Abstract
Enhancement of cellular senescence in tumours triggers a stable cell growth arrest and activation of an antitumour immune response that can be exploited for cancer therapy. Currently, there are only a limited number of targeted therapies that act by increasing senescence in cancers, but the majority of them are not selective and also target healthy cells. Here we developed a chemogenomic screening to identify compounds that enhance senescence in PTEN-deficient cells without affecting normal cells. By using this approach, we identified casein kinase 2 (CK2) as a pro-senescent target. Mechanistically, we show that Pten loss increases CK2 levels by activating STAT3. CK2 upregulation in Pten null tumours affects the stability of Pml, an essential regulator of senescence. However, CK2 inhibition stabilizes Pml levels enhancing senescence in Pten null tumours. Taken together, our screening strategy has identified a novel STAT3–CK2–PML network that can be targeted for pro-senescence therapy for cancer.
Similar content being viewed by others
Introduction
Both genetic and epigenetic losses of the tumour suppressor gene PTEN are frequently observed in human cancers1. Interestingly, while monoallelic loss or mutation of PTEN drives cellular proliferation, complete inactivation of PTEN induces a senescence response, termed PTEN loss-induced cellular senescence (PICS) that opposes tumour development1. However, this response is not complete since upon Pten inactivation in vivo, a fraction of Pten null cells escapes senescence promoting tumor formation1,2. We have previously demonstrated that inhibition of either MDM2 or PTEN by means of small-molecule inhibitors enhances PICS thereby blocking tumour progression1. While this response depends on the functional activation of p53, recent findings demonstrate that senescence can also be activated in Pten; Trp53 double null tumours by enhancing p27 levels through Skp2 inhibition3. These findings, together with the recent demonstration that senescent tumour cells activate an anti-tumour immune response4, have gained increasing interest for the clinical development of compounds that enhance senescence in cancer. However, the use of pro-senescence compounds in the clinic has been limited by their poor tolerability in cancer patients3. Indeed, the majority of these compounds block genes such as CDKs or MYC which are essential for the function of normal cells3. Identification of novel genes that regulate senescence in PTEN-deficient tumours without affecting normal cells would help to develop more effective and selective pro-senescence compounds for the treatment of different types of cancer3. This may be particularly relevant for prostate cancer therapy since PTEN-deficient prostate tumours conserve an intact p53 response even at late stage of tumorigenesis and may therefore benefit from treatments that enhance senescence by targeting p53 or its regulators. Here we report the results of a screening designed to identify ‘druggable’ targets regulating senescence in PTEN-deficient cells. Using chemogenomic and short hairpin RNA (shRNA) approaches, we found that the inhibition of CK2 enhances PICS without affecting normal cells. Our results demonstrate that CK2 is selectively upregulated in PTEN null cells and tumours and opposes PICS by affecting the stability of the promyelocytic leukaemia protein, PML, a known regulator of senescence in cancer5. Interestingly, our study also demonstrates that the upregulation of CK2 in Pten null tumours is mediated by activation of Stat3, which is highly phosphorylated in Pten null tumours. Taken together, our data provide new insight into the mechanisms that drive senescence evasion in tumour cells and validate the effectiveness of an integrated approach for the identification of ‘pro-senescence’ compounds in PTEN-deficient tumours.
Results
Identification of pro-senescence compounds
For the identification of novel pro-senescence compounds that could be rapidly moved to the clinic, we took advantage of a drug library of 500 compounds including small-molecule inhibitors in advanced phase of preclinical development, or marked in the clinic for the therapy of cancer and additional disorders (Supplementary Fig. 1a). This library was created from a larger library of 90,000 compounds comprising: (1) New chemical entities, (2) compounds in advanced phase of development, (3) compounds already in the clinic for the treatment of cancer and different disorders. The rationale behind the selection of compounds in advanced phase of development was based on the fact that these compounds have already passed a significant number of tests both in vitro and in vivo (safety, toxicity and pharmacokinetics), thereby decreasing the risks of drug development failure due to adverse toxicity which is commonly observed in screenings based on new chemical entities6. Next, we set up a ‘pro-senescence’ screening platform, to evaluate the pro-senescence efficacy of these compounds in Pten−/− mouse embryonic fibroblasts (MEFs; Fig. 1a). As previously reported, upon inactivation of Pten, 30% of the cells undergo senescence whereas the remaining 70% of Pten−/− cells bypass this arrest and continue to proliferate7 (Supplementary Methods). This provides a screening window to identify compounds that enhance senescence by further decreasing proliferation of Pten−/− cells, as previously demonstrated for Nutlin-3, a MDM2 inhibitor1. Compounds that increased the percentage of senescent cells in the screening platform were designated as pro-senescence compounds based on two parameters: 1) inhibition of cell proliferation and 2) increase of SA-β-galactosidase staining (SA-β-gal), a prototypical senescence marker1 (Fig. 1a). Experimentally, the screening was carried out in three steps. In the first step, 500 compounds were tested in triplicates using a single drug concentration (10 μM). Compounds that promoted a strong growth arrest in Pten null MEFs passed the first filter (efficacy filter; n=144/500). Positive hits were retested in triplicates in single concentration in both Ptenfl/fl and Pten−/− MEFs to assess growth arrest in both cell types (specificity filter). Finally, the selective compounds (n=33/144) were validated in duplicate in a concentration response assay (dose optimization filter). To perform IC50, data were normalized on each plate according to the negative control on Ptenfl/fl and Pten−/− cells, respectively. IC50s were then evaluated on normalized data by using a four-parameter logistic model independently on each cell line. Twelve out of 33 compounds had a marked difference in the IC50 between Ptenfl/fl and Pten−/− cells (Pten−/−IC50≤ 60% Ptenfl/flIC50) and were selected for the final step of the screening. Examples of compounds that did not pass the optimization filter despite a previous selectivity at single dosage were: Clotrimazole8,9, Clomipramine hydrochloride10, Nifedipine11, Trichostatin A12,13 and Garcinol14 (not specific cell growth arrest between Ptenfl/fl and Pten−/− cells at the IC50). Finally, compounds were tested in SA-β-gal assay using both the cell lines to test for cellular senescence induction (SA-β–gal filter). Compounds that increased the SA-β-gal staining more than 30% as compared to dimethyl sulfoxide in Pten−/− cells passed this filter and were further validated by additional senescence markers (PAI-1 and p21) using RT–PCR and western blot. Seven out of 12 compounds were found to have significantly enhanced senescence in Pten−/− cells as compared to the untreated control (Supplementary Table 1a). Among these seven compounds, we found two bona fide (quinalizarin/CK2i-Q, CX-4945/CK2i-CX)16,24,25 and one predicted (CK2i-S)17 casein kinase 2 (CK2) small-molecule inhibitors, two cytotoxic compounds already available in the clinic (Carboplatin, Methotrexate18,19), a KISS120,21 and a DKK1 inhibitor (proprietary data of Sibi Spa; Supplementary Table 1a, Supplementary Table 2b and Supplementary Fig. 1b). Examples of compounds that did not pass the SA-β–gal filter were: Penfluridol22 and 5-Iodotubericidin23 (did not induce cellular senescence in the SA-β-gal assay in both cell lines). Interestingly, bioinformatics analysis revealed that CK2 and DKK1 inhibitors targeted common genetic pathways, whereas Carboplatin, Methotrexate and KISS1 targeted distinct and independent pathways (Supplementary Table 1b). These compounds were also highly selective for Pten−/− cells (inhibition of proliferation for Ptenfl/fl≤20% and for Pten−/− ≥50%) as compared to a panel of anticancer compounds, already available in the clinic and included in the library (Supplementary Fig. 1b,c).
