cells, and examined miR-31 expression in sorted CD25+FoxP3− and CD25+FoxP3+ cells. The miR-31 expression in CD25+FoxP3+ cells was ∼90-fold lower than that in CD25+FoxP3− population (Fig. 1d,e). These data demonstrate that miR-31 is preferentially diminished in iTreg cells. Although the expression of miR-31 was slightly increased in tTreg and pTreg cells compared with iTreg cells, its expression in either tTreg or pTreg was not significantly different between control and EAE mice, suggesting that Treg cells maintain baseline miR-31 expression in vivo (Supplementary Fig. 1a). To further identify why iTreg cells exhibit diminished levels of miR-31, we activated naive T cells with CD3- and CD28-specific antibodies in the absence or presence of TGF-β1, and measured the time-dependent appearance of miR-31. miR-31 abundance was gradually increased during the stimulation with CD3 and CD28 antibodies in the absence of TGF-β1, however, decreased at 12 h when FoxP3 was induced by adding TGF-β1 (Fig. 1f). Moreover, TGF-β1 dose-dependently decreased miR-31 expression in iTreg-cell differentiation (Supplementary Fig. 1b,c). Together, these data indicate that miR-31 expression might be downregulated by TGF-β1-induced FoxP3 during iTreg-cell induction in vitro. Database analysis revealed one potential FoxP3-binding site in the promoter element at −1919 upstream from the transcription start site (TSS) of mouse miR-31 (Fig. 1g upper panel). To establish the possible binding of FoxP3 to the putative binding site in the promoter element of miR-31, we carried out chromatin immunoprecipitation (ChIP) assays. These assays showed a significant recruitment of FoxP3 to the putative miR-31 promoter (Fig. 1g lower panel). Thus, our results suggest that FoxP3 possibly binds miR-31 promoter and downregulates its expression during iTreg-cell in vitro differentiation.
miR-31 conditional deletion ameliorates autoimmune disease
To determine whether miR-31 expressed by pathogenic CD4+ T cells is a functionally relevant regulator for the development of autoimmune inflammation, we used homologous recombination to generate mice with a miR-31 allele flanked by loxP sites (floxed; Fig. 2a upper panel). The germline-transmitted mice were crossed with CD4Cre transgenic mice to achieve a conditional knockout mouse model with a deleted miR-31 allele in CD4+ T cells (Fig. 2a lower panel). To verify a specific deletion of miR-31 in CD4+ T cells, we designed primers (P1 and P2) spanning the loxP sites (floxed allele, 1,195 bp; deleted allele, 474 bp) and genotyped mice using DNA of either splenocytes or sorted CD4+ T cells derived from miR-31fl/flCD4Cre (cKO) and miR-31fl/fl control mice (Fig. 2a). We detected both floxed and deleted alleles in splenocytes of cKO mice, while only a floxed allele in splenocytes of miR-31fl/fl control mice, a deleted allele in CD4+ T cells of cKO mice and a floxed allele in CD4+ T cells of miR-31fl/fl control mice (Fig. 2b). Quantitative real-time PCR (qPCR) analysis confirmed a specific miR-31 ablation in CD4+ T cells in cKO mice (Fig. 2c). These mice remained healthy without any detectable immune-mediated pathology at least for 32 weeks. By inducing EAE, we demonstrated that the specific miR-31 ablation in CD4+ T cells significantly decreased its severity accompanied by an evident prevention of weight loss in cKO mice compared with miR-31fl/fl controls (Fig. 2d,e). Moreover, the deletion of miR-31 led to a marked decrease in infiltration of inflammatory cells and demyelination in spinal cord of cKO mice with EAE (Supplementary Fig. 2a–c). Thus, using genetic approach, we clearly demonstrated the significant impact of miR-31 expressed by CD4+ T cells on the development of autoimmunity.
(a) Schematic representation of the miR-31 locus and targeting strategy. Cre-mediated recombination of loxP sites in mice. Primers (P1 and P2) spanning the loxP sites were designed for genotyping floxed allele (1,195 bp) and deleted allele (474 bp). (b) PCR products of splenocytes and sorted CD4+ T cells derived from either miR-31fl/fl control or miR-31fl/flCD4Cre (cKO) mice. (c) qPCR analysis to confirm the deletion of miR-31 in CD4+ T cells derived from cKO mice. (d,e) Clinical scores and weight loss (mean±s.e.m.) of miR-31fl/fl or cKO mice after the induction of EAE were assessed every day (n=7 per group). ***P<0.001, two-tailed Student’s t-test for c, one-way analysis of variance for d and e. Data are representative of two (c) or three (b,d and e) independent experiments (mean±s.e.m.).
miR-31 skews the CD4 T-cell-mediated immune balance
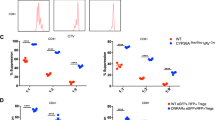
To assess how deletion of miR-31 in CD4+ T cells reduced the severity of progressive EAE, we analysed T-cell frequency and activation in non-immunized cKO mice. By flow cytometric analysis, we observed no substantial changes in T-cell numbers and activation status in the thymus in cKO mice compared with miR-31fl/fl controls (Supplementary Fig. 3a,b). T-cell proliferation in response to stimulation via TCR-CD28 was also similar in miR-31fl/fl and cKO T cells as determined by CellTrace Violet (CTV) fluorescence (Fig. 3a). These data suggest that miR-31 is dispensable for T-cell development, activation and proliferation. We next analysed Treg-cell frequency in non-immunized mice, and found that miR-31 deficiency did not change the proportion of Treg cells in the thymus and periphery, indicating that miR-31 had no impact on tTreg and pTreg-cell development (Supplementary Fig. 4a–c). TH1 and TH17 cells are inflammatory cells that develop during tissue-specific inflammatory responses and play a critical role in enhancing tissue inflammation25,27. Therefore, we investigated inflamed spleen and central nervous system (CNS) from miR-31fl/fl and cKO mice for the presence of IFN-γ-producing (TH1) and IL-17-producing (TH17) CD4+ T cells during EAE. In contrast to miR-31fl/fl controls, cKO mice showed a significant reduction of TH1 and TH17 cell proportion not only in inflamed spleen (Fig. 3b,c) but also in CNS (Fig. 3d,e) 14 days post immunization with MOG35–55. These data suggest that the development of inflammatory TH1 and TH17 cells in cKO mice is impaired during the induction phase of autoimmune disease. We next investigated whether the decreased encephalitogenic potential of CD4+ T cells in cKO mice was a consequence of increased peripheral Treg-cell generation during EAE. On day 14 post immunization, we observed a marked increase in the proportion and absolute numbers of Treg cells in the periphery of cKO mice (Fig. 3f–h). Moreover, there was no significant difference of tTreg frequency in miR-31fl/fl and cKO mice with EAE (Supplementary Fig. 4d). Together, these results demonstrate that miR-31 skews the balance between pathogenic TH1/TH17 cells and Treg cells in the periphery during autoimmune inflammation.