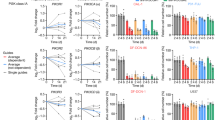
(a) Graphical representation of the pro-senescence screening cascade using Ptenfl/fl and Pten−/− primary MEFs (left panel). The assay is based on the assessment of cellular proliferation and senescence-associated SA-β-gal activity using compounds administered at single dose or in a dose–response assay (right panel). (b) Proliferation of primary Pten−/− MEFs treated with vehicle, the MDM2 inhibitor (MDM2i) Nutlin-3, the CK2 inhibitors quinalizarin (CK2i-Q) and CX-4945 (CK2i-CX) over a period of 5 days. (c) Dose–response assay in primary Ptenfl/fl and Pten−/− MEFs to determine the IC50 of CK2i-Q and CK2i-CX. (d) Quantification of SA-β-gal positivity in Ptenfl/fl and Pten−/− MEFs treated with CK2i-Q, CK2i-CX, Nutlin-3 and Rapamycin (Rapa; upper Panel). Representative images of SA-β-gal staining in cells treated as indicated above (lower panel). Magnification is × 20. Scale bar, 10 μM (e) Proliferation and SA-β-gal positivity of different cancer cell lines treated with CK2i-Q. (f) Proliferation (upper panel), representative images and quantification of SA-β-gal positivity (lower panel) in HCT 116 isogenic cells treated with CK2i-Q. (c,d–f). Scale bar, 10 μM. Data are represented as mean±s.d., n≥3. (***P<0.001, **P<0.01). P value indicates that the statistical significance was measured by Student’s t-test.
A kinome shRNA screen validates CK2 as a pro-senescence target
As an independent approach to validate the targets identified in the chemical screening, we employed ‘loss-of-representation’ or ‘drop-out’ genetic screens that allow the rapid screening of the effects of individual shRNAs in an shRNA library used to infect populations of Ptenfl/fl and Pten−/− primary MEFs (Supplementary Fig. 2a). To this end, we obtained a lentiviral kinome knockdown library comprising 3,280 shRNAs targeting 672 mouse kinases. After infection of Ptenfl/fl MEFs with the lentiviral library and Adeno-GFP or Adeno-Cre, PCR was used to isolate the integrated shRNAs from genomic DNA and deep sequencing of the shRNAs enabled the quantitative assessment of the relative abundance of these shRNAs in the cell populations before and after passaging of the cells (Supplementary Fig. 2a). We first filtered out shRNAs that were represented by less than 200 reads in the deep sequencing analyses of the early time points and then selected individual shRNAs whose representation in the shRNA population displayed little change or only modest changes in Ptenfl/fl cells (less than an 8-fold change in representation at the late versus the early time point) but exhibited between 64- and 1,200-fold decreases in representation over time in the Pten−/− cells. A specific reduction in the representation of shRNAs over time in the Cre-infected cells was indicative of anti-proliferative or anti-survival effects of those knockdowns in the Pten−/− cells (Supplementary Fig. 2b). We conducted this screen twice and identified 75 shRNAs that fulfilled these criteria in both screens (Supplementary Table 2a). Consistent with the results of our chemical screen, among these hits was an shRNA targeting Csnk2a1, which encodes CK2α, a subunit of CK2. Western blotting confirmed that CK2α was indeed downregulated in both Ptenfl/fl and Pten−/− MEFs infected with the CK2 shRNA used in the screening (Supplementary Fig. 2c). Next we assessed senescence and found an increase in both, SA-β-gal staining and p53 protein levels using different CK2 shRNAs (Supplementary Fig. 2d–f). Additional hits included shRNAs targeting Akt2 and Akt3, known downstream effectors of the PI3K pathway, and Plk1, a gene previously shown to be essential for the viability of PTEN-deficient cells, suggesting that the screen was able to functionally identify genetic interaction partners in Pten mutant cells. However, these and additional targets were not identified by the chemical screening and were not tested for senescence induction in Pten null cells.
CK2 inhibitors act as selective pro-senescence compounds
CK2 inhibitors (CK2i) triggered a strong cell growth arrest selectively in Pten−/− MEFs with no detectable level of apoptosis at dosage below 30 μM. Inhibition of proliferation in cells treated with two CK2 inhibitors, quinalizarin24 and CX-4945 (ref. 25) was similar to that of the cells treated with the MDM2 inhibitor, Nutlin-3, used as positive control26 (Fig. 1b). Dose–response experiments performed with quinalizarin and CX-4945 and three additional CK2 inhibitors confirmed the high selectivity of these compounds for Pten−/− as compared to Ptenfl/fl MEFs (Fig. 1c and Supplementary Table 2b).
Moreover, CK2 inhibition by quinalizarin strongly increased SA-β-gal staining and both p21 and PAI-1 expression in Pten−/− as compared to Ptenfl/fl cells (Fig. 1d and Supplementary Fig. 3a,b). Next we checked whether this selectivity was also observed in human cancer cell lines expressing different PTEN levels (Supplementary Table 3). Interestingly, CK2 inhibition resulted in an enhanced senescence response in PTEN-deficient cells when compared with human cancer cell lines expressing high levels of PTEN (Fig. 1e). These findings were also validated in a pair of HCT116 isogenic cancer cells with and without PTEN27 (Fig. 1f). Finally, the selectivity of these compounds was tested in MEFs of different genetic backgrounds. Although CK2 inhibition induced a strong senescence response in Pten−/− cells, it did not selectively affect senescence in Trp53−/− and H-RasV12 infected MEFs (Supplementary Fig. 3c,d). Taken together, these data suggest that Pten−/− cells and PTEN-deficient human cancer cell lines are more sensitive than cells of other genetic backgrounds to CK2 inhibition.
CK2 transcription and protein levels are controlled by STAT3
Next, we sought the mechanism underlying the selectivity of CK2 inhibitors for PTEN-deficient cells. CK2 is a serine/threonine protein kinase that functions as a tetramer of two α and two β subunits. The α subunits contain the catalytic activity, while the β subunits function as regulatory subunits, undergoing autophosphorylation28. Interestingly, while CK2 levels are consistently elevated in some subsets of human cancers, the mechanisms involved in CK2 upregulation remain unknown29. We detected a strong upregulation of CK2α protein (hereafter referred to as CK2) and mRNA levels in both Pten−/− cells and Pten null prostate conditional (Ptenpc−/−) tumours but not in Ptenfl/fl cells and Pten normal prostates (Ptenpc+/+), (Fig. 2a,b and Supplementary Fig. 4a). Of note, CK2 protein upregulation was also validated in PTEN−/− HCT116 human isogenic cancer cells (Supplementary Fig. 4b). The increased CK2 levels in Ptenpc−/− tumours correlated with an increased CK2 kinase activity as demonstrated by phosphorylation of AKT(Ser129), a specific marker of CK2 activity30 (Fig. 2c). We next performed a bioinformatics analysis in Ptenpc+/+ and Ptenpc−/− tumours to identify transcription factors (TFs) responsible for CK2 mRNA upregulation. Towards this end, we analysed which of the most upregulated TFs in Ptenpc−/− tumours could also bind the CK2 promoter and identified three different STAT family members as candidate TFs (Fig. 2d). In response to cytokines and growth factors, STAT proteins are phosphorylated by receptor-associated kinases and then form homo- or heterodimers that translocate to the nucleus, where they act as transcriptional activators31. Amongst these three STAT family members, signal transducer and activator of transcription 3, Stat3 was strongly phosphorylated at both Ser727 and Tyr705 in Ptenpc−/− tumours as compared to the Ptenpc+/+ normal prostates (Fig. 2e,f), whereas Stat1 and Stat2 were barely detectable by western blot analysis. Interestingly, Stat3 protein and mRNA levels were also strongly increased in Ptenpc−/− tumours (Fig. 2e and Supplementary Fig. 4c). These data suggest that the activation of Stat3 in Ptenpc−/− tumours may control CK2 transcription. To genetically validate this hypothesis, we intercrossed Ptenlx/lx; PbCre+ mice with Stat3lx/lx mice to generate Ptenpc−/−; Stat3pc−/− mice. Inactivation of Stat3 in Ptenpc−/− tumours (Supplementary Fig. 4d,e) decreased the mRNA levels of CK2 to the levels of the Ptenpc+/+ normal tissue (Fig. 2g). CK2 mRNA levels were also strongly decreased in Pten−/− MEFs treated with NVP-BSK805, an inhibitor of the JAK2–STAT3 pathway32 (Fig. 2h). To further validate the role of Stat3 in the regulation of CK2 transcription, we carried out a chromatin immunoprecipitation experiment in Pten−/− cells and detected a significant enrichment of Stat3 at the CK2 promoter in correspondence to a predicted STAT responsive element (Fig. 2i). As a positive control for this experiment, we used Socs3, a previously characterized STAT3 target gene33 (Supplementary Fig. 4f). Experiments using a human CK2 promoter reporter confirmed that CK2 promoter activity was high in Pten−/− cells and that the inhibition of the Jak2/Stat3 pathway reduced CK2 promoter activity (Fig. 2j). Finally, we mutated five Stat3-predicted binding sites on the human CK2 promoter and we found that three out of the five mutants showed a profound reduction in the CK2 luciferase activity (Fig. 2k). In summary, these data demonstrate that Stat3 binds to the CK2 promoter and increases CK2 transcription in both Pten−/− cells and tumours.
(a) Western blot analysis and quantification of CK2 and Pten protein levels in primary Ptenfl/fl and Pten−/− MEFs (upper panel) and Ptenpc+/+ normal prostates and Ptenpc−/− prostate tumours (10–12weeks of age) (lower panel). (b) mRNA levels of CK2 in primary Ptenfl/fl and Pten−/− MEFs (upper panel) and Ptenpc+/+ normal prostates and Ptenpc−/− prostate tumours (10–12weeks of age) (lower panel). (c) Immunofluorescence staining for pAKTSer129 in Ptenpc+/+ and Ptenpc−/− tumour sections. Western blot shows the protein level of Pten and pAKTSer129 in Ptenpc+/+ normal prostates and Ptenpc−/− prostate tumours (10–12weeks of age). Scale bar: 100 μM. (d) Heat map representation of the most upregulated and downregulated TFs predicted to bind to the CK2 promoter in Ptenpc+/+ normal prostates and Ptenpc−/− prostate tumours (n=6). (e) IHC analysis of 12-week-old prostates from Ptenpc+/+ and Ptenpc−/− tumours, stained for total STAT3, pSTAT3Ser727 and pSTAT3Tyr705. Magnification in all panels is × 40. Insets in all panels show the × 10 magnification Scale bar, 50 μM. (f) pSTAT3Ser727 and pSTAT3Tyr705 quantification of the experiment shown in e (n=4). (g) mRNA levels of CK2α in normal prostates and prostate tumours of the indicated genotypes. (h) Transcript levels of CK2α in Ptenfl/fl and Pten−/− MEFs treated with or without the JAK2/STAT inhibitor, NVP-BSK805. (i) Schematic representation of the CK2α promoter region (upper panel) and the ChIP assay performed in Pten−/− MEFs to confirm the Stat3 binding to the CK2α promoter (lower panel). Arrows indicate the length and position of the primers used in the experiment (j) Luciferase assay showing the promoter activity of human CK2α in cells treated with and without the JAK2/STAT inhibitor, NVP-BSK805. (k) Schematic representation of the mutated human STAT3-binding sites in the CK2 promoter region (upper panel) and luciferase activity (lower panel). Images (a,b,f,g,h,j,k) Data are represented as mean±s.d., n≥3 (*P<0.05,**P<0.01, ***P<0.001). P value indicates the statistical significance as measured by Student’s t-test.
As PTEN is frequently lost in patients with prostate cancer, we checked the correlation between PTEN, CSNK2A1 and STAT3 mRNA levels in patients affected by advance prostate tumours by analysing two large microarray data sets. In the majority of invasive prostate cancers (n=121), PTEN expression was significantly decreased as compared to the normal prostate samples (n=72; Supplementary Fig. 5a,b). These data reflect the high frequency of PTEN loss of heterozygosity in advanced prostate tumours. Interestingly, we found a correlation between decreased PTEN expression and increased STAT3 and CSNK2A1 mRNA levels in prostate cancer samples, in agreement with our findings in mouse tumours (Supplementary Fig. 5a,b). Moreover, CSNK2A1 overexpression and PTEN downregulation correlated with poor prognosis in patients affected by prostate cancer (both disease-free survival and overall survival; Supplementary Fig. 5c and Supplementary Table 4). Taken together, these data demonstrate that the upregulation of CSNK2A1 is a common feature of PTEN-deficient prostate tumours, emphasizing the need to target CK2 for cancer prevention and therapy in these tumours.
CK2 inhibition enhances senescence by stabilizing Pml level
We next sought to characterize the mechanism responsible for the increased senescence in Pten−/− MEFs upon CK2 inhibition. Recent evidence suggests that CK2 control the ubiquitin mediated degradation of PML, a known regulator of senescence in cancer34,35. Given the high CK2 levels in Pten−/− cells, we therefore checked Pml protein levels and found that Pml was decreased in Pten−/− cells as compared to the Ptenfl/fl cells (Fig. 3a and Supplementary Fig. 6a). Interestingly, treatment of Pten−/− cells with quinalizarin24 increased the protein levels of Pml, the SA-β-gal staining and enhanced the average size of Pml nuclear bodies (Fig. 3b,c). Next, we assessed Pml stability in Pten−/− MEFs and found that Pml half-life was significantly decreased as compared to the Ptenfl/fl MEFs (Fig. 3d). However, inhibition of CK2 by quinalizarin increased the stability of Pml protein in Pten−/− cells treated with cycloheximide, restoring Pml and p53 levels (Fig. 3e). Note that under cycloheximide and quinalizarin treatment, the mRNA levels of Pml did not change (Supplementary Fig. 6b). Along this line, when Ptenfl/fl and Pten−/− cells were treated with the proteasome inhibitor MG132, the protein levels of Pml increased, demonstrating that Pml is degraded through the proteasome as previously reported34 (Supplementary Fig. 6c). Importantly, Pml knocked down by means of short interfering RNAs (siRNAs) blunted the senescence response in Pten−/− MEFs (Fig. 3f). Moreover, treatment with the CK2 inhibitor, quinalizarin, in cells transfected with different siPML failed to induce any pro-senescence effect in Pten null cells (Fig. 3f and Supplementary Fig. 6e,f).