(a) Representative flow cytometric analysis of CTV fluorescence dilution of miR-31fl/fl or cKO naive T cells cultured with anti-CD3, anti-CD28 mAb and rmIL-2 for 3 days. Red lines indicate non-stimulated controls. (b,c) Flow cytometry of TH1 and TH17 cells in inflamed spleen of miR-31fl/fl and cKO mice 14 days after the induction of EAE. (d,e) Flow cytometry of TH1 and TH17 cells in CNS of miR-31fl/fl and cKO mice 14 days after the induction of EAE. (f,g) Flow cytometric analysis of Treg cells in the spleen of miR-31fl/fl/FoxP3gfp and cKO/FoxP3gfp mice 14 days after the induction of EAE (n=5, gated on total cells). Numbers in quadrants or adjacent to outlined areas indicate per cent cells in each. (h) Absolute numbers of Treg cells in the spleen of miR-31fl/fl/FoxP3gfp and cKO/FoxP3gfp mice 14 days after the induction of EAE (n=7). *P<0.05, **P<0.01, two-tailed Student’s t-test. Data are from one experiment representative of at least two independent experiments (mean±s.e.m.).
miR-31 limits pTreg-cell induction
Because miR-31 exhibited a distinct expression pattern in differentiated CD4+ T-cell subsets, we postulated that the intrinsic miR-31 may regulate their in vitro generation. We sorted naive T cells from miR-31fl/fl and cKO mice, and polarized them into TH1, TH17 and iTreg cells under lineage-specific conditions in vitro. After 4 days culture, we found no significant change for the differentiation of TH1 and TH17 cells from naive T cells of cKO mice compared with miR-31fl/fl control mice (Fig. 4a–d). Of note, the lack of miR-31 markedly induced iTreg-cell differentiation in culture (Fig. 4e). Thus, these data suggest that miR-31 deficiency preferentially enhanced the generation of TGF-β1-induced iTreg cells in vitro. Helios is potentially a marker, which could distinguish tTreg cells from pTreg cells28. We injected intravenously bone marrow cells (5 × 106) from either miR-31fl/fl or cKO mice into lethally irradiated C57BL/6J recipient mice to generate bone marrow chimeric mice. Eight weeks after bone marrow transplantation, EAE was induced in all chimeric mice. On day 14 post immunization, we analysed the frequency of Helios−FoxP3+ pTreg cells and Helios+FoxP3+ tTreg cells in the spleen of chimeric mice by flow cytometry. By Helios staining, we demonstrated that the conditional deletion of miR-31 led to increased numbers of Helios− pTreg cells, whereas Helios+ tTreg cells had no significant change in vivo (Supplementary Fig. 5). Together, our data indicate that miR-31 limits pTreg-cell induction in autoimmunity. We next examined the suppressive capacity of miR-31fl/fl and cKO Treg cells. CTV dilution determined that cKO Treg cells inhibited T-cell proliferation to the same extent as miR-31fl/fl Treg cells (Fig. 4f). pTreg cells had robust suppression and enhanced stability, suppressed ongoing EAE29. Given the fact that pTreg cells converted from conventional T cells play a critical role in the control of development of EAE or other autoimmune diseases8,26,30,31,32, we here provided strong evidence that promoting generation of pTreg cells by disrupting miR-31 was likely responsible for the observed phenotype.
FoxP3gfp reporter mice were crossed with either miR-31fl/fl or cKO mice. (a–d) Flow cytometry of TH1 and TH17 cells polarized from naive T cells of miR-31fl/fl/FoxP3gfp and cKO/FoxP3gfp mice (n=4–5). Numbers in quadrants indicate per cent cells in each. (e) Flow cytometric analysis of in vitro-induced iTreg cells from naive T cells of miR-31fl/fl/FoxP3gfp and cKO/FoxP3gfp mice (n=7). Numbers adjacent to outlined areas indicate per cent cells in each. (f) FoxP3gfp+ Treg and naive T cells (Teff) were sorted by flow cytometry. The CTV-labelled T cells (1 × 105) were cultured in 96-well plates for 72 h together with a decreasing ratio of sorted miR-31fl/fl or cKO Treg cells in the presence of anti-CD3 (1 μg ml−1) plus γ-irradiated antigen-presenting cells (1 × 105). The suppressive function of Treg cells was determined by the proliferation of activated Teff cells on the basis of CTV dilution. NS, not significant, ***P<0.001, two-tailed Student’s t-test. Data are from one experiment representative of at least two independent experiments (mean±s.e.m.).
Gprc5a is a target of miR-31
To elucidate the mechanisms, we combined microarray gene expression analysis and target prediction to look for putative targets of miR-31. Using a combination of these two approaches, we identified seven predicted target genes that were upregulated in polarized iTreg cells derived from cKO mice (Fig. 5a–c). To confirm accuracy of the microarray data, we validated these potential target genes by increasing sample numbers. We found one predicted target of miR-31, Gprc5a, was significantly upregulated at mRNA levels, and increased by more than 5.0-fold in cKO iTreg cells compared with miR-31fl/fl controls (Supplementary Fig. 6 and Fig. 5d). In contrast to miR-31fl/fl iTreg cells, Gprc5a protein level was also increased by 1.96-fold in cKO iTreg cells (Fig. 5e). Gprc5a was reported to be regulated directly by RA via its receptors, RARs and RXRs33,34. These nuclear RA receptors bind to the Gprc5a promoter for its transcriptional activity24. We next generated a reporter construct that includes the 3′-UTR of Gprc5a mRNA. In contrast to a control construct lacking the target sequence, miR-31 overexpression led to significantly decreased luciferase activity derived from the construct expressing the target sequence (Fig. 5f,g). Thus, our data demonstrate that miR-31 is capable of directly targeting a sequence within the 3′-UTR of Gprc5a mRNA and that Gprc5a is one of the key targets of miR-31 in Treg-cell differentiation.