(a) Western blot analysis of prostate tissues for Pml in Ptenfl/fl and Pten−/− MEFs. (b) Western blot analysis for Pml protein levels (upper panel) and quantification (lower panel) in Ptenfl/fl and Pten−/− MEFs before and after treatment with the CK2i-Q. Representative images and quantification of SA-β-gal positivity in MEFs treated with vehicle and CK2i-Q. (c) Immunofluorescence staining for Pml in Ptenfl/fl and Pten−/− MEFs before and after treatment with the CK2i-Q. Quantification of the Pml bodies average size and s.d. is included in the panels. Scale bar, 5μM (d) Western blot analysis (upper panel) and quantification (lower panel) of Pml protein levels in primary Ptenfl/fl and Pten−/− MEFs after treatment with cycloheximide at different time points (0, 2, 4, 6 h) (e) Western blot analysis for PML and p53 in primary Ptenfl/fl and Pten−/− MEFs treated with both CK2i and cycloheximide at two different time points 4 and 6 h. (f) WB for Pml and Pten (upper panel) and SA-β-gal quantification for Ptenfl/fl and Pten−/− MEFs transfected with a siPml and treated with the CK2i-Q. WBs have been run on two different gels (g) IHC staining and quantification for PML, Ki-67 and pAKTSer129 in both PIN and invasive areas from Ptenpc−/− tumours at 15 weeks of age. Six samples per group were analysed. Invasive areas were defined using the following criteria: high percentage of Ki-67 staining and invasion of the stroma by the tumour epithelial cells. Scale bar, 50 μM (h) Percentage of mice affected with invasive prostate cancer in Ptenpc−/− tumours at different stages (n=10). (i) Western blot analysis for Pml and Pten in Ptenpc−/− tumours at different stages (1, 4 and 6 months). Ptenpc+/+ normal prostate at 6 months is used as control (j) IHC staining (left panel) and quantification (right panel) of p16-positive cells in Ptenpc−/− tumours at different stages (1, 4 and 6 months). Inset shows the negativity for p16 staining in Ptenpc+/+ normal prostate at 6 months. Scale bar: 50 μM. Images (b,e,f,g,h). Data are represented as mean±s.d. n≥3 (**P<0.01, ***P<0.001). P value indicates the statistical significance as measured by Student’s t-test. H&E, haematoxylin and eosin; Pos., positive.
Taken together, these data suggest that CK2 upregulation in Pten−/− cells favours the degradation of Pml, enabling a fraction of cells to bypass PICS. Consistently, CK2 inhibition potentiates senescence driven by Pten loss by stabilizing Pml protein levels. Finally, we checked the levels of PML and PTEN in a panel of prostate cancer cell lines and found a direct correlation between the level of PTEN and PML (Supplementary Fig. 6g). This was also associated with an increase in CSNK2A1 mRNA levels in PTEN-deficient cells (Supplementary Fig. 6h).
We then checked in vivo whether CK2 upregulation in Ptenpc−/− tumours could drive attenuation of senescence in vivo. During an early stage of tumorigenesis, Ptenpc−/− tumours develop high-grade prostatic intraepithelial neoplasia, characterized by a strong senescence response. However, at late stage, Ptenpc−/− tumours progress into more aggressive and invasive tumours2. We therefore monitored CK2 activity and Pml levels in Pten null tumours at the onset of tumour invasiveness (mice at 15 weeks of age). Immunohistochemistry (IHC) revealed that within the Ptenpc−/− tumours the most invasive tumour glands were characterized by high CK2 activity (assessed by pAKTSer129), high Ki-67 staining and low Pml protein levels, whereas the less aggressive glands displayed high levels of Pml and both low CK2 activity and Ki-67 positivity (Fig. 3g). A further time-course analysis in Ptenpc−/− tumours at different stages, demonstrated that Pml levels progressively decreased during tumour progression (Fig. 3h,i). Moreover, decreased Pml levels correlated with a reduced percentage of p16-positive cells in the tumours and an increased tumour invasiveness (Fig. 3j). These data suggest that activation of CK2 in Ptenpc−/− tumours drives a concomitant downregulation of Pml resulting in attenuation of senescence and promotion of tumour progression.
Inhibition of CK2 enhances senescence in Ptenpc−/− tumours
We next evaluated in a preclinical trial in vivo, whether the inhibition of CK2 could block tumour progression in Ptenpc−/− mice by stabilizing Pml protein levels and promoting senescence. To block CK2 in vivo, we used CX-4945, a novel CK2 inhibitor in clinical development25. CX-4945 was administered to Ptenpc–/– mice at 8 weeks of age for 5 weeks (Fig. 4a). Interestingly, CX-4945 treatment decreased both tumour size and the number of PIN-affected glands in all the mouse prostate lobes analysed (Fig. 4b). This was associated with decreased staining of Ki-67, pStat3Tyr705, pStat3Ser727 and increased p16 staining (Fig. 4c,d). Disappearance of pAKTSer129 staining in Ptenpc−/− tumours treated with CX-4945 was used as sign of impaired CK2 activity (Fig. 4c,d). Strikingly, Pml and p53 protein levels and SA-β-gal staining were also increased in Ptenpc–/– tumours treated with CX-4945 (Fig. 4e,f). Finally, staining by caspase-3 revealed no major difference (0.5 versus 1.5% of positive cells per section) in apoptosis between the treatment groups, suggesting that the main function of CX-4945 in Ptenpc–/– tumours is to enhance senescence rather than apoptosis (Fig. 4g). These findings substantiate the pro-senescence potential of CX-4945 in a genetically engineered mouse prostate model and identify a novel effective pro-senescence compound for prostate cancer therapy. Our findings also point to CK2-induced PML downregulation as a prominent mechanism of senescence attenuation and tumour progression in Pten null tumours. This is particularly noteworthy for human prostate cancer, where p53 function often remains functional even in advanced tumours, whereas PML levels are often found decreased36.
(a) Schedule of treatment used in the preclinical trial with the timing of the drug administration in the Ptenpc−/− mice (n=12). (b) Quantification (left panel) of relative tumour size (APs) and percentage of PIN-affected glands in the anterior (AP), ventral (VP) and dorsolateral (DLPs) prostates of Ptenpc−/− mice in the two treatment groups. (c) Histopathological analysis of Ptenpc−/− prostate tumours before and after treatment with CK2i-CX and stained as indicated: H&E, Ki-67, p16, Pml, pAKTSer129, pStat3Tyr705, pStat3Ser727 and CK2. Original magnification, × 40; insets show the × 10 magnification. Scale bar, 50 μM. (d) Quantification of c. Sections from three mice were counted for each treatment group. (e) Representative western blot analysis for Pml and p53 protein levels in Ptenpc−/− tumours treated with vehicle or CK2i-CX and quantification of protein levels. (f) SA-β-gal staining of Ptenpc−/− tumours from vehicle and CK2i-treated mice and the percentage of SA-β-gal positive cells is represented as a bar graph. (g) IHC staining and quantification of cleaved caspase-3 in Ptenpc−/− tumours before and after treatment with CK2i-CX. Imaged (b,d,g) data are represented as mean±s.d. n≥3 (*P<0.05,**P<0.01, ***P<0.001). P value indicates the statistical significance as measured by Student’s t-test. H&E, haematoxylin and eosin; Pos., positive.