(a) Total RNAs of polarized iTreg cells derived from 3 miR-31fl/fl controls and 3 cKO mice were used for a microarray analysis (41,174 genes in total). Transcripts of top 63 genes were found to be upregulated in iTreg cells of cKO mice. (b,c) TargetScan and miRnada predicted 1,305 potential targets of miR-31. The overlapping seven genes were defined as ACCEPT genes. (d,e) qPCR or western blot analysis of Gprc5a expression in polarized iTreg cells derived from miR-31fl/fl and cKO mice. (f) WT and point-mutated 3′-UTR reporter constructs. (g) Luciferase activity was determined in NIH3T3 cells that were transfected with miR-31 mimics and the indicated 3′-UTR reporter construct or with the indicated WT or point-mutated 3′-UTR reporter construct (WT UTR or mutant UTR). Results (d) are presented as the ratio of mRNA to the β-actin, relative to that in controls. *P<0.05; ***P<0.001, NS, not significant, two-tailed Student’s t-test. Data (d,e and g) are representative of at least two independent experiments (mean±s.e.m.).
Gprc5a is critical for pTreg-cell differentiation
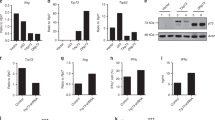
Gprc5a was reported to be expressed preferentially in lung tissue and to be a putative lung tumour suppressor gene35. The functional analysis of Gprc5a in T-cell differentiation and autoimmunity is not yet performed. We induced EAE, and measured Gprc5a expression in inflamed spleen and sorted CD4+ T cells. In contrast with non-immunized controls, Gprc5a expression was significantly decreased in spleen and CD4+ T cells in EAE mice, and this might be the consequence of increased miR-31 under inflammatory conditions (Fig. 6a,b). Western blot analysis confirmed that expression of Gprc5a protein was much lower in spleen and CNS of EAE mice than those of healthy controls (Fig. 6c,d). We assessed the role of Gprc5a in pTreg-cell generation using Gprc5a−/− mice35. Gprc5a deficiency resulted in a marked decrease in the TGF-β1-mediated induction of iTreg cells, but had no impact on the induction of TH1 and TH17 cells (Fig. 6e and Supplementary Fig. 7a,b), suggesting that Gprc5a preferentially regulates FoxP3 expression. Thus, our data demonstrate that Gprc5a is a novel regulator in iTreg-cell generation. Interestingly, consistent with the observation in miR-31 cKO mice, we found that Gprc5a deficiency did not affect tTreg-cell generation in healthy mice (Fig. 6f). To test the impact of Gprc5a on the Treg-cell response during inflammation, we analysed the frequency of Treg cells in inflamed spleen in Gprc5a−/− mice after the induction of EAE. Gprc5a deficiency resulted in a significant decrease in the percentage of pTreg cells compared with WT controls (Fig. 6g, h), suggesting that Gprc5a is critically required for pTreg-cell generation in vivo in autoimmune disease. Notably, Gprc5a−/− mice developed EAE not only much earlier, but also more severe than Gprc5a+/+ mice (Fig. 6i). Moreover, an excessive weight loss was displayed in Gprc5a−/− mice compared with Gprc5a+/+ mice (Fig. 6j). To further investigate whether miR-31 affects EAE development via regulating Gprc5a in vivo, we generated double knockout (DKO) mice by crossing miR-31 cKO mice with Gprc5a−/− mice. We demonstrated that the severity of EAE was significantly reduced in cKO mice compared with Gprc5a−/− mice, however, the disease phenotype was completely restored when Gprc5a was deleted in the cKO mice (Supplementary Fig. 7c). Collectively, our observations indicate that Gprc5a is regulated by miR-31, and functionally involved in the development of EAE. The beneficial effect of RA is possibly due to its stimulation of Gprc5a expression to promote pTreg-cell generation in tissue-specific autoimmune inflammation.
(a,b) Splenocytes were prepared from unimmunized control mice (Ctr) or EAE mice 10 days after immunization (n=3–4). qPCR analysis of Gprc5a expression in total splenocytes or in sorted CD4+ T cells. (c,d) Western blot analysis of Gprc5a in splenocytes and CNS infiltrating cells derived from healthy controls (Ctr) or EAE mice. (e) Naive T cells were sorted from Gprc5a+/+ and Gprc5a−/− mice. Flow cytometry of polarized iTreg cells in the presence of different concentrations of TGF-β1. (f) Flow cytometric analysis of Treg cells in thymus from 6-week-old Gprc5a+/+ and Gprc5a−/− mice (gated on CD4+ T cells). (g,h) Flow cytometric analysis of Treg cells in the inflamed spleen and lymph nodes Gprc5a+/+ and Gprc5a−/− mice 14 days after the induction of EAE (gated on CD4+ T cells). Numbers adjacent to outlined areas indicate per cent cells in each. (i,j) Clinical scores and weight loss (mean±s.e.m.) of Gprc5a+/+ or Gprc5a−/− mice after the induction of EAE were assessed every day (n=7). *P<0.05, **P<0.01, ***P<0.001, NS, not significant, two-tailed Student’s t-test for a,b,f,g and h; one-way analysis of variance for i,j. Data are representative of at least two independent experiments (mean±s.e.m.).
Discussion
Treg cells have been reported to be capable of controlling CNS autoimmunity in several CD4+ T-cell-driven EAE models. Treg-cell frequency within the CNS was increased during the recovery phase of actively induced EAE36,37, and the adoptive transfer of Treg-cells ameliorated EAE symptoms36,38. Furthermore, depletion of Treg cells with anti-CD25 mAb has been demonstrated to exacerbate EAE36. More importantly, reduced Treg-cell proliferative potential and cloning frequency were identified in patients with MS12,39. Thus, regulators of Treg-cell generation are considered to harbour valuable potential for clinical applications in the treatment of autoimmune disorders. Here, we report that miR-31 expression in CD4+ T cells was triggered by TCR signalling, and downregulated by TGF-β1-mediated FoxP3. Its conditional deletion substantially enhanced the pTreg-cell induction and ameliorated disease severity in the EAE model. Mechanistically, we have proven that by targeting Gprc5a, a known retinoic acid-inducible protein, miR-31 promoted the generation of pTreg cells in vivo. Gprc5a is a functional target of miR-31, and its deficiency resulted in impaired pTreg-cell generation and increased EAE severity.
Antigen-specific stimuli delivered through the TCR cooperates with antigen-nonspecific cytokines to support proliferation and differentiation of distinct TH cell subsets40. However, it has become increasingly clear that miRNAs, post-transcriptional regulators, are involved in driving TH cell differentiation and lineage commitment41. A selective effect of miR-31 on pTreg-cell differentiation could be explained by the differential requirement of TCR signalling in the induction of these T-cell lineages. A low antigen dose of a high-affinity TCR ligand favours to induce pTreg cells in vivo9, whereas high doses of TCR stimulation prevents FoxP3 induction and pTreg-cell generation through activating NF-κB signalling42. Indeed, we have determined that the activation of NF-κB induced miR-31 expression through a direct binding of p65 to its promoter (to be published elsewhere). Thus, it is possible that in the absence of TGF-β1, TCR stimulation at high doses elicits activation of NF-κB, which directly triggers the expression of miR-31 inhibiting FoxP3 levels in CD4+ T cells. However, TCR stimulation at low doses induces FoxP3, which may downregulate miR-31 expression through binding to its promoter, providing a feedback loop during pTreg-cell differentiation.
Several miRNAs have been reported to impact Treg-cell development and function. miR-155 is highly expressed in Treg cells, facilitates Treg-cell homeostasis by repressing Socs1 and its deficiency results in decreased numbers of both tTreg cells and pTreg cells43. miR-21 indirectly acts as a positive regulator of human FoxP3 expression21. miR-146a is critical for Treg-cell-mediated control of TH1 responses via targeting Stat1 (ref. 44). Despite that miR-17∼92 is dispensable for the development of tTreg cells in vivo, miR-17∼92 ablation reduces the frequency of MOG35–55-specific pTreg cells during EAE45. miR-10a is induced by TGF-β1 and RA, and promotes the differentiation of pTreg cells through inhibiting Bcl-6 (ref. 46). Collectively, these Treg-cell-associated miRNAs are all enriched in Treg cells compared with conventional T cells, and function as positive regulators. Our data demonstrated that miR-31 was preferentially diminished in Treg cells, was downregulated by FoxP3, and negatively regulated naive CD4+ T-cell differentiation into pTreg cells. The conditional deletion of miR-31 in CD4+ T cells resulted in enhanced induction of pTreg cells in the periphery, and decreased severity of autoimmune disease. Thus, we highlight miR-31 acts as a negative regulator for pTreg-cell generation in vivo. Although different targets of miR-31 were identified, its similar effect was also reported previously for human Treg cells21. Our findings are inconsistent with a recent report which demonstrated that growth factor independent 1 (Gfi-1) was underexpressed in pTreg cells, downregulated by TGF-β1 and limited the pTreg-cell differentiation47.
miR-31 is the only member of a broadly conserved miRNA ‘seed family’ that is present in vertebrates and Drosophila48. miR-31 regulates keratinocyte differentiation through inhibiting hypoxia-inducible factor 1 (ref. 49). Furthermore, in contrast to other T-cell subsets, miR-31 has been shown to be downregulated in human Treg cells21. This raises the intriguing possibility that miR-31 may be preferentially diminished in Treg cells and its upregulation in CD4+ T cells under inflammatory stress may limit pTreg cell induction in human autoimmune diseases.
RA has been proven to facilitate pTreg-cell generation23. RA regulates Gprc5a transcriptional activity by binding to its receptors, RARs and RXRs24,33,34. So far, the role of Gprc5a in the T-cell differentiation programme is not investigated. We here showed that Gprc5a deficiency led to a severe defect in in vitro- and in vivo-generation of Treg cells, as well as increased severity of inflammatory CNS phenotypes, indicating that this may be one of the mechanisms by which RA inhibits autoimmune reactions in vivo. However, the molecular mechanism by which Gprc5a promotes Treg-cell generation and suppresses autoimmune disease is subjected to further investigation. Nevertheless, by generating miR-31 and its target Gprc5a DKO mice we clearly show that miR-31 promotes the development of autoimmune disease through inhibiting Gprc5a.
In summary, our results demonstrate that miR-31 inhibits pTreg-cell generation through directly targeting Gprc5a, a retinoic acid-inducible protein and promotes autoimmunity, therefore, providing the first in vivo genetic evidence that miR-31 and its novel target Gprc5a are critical intrinsic factors for controlling physiological and pathological immune responses regulated by pTreg cells.
Methods
Mice
C57BL/6J mice (stock number: 000664), B6.Cg-FoxP3tm2Tch/J mice (stock number: 006772, designated as FoxP3gfp) and CD4Cre mice (stock number: 017336) were purchased from The Jackson Laboratory (Bar Harbor, ME). B6.129S6-Rag2tm1Fwa N12 (RAG2−/−) mice were purchased from Taconic Labs (Hudson, NY). Gprc5a−/− mice were generated as previously reported35. Mice were kept under specific pathogen-free conditions in compliance with the National Institutes of Health Guide for the Care and Use of Laboratory Animals with the approval (SYXK-2003-0026) of the Scientific Investigation Board of Shanghai Jiao Tong University School of Medicine, Shanghai, China. To ameliorate any suffering of mice observed throughout these experimental studies, mice were euthanized by CO2 inhalation.