Discussion
Since the first discovery that senescence could be enhanced in cancer cells, great effort has been made to identify novel pro-senescence compounds for cancer therapy. However, few small molecules have been identified so far that enhances senescence in tumours. Moreover, the majority of these molecules do not show selectivity for tumour cells and the risk of toxicity for cancer patients remains high. These targeted compounds include inhibitors of MDM2, MYC or CDK4, genes frequently deregulated in cancer, which also play a central role for the functional integrity of normal cells3. Senescence enhancement in normal cells may therefore increase the risk of organ toxicity in patients affected by cancer. Thus, identifying small-molecule inhibitors that increase senescence in cancer cells without affecting normal cells is an attractive strategy for cancer therapy. Senescence is a stable cell growth arrest which promotes tumour inhibition. Moreover, recent findings demonstrate that senescent tumour cells can be recognized and removed from cancerous tissues by components of both the adaptive and the innate immunity37,38. Indeed, senescent tumours cells are characterized by a senescence-associated secretory phenotype that can recruit and activate the tumour immune response37,39. Therefore, in stark contrast with current therapies (for example, chemotherapy) that mainly induce an apoptotic response, an effective pro-senescence therapy is expected to induce both a potent cell growth arrest and a specific antitumour immune response, thereby resulting in immune mediated tumour clearance. Here, we report the results of a screening strategy developed to identify pro-senescence compounds acting on a specific and relevant genetically defined subset of cancers, such as PTEN null tumours. The validity of this approach is supported by the effective identification of drug–genotype associations already established in previous preclinical models26,40,41 and from the identification of CK2 inhibitors that show high selectivity in PTEN-deficient cells and tumours. Our screening approach coupling a small-molecule screening with a shRNA screen has allowed us to focus on a target that has been identified in both the screens, using complementary methodologies. Given that the tumour immune response is an essential component of the tumour suppressive function of senescence in cancer, these compounds could be tested in the clinic either alone or in combination with compounds that activate the tumour immune response or reprogramme the senescence-associated secretory phenotype as recently demonstrated from our group7,42. In this respect, we envisioned that our screening platform may also be implemented with different immune assays. For example, the conditioned media obtained from MEFs treated with selected pro-senescence compounds might be transferred on different immune cell subtypes to assess their immunostimulatory capabilities.
Mechanistically, our findings have also identified a previously unknown STAT3–CK2–PML network that can be targeted to enhance senescence in cancer. While CK2 acts by promoting tumorigenesis in different settings, its role as senescence regulator in PTEN-deficient tumours remained elusive. It has been shown that CK2 controls proliferation and apoptosis by promoting the phosphorylation of several proteins including p53 (ref. 43). Intriguingly, although CK2 is upregulated in several cancers, the mechanism underlying such upregulation was ambiguous. In this respect, our findings demonstrate that activity of the JAK/STAT3 pathway, one of the most commonly deregulated pathways in cancer, resulted in CK2 overexpression, thus providing one of the mechanisms by which CK2 level is increased in cancer. Another intriguing finding of this study is the discovery that Pten null tumours at the late stage of tumorigenesis exhibit decreased levels of the tumour suppressor Pml, an essential regulator of oncogene induced senescence5. Previous evidence demonstrates that the Ptenpc−/− mice develop senescence in tumours at an early stage; however, the majority of these tumours eventually progress to invasive prostate cancer bypassing the initial senescence response driven by Pten loss. Our time-course analysis in Pten null tumours analysed at different stages demonstrates that Pml levels decrease during tumour progression in parallel with p16 staining positivity a marker of senescence. Importantly, we also detected increased levels of pAKTSer129 staining, a marker of CK2 kinase activation in tumour regions with low level of Pml. This is consistent with a model in which CK2 upregulation and Pml downregulation contribute to senescence evasion in Pten null tumours. Thus, in agreement with a model of intratumor heterogeneity, Pten loss promotes a senescence response that opposes tumour progression. However, a subset of Pten null cells upregulates CK2 that in turn promotes the evasion of cellular senescence by decreasing the level of Pml (Supplementary Fig. 6i). We believe that this mechanism may further contribute to the senescence evasion mediated by tumour-infiltrating myeloid cells in Pten null tumours7. Therefore, both cell autonomous and non-autonomous mechanisms can promote senescence evasion in tumours, suggesting that to design more efficient pro-senescence therapies for cancer, novel compounds should target both cell intrinsic and extrinsic factors.
The existence of this novel PTEN–STAT3–CK2 network has also been validated in patients affected by prostate cancers through a bioinformatics analysis. Although we cannot rule out the possibility that additional genes may cooperate with PTEN loss to drive tumour progression, and demonstrate that the inhibition of CK2 is highly effective in blocking tumour development by enhancing senescence. Our results also indicate the need to monitor PTEN and CK2 status in clinical studies and highlight the importance of testing CK2 inhibitors alone or in combination with standard therapies in cancer patients. These drugs have acceptable toxicity profiles and are currently in clinical trials for different types of cancer25,44,45. In sum, our findings provide new insight into the mechanisms that regulate senescence evasion and encourage further efforts to screen for small molecules that could prove useful for pro-senescence therapy for cancer.
Methods
Mouse crosses and MEF production
Ptenloxp/loxp mice were crossed with PB-Cre4 transgenic mice to generate prostate-specific deletion of Pten as previously reported1. MEFs were obtained by crossing male Ptenloxp/loxp mice with female Ptenloxp/loxp female. A pregnant mouse at 13 or 14 day post coitum was sacrificed by cervical dislocation. Embryos were harvested and the individual MEFs were cultured in DMEM containing 10% fetal bovine serum and 1% PenStrep. Primary Ptenlox/lox MEFs were infected with retroviruses expressing either pMSCV-CRE-PURO-IRES-GFP or pMSCV-PURO-IRES-GFP for 48 h and selected with Puromycin at a concentration of 3 μg ml−1 and as previously described1. All mice were maintained under specific pathogen-free conditions in the animal facilities of the IRB institute, and the experiments were performed according to the state guidelines and approved by the local ethical committee.
Human cancer cell lines
Human prostate cancer cell lines (Supplementary Table 3) were obtained from ATCC and were cultured according to manufacturer instructions. HCT116 isogenic cell line was a gift from Waldman lab.
Pro-senescence screening platform
The screening has been carried out in three steps. In the first step of the screening, compounds have been tested in triplicate in Pten−/− MEFs, plated in 96-well dishes, using a single drug concentration (10 μM). Compounds that promoted a strong growth arrest in Pten null MEFs (inhibition of proliferation ≥50%) have been considered positive hits and passed the first filter of the screening (efficacy filter). Positive hits were retested in triplicate at a single concentration (10 μM) in both Ptenfl/fl and Pten−/− MEFs to assess growth arrest in both cell types (specificity filter). Compounds that pass the cell_specificity filter (inhibition of proliferation for Ptenwt≤20% and for Pten−/− ≥50%) were validated in duplicate in a dose–response assay in MEFs plated in 96-well dishes, to evaluate specificity at different concentrations (0.03–100 μM). Only the compounds having an optimal window of inhibition passed this filter (dose optimization filter). Finally, the compounds that passed all the filters were evaluated for SA-β-gal staining at three different concentrations in MEFs plated in 12-well dishes. Data shown refer to a single concentration (compound IC50). Data were normalized according to the negative control (dimethyl sulfoxide 100%). Nutlin-3, a previously characterized pro-senescence compound26 was used as a positive control. Concentration response was analysed with XLfit (IDBS) by using four parameters logistic model.