Generation of miR-31fl/fl and miR-31fl/flCD4Cre mice
The miR-31 locus (mmu-mir31 ENSMUSG00000065408, http://www.ensembl.org/index.html) is on the chromosome 4 (Mus musculus) and encodes the miR-31. To create loxP-miR-31-loxP mice, a targeting vector was designed to insert with an frt-flanked PGK-neo cassette and a loxP site upstream of miR-31, and a second loxP site downstream. LoxP site is a 34 bp length DNA sequence that can be recognized by Cre recombinase catalyses. If two loxP sites are introduced in the same orientation into a genomic locus, expression of Cre results in the deletion of the loxP-flanked DNA sequence. After linearization, the vector was electroporated into 129S6-derived embryonic stem (ES) cells. The collected ES cells were screened with 300 μg ml−1 G418 and 2 μM Gan C for 8 days and ascertained by PCR. The ES cells with right homologous recombination were injected into blastocyst. After birth, the chimeric mice were bred with 129S mice to generate the heterozygotes. At this point, mutant mice were bred with Flp recombinase-expressing mice to remove the frt-flanked neo cassette. The resulting loxP-miR-31-loxP mice were backcrossed into C57BL/6J background for eight generations and bred with CD4Cre transgenic mice. P1 and P2 were used to genotype the miR-31 floxed allele (1,195 bp) and the miR-31 deleted allele (474 bp). P1, 5′- TTTAAGGGCTCATGGAGCAA -3′; P2, 5′- TGAGGACTTGCAAACGTCAG -3′. Excision by CD4Cre was complete for all pups used in experiments. In some experiments, these mice were further crossed with FoxP3gfp mice to generate mice that express green fluorescent protein (GFP) in their Treg cells.
Induction of EAE
EAE was induced by complete Freund’s adjuvant (CFA)-MOG35–55 peptide immunization (China Peptides Biotechnology) and scored daily. Briefly, C57BL/6J mice were injected subcutaneously into the base of the tail with a volume of 200 μl containing 300 μg MOG35–55 peptide emulsified in CFA (Sigma-Aldrich). Mice were also injected intravenously with 200 ng of pertussis toxin (Merck-Calbiochem) on day 0 and 2 post immunization. All the reagents used for in vivo experiments were free of endotoxin. Mice were monitored daily for the development of disease which was scored according to the following scale: 0, no symptoms; 0.5, partially limp tail; 1, completely limp tail; 1.5, impaired righting reflex; 2, hind limb paresis; 2.5, hind-limb paralysis; 3, forelimb weakness; 4, complete paralysis; 5, moribund or death.
T-cell isolation and sorting
Peripheral T cells were obtained from the spleen and lymph nodes of 6-week-old mice. Naive CD4+ T cells (CD4+CD25−CD62Lhigh) were sorted by FACSAria III (BD Biosciences) after enrichment of CD4+ T cells by the mouse CD4+ T cell Isolation Kit (Miltenyi). For the isolation of naive CD4+ T cells and nTreg cells from Foxp3gfp mouse, CD4+CD25−GFP−CD62Lhigh cells and CD4+CD25+GFP+ cells were isolated, respectively. Cell purity was >94% as determined by flow cytometry. CNS-infiltrating mononuclear cells from EAE mice were prepared by Percoll (GE Healthcare) gradient separation.
In vitro Treg-cell differentiation
All the cultures of T cells used RPMI-1640 medium (Gibco) supplemented with 10% heat-inactivated fetal bovine serum (Gibco), 2 mM L-glutamine (Gibco), 100 IU ml−1 penicillin, 100 μg ml−1 streptomycin, 10 mM HEPES (Gibco) and 5 mM β-mercaptoethanol (Gibco). The naive CD4+CD25−Foxp3gfp-CD62Lhi T cells were activated with plate-bound anti-CD3 (5 μg ml−1; 145-2C11; BD Biosciences) plus soluble anti-CD28 (2 μg ml−1; 37.51; BD Biosciences). TH1-cell differentiation conditions included 10 ng ml−1 rmIL-12 (R&D Systems) and 10 μg ml−1 anti-IL-4 (11B11; Biolegend). The TH17 cell differentiation conditions included 20 ng ml−1 rmIL-6 (R&D Systems), 3 ng ml−1 rmTGF-β1 (R&D Systems), 10 μg ml−1 anti-IL-4 (11B11; Biolegend) and 10 μg ml−1 anti-IFN-γ (XMG1.2; eBioscience). The iTreg-cell differentiation conditions included 5 ng ml−1 rmTGF-β1 (R&D Systems) and 10 ng ml−1 rmIL-2 (R&D Systems).
Flow cytometry
Cytokines, transcriptional factors and surface markers were evaluated by flow cytometry with a FACSCanto II (BD Biosciences). To detect intracellular expression of IL-17A, IFN-γ in CD4+ T cells, lymph nodes or CNS (purified with Percoll) were first treated with 750 ng ml−1 ionomycin (Sigma), 50 ng ml−1 phorbol 12-myristate 13-acetate (PMA) (Sigma) and GolgiPlug (BD Biosciences) for 4–6 h at 37 °C. Cells were fixed and permeabilized with the Foxp3 Staining Buffer Set (eBioscience) or BD Cytofix/Cytoperm (BD Biosciences) and were stained with fluorescent antibodies. After washing, stained cells were assayed with a BD Biosciences FACSCanto II flow cytometer and data were analysed with FlowJo software. For flow cytometry, monoclonal antibodies against CD4 (clone GK1.5), CD8 (clone 53-6.7), CD62L (clone MEL-14), CD44 (clone IM7), CD25 (clone PC61.5), IL-17A (clone eBio17B7), IFN-γ (clone XMG1.2), Helios (22F6, Biolegend) and FoxP3 (clone FJK-16s) were from eBioscience and CD3 (clone 145-2C11) was from Biolegend.
RNA reverse transcription and real-time quantitative PCR
Total RNA was isolated using Trizol (Invitrogen) according to the manufacturer’s instructions. RNA was quantified spectrophotometrically, and 1 μg of total RNA was reverse transcribed into cDNA using SuperScript III (Invitrogen, Carlsbad, CA) in the presence of random hexamers and oligo dT primers (Invitrogen). The cDNA samples were distributed on plates at 200 ng per well and run in triplicate. qPCR was carried out with the FastStart Universal SYBR Green Master (Roche) in a ABI 7500 Fast Real-Time PCR system or ViiA 7 Real-Time PCR System (Applied Biosystems). Primer sequences were listed in Supplementary Table 1. To measure mature miR-31 levels, 50 ng of total RNA was reverse-transcribed using the TaqMan miRNA reverse transcription kit, miR-31 RT primers and U6 snRNA (Applied Biosystems). The cDNAs were then analysed by qPCR using the TaqMan probes for miR-31 and U6 snRNA (Applied Biosystems). Quantification of relative miRNA expression was measured by the comparative CT (critical threshold) method, normalized to endogenous U6 expression and determined by the formula 2−ΔΔCT.
Chromatin immunoprecipitation assay
ChIP assays were performed using the SimpleChIP enzymatic chromatin immunoprecipitation kit (Cell Signaling Technology) according to the manufacturer’s protocol with minor modifications. In brief, the cells were collected and crosslinked with 1% (v/v) formaldehyde for 10 min at room temperature. Subsequently, nuclei were isolated by the lysis of cytoplasmic fraction and chromatin was digested into fragments of 150–900 bp by micrococcal nuclease (400 gel units) for 20 min at 37 °C, followed by ultrasonic disruption of the nuclear membrane using a standard microtip and a Branson W250D Sonifier (four pulses, 60% amplitude, duty cycle 40%). The sonicated nuclear fractions were divided for input control and for overnight incubated at 4 °C with 5 μg either anti-FoxP3 Ab (FJK-16 s; eBioscience) or the negative control IgG (Cell Signaling Technology). After incubation with 30 μl of ChIP grade protein G-agarose beads for 2 h at 4 °C, the antibody-protein–DNA complexes were then eluted from the beads and digested by Proteinase K (40 μg) for 2 h at 65 °C, followed by spin column-based purification of the DNA. Finally, genomic DNA recovered from the ChIP assays were qPCR amplified with primers specific to the FoxP3-binding elements of the miR-31 promoter region. The primers used for detection of miR-31 promoter sequences were listed (Supplementary Table 1). The specificity of each primer set was verified by analysing the dissociation curve of each gene-specific PCR product.
In vitro Treg-cell suppression assay
Treg cells were isolated from the spleen of mice by sorting with flow cytometry based on cell surface markers (CD4, CD25 and FoxP3gfp). Naive CD4+ T cells were isolated from the spleen of mice by sorting with flow cytometry based on cell surface markers (CD4+, CD25−, CD62Lhi and FoxP3gfp). Splenocytes from RAG2−/− mice lacking mature T and B lymphocytes were used as antigen presenting cells. The purified naive CD4+ T cells were labelled for 15 min at 37 °C with 10 μM CTV (Life Technologies) and the CTV-labelled T cells (1 × 105) were cultured in 96-well plates for 72 h together with an increasing ratio of sorted Treg cells in the presence of anti-CD3 (1 μg ml−1) plus γ-irradiated antigen-presenting cells (1 × 105). The suppressive function of Treg cells was determined by measurement of the proliferation of activated CD4+ effector T cells on the basis of CTV dilution.
Generation of bone marrow chimeric mice
Bone marrow cells were flushed from miR-31fl/fl or cKO donor mice, and 5 × 106 T-cell-depleted bone marrow cells were transplanted into each C57BL/6J host mouse with total-body irradiation of 950 cGy in two divided doses. Chimeric mice reconstituted with bone marrow cells derived from either miR-31fl/fl or cKO were subjected for EAE induction 8 weeks after the transplantation.
Histology
Spinal cords from miR-31fl/fl or cKO EAE mice were fixed in 4% paraformaldehyde and paraffin embedded. Paraffin-embedded 5-μm sections of spinal cord were stained with haematoxylin and eosin or Luxol fast blue and then examined by light microscopy (Axio scope A1, Zeiss).
Luciferase reporter plasmid
The Gprc5a 3′-UTR was amplified using primers Gprc5a Forward, 5′- AATCTCGAGCTGTTGGGAAGAGTGGGAC -3′, Reverse, 5′- TCGGCGGCCGCAATAGTTGTGACCACATCTTTATTG -3′. The Gprc5a 3′-UTR genomic fragment was digested with XhoI-NotI and inserted into the corresponding sites of the psiCheck-2 Synthetic firefly luciferase reporter plasmid (Promega). This construct was also used to generate a miR-31 ‘seed’ mutant plasmid. The mutagenic primers used for Gprc5a were Mutant Forward, 5′- AATCTCGAGCTGTTGGGAAGAGTGGGAC -3′, Mutant Reverse, 5′- CAGCCCCACCGTTCTCGGCGGTG -3′. The correctness of all the plasmids was confirmed by sequencing.
Luciferase assays
All 3′ UTR reporter vectors were prepared by amplifying the 3′ UTRs of Gprc5a, followed by insertion into the psiCHECK-2 vector (Promega). Site-specific mutants were generated by PCR in the psiCHECK-2 vector (Promega). NIH3T3 cells were maintained in DMEM (HyClone) supplemented with 10% fetal bovine serum (HyClone), 2 mM glutamine, 100 IU ml−1 penicillin, 0.1 mg ml−1 streptomycin. NIH3T3 cells were seeded in the 24-well plates (1 × 105 cells per well) one day before transfection and then each well was transfected with a mixture of 100 ng 3′-UTR luciferase reporter vector and 50 pmol miRNA mimics or controls. Twenty four hours post transfection, the cells were lysed. Then, luciferase activity was measured using the Dual-Luciferase Reporter Assay System (Promega), using a Lumat3 LB 9508 Single Tube Luminometer instrument (Berthold Technologies). Each experiment was performed in triplicate. The ratio of Renilla luciferase to Firefly luciferase was calculated for each well.
Western blotting
Cultured T cells and the CNS of mice were lysed in radio immunoprecipitation assay buffer supplemented with protease and phosphatase inhibitor cocktail (Thermo Scientific). Mouse anti-actin Ab (1:3,000) (Cell Signaling Technology), rabbit anti-Gprc5a Ab (1:1,000) (Dr Jiong Deng provided) were used. The signal was detected with Pierce ECL Western Blotting Substrate (Thermo Scientific) and GE ImageQuant LAS 4000 (GE Healthcare). Images have been cropped for presentation. Full size images are presented in Supplementary Fig. 8.