Proliferation and senescence assay
Four days after treatment, MEFs were fixed for 20 min in a solution of buffered formalin (3.7%), washed with PBS (pH 7.3) and subsequently stained with a 0.01% crystal violet solution. After removing excess stain, the crystal violet stained cells were dissolved in 1 ml of 10% SDS solution overnight and the optical density of the extracted dye was read with a spectrophotometer at 590 nm. In the senescence assay, cells were stained for SA-β-gal activity, four days after treatment, using a senescence detection kit (Calbiochem). The number of blue stained cells were counted and normalized to the DAPI-positive cells.
Generation of lentiviral shRNA kinome library
An shRNA library in the pLKO.1 vector backbone comprising 3,280 shRNAs targeting 672 mouse kinases was obtained from Sigma. A pooled DNA preparation of the library was generated by growing bacterial cultures in deep-well 96-well plates inoculated from 96-well format bacterial glycerol stocks of the library. Culture medium was pooled and a midi-prep (Machery Nagel Nucleobond 500 kit) was performed to isolate plasmid DNA. Lentiviruses were produced by transfection of 293T cells with the pooled library DNA together with psPAX2 and pMD2G (gift from Didier Trono, EPFL Lausanne) and harvesting of culture supernatant between 48 and 72 h after transfection. Medium was filtered through a 0.45-μm filter and aliquots were frozen at −80 °C. Virus titre was determined by infecting NIH3T3 cells with serial dilutions of the virus and assaying colony formation in the presence of puromycin (4 μg ml−1).
Kinome library screening
Primary MEFs (5 × 106) derived from Ptenfl/fl mice were cultured in an incubator maintained at 5% CO2 and infected with the pooled mouse kinome lentiviral library at a multiplicity of infection of one functional viral particle per cell in the presence of polybrene (8 μg ml−1). Two days later, cells were split in three into medium containing puromycin (4 μg ml−1) alone to select for virally infected cells or puromycin together with Adenovirus expressing GFP (Vector Biolabs, 1060) or Cre-GFP (Vector Biolabs, 1700). Genomic DNA from the non-Adenovirus-infected cells after two further days of culture was isolated by incubation for 4 h with lysis buffer (200 mM NaCl, 50 mM Tris, 10 mM EDTA, 1% SDS) containing proteinase K (500 μg ml−1) and RNase A (83 μg ml−1) at 58 °C, followed by precipitation with NaCl/isopropanol and washing with 70% ethanol. The Adenovirus-infected cells were passaged every 3 days for a further 16 days before genomic DNA isolation. To maintain representation of the shRNA library in the population, at least 5 × 106 cells were utilized for genomic DNA isolation. To amplify the integrated shRNAs from the genomic DNA, PCR was conducted using a common forward primer containing sequencing adaptors and one of three reverse primers containing a barcode sequence specific for the experimental condition, as well as sequencing adapters. Forward primer 5′–3′ (bold highlights sequence of homology to pLKO.1):
5′-AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCTATCTTGTGGAAAGGACGA-3′
Reverse primers 5′–3′ (bold highlights Illumina Index six-nucleotide barcode sequences and sequence of homology to pLKO.1):
5′-CAAGCAGAAGACGGCATACGAGATATCACGGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCTATTCTTTCCCCTGCACT-3′
5′-CAAGCAGAAGACGGCATACGAGATTGGTCAGTGACTGGAGTTCAG ACGTGTGCTCTTCCGATCTATTCTTTCCCCTGCACT-3′
5′-CAAGCAGAAGACGGCATACGAGATCACTGTGTGACTGGAGTTCAGACGTGTGCTCTTCCGATCTATTCTTTCCCCTGCACT-3′
Genomic DNA (15 ng) was used as PCR template in a 50-μl reaction containing 2.5 nmol of each primer, 5 U of Dream Taq (Thermo Scientific), 5 nmol dNTPs and 34 cycles at annealing temperature 56 °C and extension temperature 72 °C. The amplified PCR product was purified by excision from an agarose gel and purification using NucleoSpin Gel and PCR Clean-up kit (Macherey-Nagel) and subjected to 100 bp paired-end deep sequencing using an Illumina HiSeq 2000 machine. Bioinformatic analysis was used to identify each experiment using the unique barcodes and to map each individual sequencing read from the experiment to an individual hairpin sequence. For identification of experimental hits, shRNAs were first filtered for those represented by at least 200 reads at the early time point.
Western blotting and histology
Protein was isolated from whole MEFs cell pellets lysed with RIPA buffer and BCA protein assay reagent was used to determine the protein concentration. The following antibodies were used for western blotting: p21, p16, PAI-1 (Santa Cruz), CK2α, Pten, Hsp90 (Cell Signalling), PML (Millipore), p53 (Clone IMX25, Accurate Chemical Corp), pAKTSer129 was produced in rabbit30, β-actin (Sigma) and E-cadherin (BD Biosciences). Secondary antibodies for mouse and rabbit were purchased from GE Healthcare. All dilutions are 1:1,000, except that β-actin antibody dilution is 1:5,000 and Secondary antibody dilution is 1:5,000. Full gel scans are shown in Supplementary Fig. 7. For IHC and immunofluorescence, the tissues were embedded in 37% formaldehyde and embedded in paraffin for sectioning.
CK2 shRNAs and PML siRNA transfection protocol
Mouse CK2 pLKO shRNA vectors were obtained from Sigma. Using 293T HEKs, lentiviral particles were produced for negative control or shRNA vectors. MEFs were harvested after 72 h of infection for western blotting and SA-β-gal activity. shRNA sequences: 5′-CCGGACCTGTCAGCAGCGCCAATATCTCGAGATATTGGCGCTGCTGACAGGTTTTTTG-3′ (Clone ID:NM_007788.3-1293s21c1) and 5′-CCGGGCAATTGTACCAGACGTTAACCTCGAGGTTAACGTCTGGTACAATTGCTTTTTG-3′ (Clone ID:NM_007788.3-627s21c1). For the experiment with the PML siRNA, MEFs were transfected 1 day after selection using a smart pool PML siRNA (GE Healthcare, Dharmacon) and sequences of the siRNA targeting Pml from Qiagen used are 5′-CTGGATGTTCGATTCATGTAA-3′ and 5′-CCCAGCATATCTACTCCTTTA-3′.
Luciferase reporter gene assay
The promoter region of CSNK2A1 between −500 bp to +120 bp was cloned into a pGL4.24 luc2 vector to generate the CSNK2A1 reporter construct. Before transfection, immortalized MEFs were plated in 48-well plates and were co-transfected with CSNK2A1 reporter construct and a control pGL4.27 renilla TK vector in triplicates. The ratio of reporter to control plasmid was 10:1. After 48 h of transient transfection, cell lysates were transferred to 96-well plates and dual luminescence was measured using a Glo-Max Luminometer (Promega). Different CSNK2A1 mutants were generated by PCR mutagenesis. Primers used are shown in Supplementary Table 5.