MicroRNA microarray analysis
Naive T cells from 3 miR-31fl/fl mice and 3 miR-31fl/flCD4Cre mice were used to induce iTreg in vitro. Total RNA was isolated using RNeasy MiniKit (Qiagen). Mouse genome-wide cDNA microarray analysis was performed by Shanghai Biotechnology (Shanghai).
Statistical analysis
The data were analysed with GraphPad Prism 5 and were presented as the mean±s.e.m. Student’s t-test was used when two conditions were compared, and analysis of variance with Bonferroni or Newman–Keuls correction was used for multiple comparisons. Probability values of <0.05 were considered significant; two-sided tests were performed.
Additional information
Accession codes: Microarray data have been deposited in the GEO database under accession code GSE61938.
How to cite this article: Zhang, L. et al. MicroRNA-31 negatively regulates peripherally derived regulatory T-cell generation by repressing retinoic acid-inducible protein 3. Nat. Commun. 6:7639 doi: 10.1038/ncomms8639 (2015).
Accession codes
References
Yamane, H. & Paul, W. E. Cytokines of the gamma(c) family control CD4+ T cell differentiation and function. Nat. Immunol. 13, 1037–1044 (2012).
Zhu, J., Yamane, H. & Paul, W. E. Differentiation of effector CD4 T cell populations (*). Annu. Rev. Immunol. 28, 445–489 (2010).
Simmons, S. B., Pierson, E. R., Lee, S. Y. & Goverman, J. M. Modeling the heterogeneity of multiple sclerosis in animals. Trends Immunol. 34, 410–422 (2013).
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Sakaguchi, S., Miyara, M., Costantino, C. M. & Hafler, D. A. FOXP3+ regulatory T cells in the human immune system. Nat. Rev. Immunol. 10, 490–500 (2010).
Abbas, A. K. et al. Regulatory T cells: recommendations to simplify the nomenclature. Nat. Immunol. 14, 307–308 (2013).
Curotto de Lafaille, M. A. & Lafaille, J. J. Natural and adaptive foxp3+ regulatory T cells: more of the same or a division of labor? Immunity 30, 626–635 (2009).
Chen, W. et al. Conversion of peripheral CD4+CD25- naive T cells to CD4+CD25+ regulatory T cells by TGF-beta induction of transcription factor Foxp3. J. Exp. Med. 198, 1875–1886 (2003).
Gottschalk, R. A., Corse, E. & Allison, J. P. TCR ligand density and affinity determine peripheral induction of Foxp3 in vivo. J. Exp. Med. 207, 1701–1711 (2010).
Venken, K. et al. Compromised CD4+ CD25(high) regulatory T-cell function in patients with relapsing-remitting multiple sclerosis is correlated with a reduced frequency of FOXP3-positive cells and reduced FOXP3 expression at the single-cell level. Immunology 123, 79–89 (2008).
Huan, J. et al. Decreased FOXP3 levels in multiple sclerosis patients. J. Neurosci. Res. 81, 45–52 (2005).
Carbone, F. et al. Regulatory T cell proliferative potential is impaired in human autoimmune disease. Nat. Med. 20, 69–74 (2014).
Buckner, J. H. Mechanisms of impaired regulation by CD4(+)CD25(+)FOXP3(+) regulatory T cells in human autoimmune diseases. Nat. Rev. Immunol. 10, 849–859 (2010).
Kim, V. N. & Nam, J. W. Genomics of microRNA. Trends Genet. 22, 165–173 (2006).
Ceribelli, A., Satoh, M. & Chan, E. K. MicroRNAs and autoimmunity. Curr. Opin. Immunol. 24, 686–691 (2012).
Rossi, R. L. et al. Distinct microRNA signatures in human lymphocyte subsets and enforcement of the naive state in CD4+ T cells by the microRNA miR-125b. Nat. Immunol. 12, 796–803 (2011).
Cobb, B. S. et al. A role for Dicer in immune regulation. J. Exp. Med. 203, 2519–2527 (2006).
Liston, A., Lu, L. F., O'Carroll, D., Tarakhovsky, A. & Rudensky, A. Y. Dicer-dependent microRNA pathway safeguards regulatory T cell function. J. Exp. Med. 205, 1993–2004 (2008).
Chong, M. M., Rasmussen, J. P., Rudensky, A. Y. & Littman, D. R. The RNAseIII enzyme Drosha is critical in T cells for preventing lethal inflammatory disease. J. Exp. Med. 205, 2005–2017 (2008).
Zhou, X. et al. Selective miRNA disruption in T reg cells leads to uncontrolled autoimmunity. J. Exp. Med. 205, 1983–1991 (2008).
Rouas, R. et al. Human natural Treg microRNA signature: role of microRNA-31 and microRNA-21 in FOXP3 expression. Eur. J. Immunol. 39, 1608–1618 (2009).
Bastien, J. & Rochette-Egly, C. Nuclear retinoid receptors and the transcription of retinoid-target genes. Gene 328, 1–16 (2004).
Hill, J. A. et al. Retinoic acid enhances Foxp3 induction indirectly by relieving inhibition from CD4+CD44hi Cells. Immunity 29, 758–770 (2008).
Ye, X., Tao, Q., Wang, Y., Cheng, Y. & Lotan, R. Mechanisms underlying the induction of the putative human tumor suppressor GPRC5A by retinoic acid. Cancer Biol. Ther. 8, 951–962 (2009).
Park, H. et al. A distinct lineage of CD4 T cells regulates tissue inflammation by producing interleukin 17. Nat. Immunol. 6, 1133–1141 (2005).
Bilate, A. M. & Lafaille, J. J. Induced CD4+Foxp3+ regulatory T cells in immune tolerance. Annu. Rev. Immunol. 30, 733–758 (2012).
Jager, A., Dardalhon, V., Sobel, R. A., Bettelli, E. & Kuchroo, V. K. Th1, Th17, and Th9 effector cells induce experimental autoimmune encephalomyelitis with different pathological phenotypes. J. Immunol. 183, 7169–7177 (2009).
Thornton, A. M. et al. Expression of Helios, an Ikaros transcription factor family member, differentiates thymic-derived from peripherally induced Foxp3+ T regulatory cells. J. Immunol. 184, 3433–3441 (2010).