Chromatin immunoprecipitation assay
Immortalized MEFs were grown in T-75 flasks to a confluence of 90–95% and cells were fixed with 37% formaldehyde for 10 min and glycine for 5 min at room temperature. Nuclear extracts were sonicated using a Misonix 3,000 model sonicator to sheer crosslinked DNA to an average fragment size of ∼500 bp. Sonicated chromatin was incubated for 16 h at 4 °C with magnetic protein A beads (Millipore) conjugated with either anti-Stat3 antibody (Cell Signalling) or mouse IgG antibody (Millipore) by incubating overnight at 4 °C on a rotor. After incubation, beads were washed thoroughly and then centrifuged. The chromatin was eluted from the beads, and crosslinks were removed by incubation at 65 °C for 5 h. DNA was then purified using the QIAquick PCR Purification Kit (Qiagen). DNA was used to detect Stat3 binding to CK2 promoter sequences by qRT-PCR. The ChIP primers (5′-CAGCTGACAAACTGTAAAAGGGG-3′ and 5′-TGGGTTATTTCTGTACCAGCCATT-3′) were designed to amplify a region containing the putative Stat3-binding site (−1,228/−1,122 bp) in the mouse CK2 promoter As an internal negative control, the primers (5′-ATCCAGGATTATCAGGTTCAATTT-3′ and 5′-AGAGGTCTATGTCTGGGGGTGTAG-3′) were designed such that there is no putative Stat3-binding site (−2,184 to −2,083 bp). Primers pair used for positive control Socs3 are 5′-CCCTTGCAGTTCTAAGTTCAACA-3′ (forward) and 5′-ACCTTTGACAAGCGGACTCTC-3′ (reverse).
Quantitative real-time PCR
Quantitative real-time PCR was performed on RNA extracted from MEFs and prostatic tissues by using Trizol (Invitrogen) in triplicates. Complementary DNA was prepared with SuperScript III First-Strand Synthesis SuperMix (Invitrogen). Primers sequences used for p21, PAI-1, STAT3 and p16 are reported previously42 and primers used for Pml forward 5′-CAGGCCCTAGAGCTGTCTAAG-3′, reverse 5′-ATACACTGGTACAGGGTGTGC-3′; Csnk2α forward 5′-AGGATAGCCAAGGTTCTGGGA-3′, reverse 5′-CCATCGCTTACGGGAGTGTC-3′; CSNK2α forward 5′-GGTGGAATGGGGAAATCAAGAT-3′, reverse 5′-TGATGATGTTGGGACCTCCTC-3′.
Half-life determination and proteasome inhibitor
MEFs were treated at 20 μg ml−1 concentration of cycloheximide and harvested at different time points for the analysis by western blotting. Proteosomal inhibitor MG132 was used at 10 μM in MEFs.
PTEN and CK2 mRNA levels in human prostate cancer data sets
Expression values for PTEN, CK2 and STAT3 were downloaded for Wallace46 and Singh47 prostate cancer data sets from Oncomine database48. Expression values for PTEN and CK2, as well as clinical information and follow-up was downloaded for Setlur49 and Glinsky50 data sets from Oncomine database. PTEN/CK2 mRNA ratio was calculated per each sample and patients were stratified into three groups by dividing the dataset into four percentiles (from highest to lowest value). Patients with lowest ratio values (1st percentile) were ascribed to group low (L), patients with intermediate values (2nd and 3rd percentiles) were ascribed to group intermediate (I) and patients with highest values (4th percentile) were ascribed to group high (H). Statistical analysis such as t-test, Pearson correlation and Kaplan–Meier curves were done using Stata 12.1 software.
Differential expression of TFs
Expression values for TFs in Ptenpc+/+ normal prostates and Ptenpc−/− tumours was downloaded from Gene Expression Omnibus database under the GSE24691 identifier, and the data have been previously reported51. t-test analysis was done to extract TFs with significant differential expression between mouse genotypes.
Treatment with CK2 inhibitors
CK2 inhibitors used were CK2i-Q quinalizarin (Sigma Aldrich), CK2i-CX (CX-4945 (Selleckchem)), K156 and K164 (ref. 15). CK2i-S (obtained in collaboration with Siena Biotech spa). CX-4945 sodium salt inhibitor administered in vivo treatment was synthesized by Shanghai Biochempartner Co. Ltd. Ptenpc−/− mice at 8 weeks of age were treated by oral gavage with 75 mg kg−1 of CX-4945 or vehicle twice daily. CX-4945 was diluted in a solution with 25 mM NaH2PO4 buffer. After 4 weeks of treatment, prostate tumours were assessed for the size, number of PIN-affected gland and IHC.
Functional annotation of targets and pro-senescence compounds
All targets of the pro-senescence compounds were converted to HGNC compliant symbols and functionally annotated with experimentally biological processes comprised in gene ontology (GO; 14.10.2014) and Reactome (18.10.2014) using ClueGO 2.1.4 (ref. 52) and CluePedia 1.1.4 (ref. 52) within Cytoscape 3.1.1 (ref. 53). ClueGO analysis settings were set as follows: (i) Hierarchy levels=4 to 20, (ii) number of targets member of a biological process for the term to be mapped ≥1, (iii) percentage of a term covered by the targets ≥3%, (iv) GO-term fusion was applied and GO-term-grouping based on kappa score of 0.4 (similarity measure between the biological processes). For each group of terms and for terms that could not be grouped, the biological process that accommodated the largest number of targets was selected as overview term. All other settings were used as per default. Senescence pathways were extracted from a total of 25,517 gene sets (GSs) (http://download.baderlab.org/EM_Genesets/May_14_2014/Human/symbol/Human_GO_AllPathways_with_GO_iea_May_14_2014_symbol.gmt; 14.05.2014 and all curated GSs comprised in MSigDB v4.0 http://www.broadinstitute.org) by filtering the GSs’ names for senescence and senescent. Any senescence pathway that had targets of a pro-senescence compound as members was mapped on the network. Thus, the network maps all pro-senescence compounds, the targets they inhibit, and the biological processes and senescence pathway.
Statistical analysis
Data analysis was performed using a two-tailed unpaired Student’s t-test. Data are represented as mean±s.d.
Additional information
How to cite this article: Kalathur, M. et al. A chemogenomic screening identifies CK2 as a target for pro-senescence therapy in PTEN-deficient tumours. Nat. Commun. 6:7227 doi: 10.1038/ncomms8227 (2015).
References
Alimonti, A. et al. Subtle variations in Pten dose determine cancer susceptibility. Nat. Genet. 42, 454–458 (2010) .
Chen, Z. et al. Crucial role of p53-dependent cellular senescence in suppression of Pten-deficient tumorigenesis. Nature 436, 725–730 (2005) .
Nardella, C., Clohessy, J. G., Alimonti, A. & Pandolfi, P. P. Pro-senescence therapy for cancer treatment. Nat. Rev. Cancer 11, 503–511 (2011) .
Xue, W. et al. Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature 445, 656–660 (2007) .
Pearson, M. et al. PML regulates p53 acetylation and premature senescence induced by oncogenic Ras. Nature 406, 207–210 (2000) .