Strainic, M. G., Shevach, E. M., An, F., Lin, F. & Medof, M. E. Absence of signaling into CD4(+) cells via C3aR and C5aR enables autoinductive TGF-beta1 signaling and induction of Foxp3(+) regulatory T cells. Nat. Immunol. 14, 162–171 (2013).
Korn, T. et al. IL-6 controls Th17 immunity in vivo by inhibiting the conversion of conventional T cells into Foxp3+ regulatory T cells. Proc. Natl Acad. Sci. USA 105, 18460–18465 (2008).
Haribhai, D. et al. A central role for induced regulatory T cells in tolerance induction in experimental colitis. J. Immunol. 182, 3461–3468 (2009).
Curotto de Lafaille, M. A. et al. Adaptive Foxp3+ regulatory T cell-dependent and -independent control of allergic inflammation. Immunity 29, 114–126 (2008).
Tao, Q., Cheng, Y., Clifford, J. & Lotan, R. Characterization of the murine orphan G-protein-coupled receptor gene Rai3 and its regulation by retinoic acid. Genomics 83, 270–280 (2004).
Robbins, M. J. et al. Molecular cloning and characterization of two novel retinoic acid-inducible orphan G-protein-coupled receptors (GPRC5B and GPRC5C). Genomics 67, 8–18 (2000).
Tao, Q. et al. Identification of the retinoic acid-inducible Gprc5a as a new lung tumor suppressor gene. J. Natl Cancer Inst. 99, 1668–1682 (2007).
McGeachy, M. J., Stephens, L. A. & Anderton, S. M. Natural recovery and protection from autoimmune encephalomyelitis: contribution of CD4+CD25+ regulatory cells within the central nervous system. J. Immunol. 175, 3025–3032 (2005).
O'Connor, R. A., Malpass, K. H. & Anderton, S. M. The inflamed central nervous system drives the activation and rapid proliferation of Foxp3+ regulatory T cells. J. Immunol. 179, 958–966 (2007).
Mekala, D. J., Alli, R. S. & Geiger, T. L. IL-10-dependent infectious tolerance after the treatment of experimental allergic encephalomyelitis with redirected CD4+CD25+ T lymphocytes. Proc. Natl Acad. Sci. USA 102, 11817–11822 (2005).
Viglietta, V., Baecher-Allan, C., Weiner, H. L. & Hafler, D. A. Loss of functional suppression by CD4+CD25+ regulatory T cells in patients with multiple sclerosis. J. Exp. Med. 199, 971–979 (2004).
Yamane, H. & Paul, W. E. Early signaling events that underlie fate decisions of naive CD4+ T cells toward distinct T-helper cell subsets. Immunol. Rev. 252, 12–23 (2013).
Kanno, Y., Vahedi, G., Hirahara, K., Singleton, K. & O'Shea, J. J. Transcriptional and epigenetic control of T helper cell specification: molecular mechanisms underlying commitment and plasticity. Annu. Rev. Immunol. 30, 707–731 (2012).
Molinero, L. L., Miller, M. L., Evaristo, C. & Alegre, M.-L. High TCR stimuli prevent induced regulatory T cell differentiation in a NF-κB–dependent manner. J. Immunol. 186, 4609–4617 (2011).
Lu, L.-F. et al. Foxp3-dependent microRNA155 confers competitive fitness to regulatory T cells by targeting SOCS1 protein. Immunity 30, 80–91 (2009).
Lu, L.-F. et al. Function of miR-146a in controlling treg cell-mediated regulation of Th1 responses. Cell 142, 914–929 (2010).
de Kouchkovsky, D. et al. microRNA-17–92 regulates IL-10 production by regulatory T cells and control of experimental autoimmune encephalomyelitis. J. Immunol. 191, 1594–1605 (2013).
Takahashi, H. et al. TGF-[beta] and retinoic acid induce the microRNA miR-10a, which targets Bcl-6 and constrains the plasticity of helper T cells. Nat. Immunol. 13, 587–595 (2012).
Zhu, J. et al. Down-regulation of Gfi-1 expression by TGF-beta is important for differentiation of Th17 and CD103+ inducible regulatory T cells. J. Exp. Med. 206, 329–341 (2009).
Valastyan, S. & Weinberg, R. A. miR-31: a crucial overseer of tumor metastasis and other emerging roles. Cell Cycle 9, 2124–2129 (2010).
Peng, H. et al. microRNA-31/factor-inhibiting hypoxia-inducible factor 1 nexus regulates keratinocyte differentiation. Proc. Natl Acad. Sci. USA 109, 14030–14034 (2012).
Acknowledgements
This work is supported by grants from 973 program and National Natural Science Foundation of China (No: 2012CB917100, No: 2014CB541905, No: 31330026, No. 81202304 and No: 91029730), by the Leading Academic Discipline Project of the Shanghai Municipal Education Commission (No: J50208 and No: J50207).
Author information
Authors and Affiliations
Contributions
L.Z. and H.W. designed the research; H.W. supervised the research; L.Z. conducted the experiments; H.W. wrote the manuscript and revised the manuscript; F.K., Z.L., J.B., J.L., S.Y., Z.X., F.L., H.Z., Y.S., W.C., Y,G., Z.H. and J.D. helped with the experiments; Q.L., X.-Z.Y., Y.Q. and Q.-J.L. commented on the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-8 and Supplementary Table 1 (PDF 2718 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhang, L., Ke, F., Liu, Z. et al. MicroRNA-31 negatively regulates peripherally derived regulatory T-cell generation by repressing retinoic acid-inducible protein 3. Nat Commun 6, 7639 (2015). https://doi.org/10.1038/ncomms8639
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms8639
This article is cited by
-
Deregulation of circ_003912 contributes to pathogenesis of erosive oral lichen planus by via sponging microRNA-123, -647 and -31 and upregulating FOXP3
Molecular Medicine (2021)
-
Derangement of cell cycle markers in peripheral blood mononuclear cells of asthmatic patients as a reliable biomarker for asthma control
Scientific Reports (2021)
-
Current insight into the functions of microRNAs in common human hair loss disorders: a mini review
Human Cell (2021)
-
MicroRNAs and immunity in periodontal health and disease
International Journal of Oral Science (2018)
-
Hit-and-run programming of therapeutic cytoreagents using mRNA nanocarriers
Nature Communications (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.