Guengerich, F. P. Mechanisms of drug toxicity and relevance to pharmaceutical development. Drug Metab Pharmacokinet. 26, 3–14 (2011) .
Di Mitri, D. et al. Tumour-infiltrating Gr-1+ myeloid cells antagonize senescence in cancer. Nature 515, 134–137 (2014) .
Liu, H., Li, Y. & Raisch, K. P. Clotrimazole induces a late G1 cell cycle arrest and sensitizes glioblastoma cells to radiation in vitro. Anticancer Drugs 21, 841–849 (2010) .
Furtado, C. M., Marcondes, M. C., Sola-Penna, M., de Souza, M. L. & Zancan, P. Clotrimazole preferentially inhibits human breast cancer cell proliferation, viability and glycolysis. PLoS ONE 7, e30462 (2012) .
Bilir, A. et al. Enhancement of vinorelbine-induced cytotoxicity and apoptosis by clomipramine and lithium chloride in human neuroblastoma cancer cell line SH-SY5Y. J. Neurooncol. 100, 385–395 (2010) .
Yang, J. L. & Friedlander, M. L. Effect of nifedipine in metastatic colon cancer with DNA mismatch repair gene defect. Lancet 357, 1767–1768 (2001) .
Meng, F., Sun, G., Zhong, M., Yu, Y. & Brewer, M. A. Anticancer efficacy of cisplatin and trichostatin A or 5-aza-2'-deoxycytidine on ovarian cancer. Br. J. Cancer 108, 579–586 (2013) .
Vigushin, D. M. et al. Trichostatin A is a histone deacetylase inhibitor with potent antitumor activity against breast cancer in vivo. Clin. Cancer Res. 7, 971–976 (2001) .
Li, F. et al. Garcinol sensitizes human head and neck carcinoma to cisplatin in a xenograft mouse model despite downregulation of proliferative biomarkers. Oncotarget 6, 5147–5163 (2015) .
Cozza, G. et al. Cell-permeable dual inhibitors of protein kinases CK2 and PIM-1: structural features and pharmacological potential. Cell. Mol. Life Sci. 71, 3173–3185 (2014) .
Martins, L. R. et al. Activity of the clinical-stage CK2-specific inhibitor CX-4945 against chronic lymphocytic leukemia. Leukemia 28, 179–182 (2014) .
Mateo, A. A.. et al. Synthesis of phenazine derivatives for use as precursors to electrochemically generated bases. A. Org. Biomol. Chem. 3, 2832–2841 (2005) .
Roberson, R. S., Kussick, S. J., Vallieres, E., Chen, S. Y. & Wu, D. Y. Escape from therapy-induced accelerated cellular senescence in p53-null lung cancer cells and in human lung cancers. Cancer Res. 65, 2795–2803 (2005) .
Dabrowska, M., Mosieniak, G., Skierski, J., Sikora, E. & Rode, W. Methotrexate-induced senescence in human adenocarcinoma cells is accompanied by induction of p21(waf1/cip1) expression and lack of polyploidy. Cancer Lett. 284, 95–101 (2009) .
Orsini, M. J. et al. Metastin (KiSS-1) mimetics identified from peptide structure-activity relationship-derived pharmacophores and directed small molecule database screening. J. Med. Chem. 50, 462–471 (2007) .
Yip, K. W. et al. Benzethonium chloride: a novel anticancer agent identified by using a cell-based small-molecule screen. Clin. Cancer Res. 12, 5557–5569 (2006) .
Chien, W. et al. Activation of protein phosphatase 2A tumor suppressor as potential treatment of pancreatic cancer. Mol. Oncol. 9, 889–905 (2015) .
Sengupta, T. K. et al. Cytotoxic effect of 5-aminoimidazole-4-carboxamide-1-beta-4-ribofuranoside (AICAR) on childhood acute lymphoblastic leukemia (ALL) cells: implication for targeted therapy. Mol. Cancer 6, 46 (2007) .
Cozza, G. et al. Quinalizarin as a potent, selective and cell-permeable inhibitor of protein kinase CK2. Biochem. J. 421, 387–395 (2009) .
Siddiqui-Jain, A. et al. CX-4945, an orally bioavailable selective inhibitor of protein kinase CK2, inhibits prosurvival and angiogenic signaling and exhibits antitumor efficacy. Cancer Res. 70, 10288–10298 (2010) .
Alimonti, A. et al. A novel type of cellular senescence that can be enhanced in mouse models and human tumor xenografts to suppress prostate tumorigenesis. J. Clin. Invest. 120, 681–693 (2010) .
Kim, J. S. et al. Mechanistic analysis of a DNA damage-induced, PTEN-dependent size checkpoint in human cells. Mol. Cell. Biol. 31, 2756–2771 (2011) .
Niefind, K., Guerra, B., Ermakowa, I. & Issinger, O. G. Crystal structure of human protein kinase CK2: insights into basic properties of the CK2 holoenzyme. EMBO J. 20, 5320–5331 (2001) .
Trembley, J. H., Wang, G., Unger, G., Slaton, J. & Ahmed, K. Protein kinase CK2 in health and disease: CK2: a key player in cancer biology. Cell. Mol. Life Sci. 66, 1858–1867 (2009) .
Di Maira, G. et al. Protein kinase CK2 phosphorylates and upregulates Akt/PKB. Cell Death Differ. 12, 668–677 (2005) .
Miklossy, G., Hilliard, T. S. & Turkson, J. Therapeutic modulators of STAT signalling for human diseases. Nat. Rev. Drug Discov. 12, 611–629 (2013) .
Baffert, F. et al. Potent and selective inhibition of polycythemia by the quinoxaline JAK2 inhibitor NVP-BSK805. Mol. Cancer Ther. 9, 1945–1955 (2010) .
Yang, J. et al. Reversible methylation of promoter-bound STAT3 by histone-modifying enzymes. Proc. Natl Acad. Sci. USA 107, 21499–21504 (2010) .
Scaglioni, P. P. et al. A CK2-dependent mechanism for degradation of the PML tumor suppressor. Cell 126, 269–283 (2006) .
Scaglioni, P. P. et al. CK2 mediates phosphorylation and ubiquitin-mediated degradation of the PML tumor suppressor. Mol. Cell. Biochem. 316, 149–154 (2008) .
Gurrieri, C. et al. Loss of the tumor suppressor PML in human cancers of multiple histologic origins. J. Natl Cancer Inst. 96, 269–279 (2004) .
Kang, T. W. et al. Senescence surveillance of pre-malignant hepatocytes limits liver cancer development. Nature 479, 547–551 (2011) .
Braumuller, H. et al. T-helper-1-cell cytokines drive cancer into senescence. Nature 494, 361–365 (2013) .
Hemann, M. T. et al. Evasion of the p53 tumour surveillance network by tumour-derived MYC mutants. Nature 436, 807–811 (2005) .
Tovar, C. et al. MDM2 small-molecule antagonist RG7112 activates p53 signaling and regresses human tumors in preclinical cancer models. Cancer Res. 73, 2587–2597 (2013) .
Rader, J. et al. Dual CDK4/CDK6 inhibition induces cell-cycle arrest and senescence in neuroblastoma. Clin. Cancer Res. 19, 6173–6182 (2013) .
Toso, A. et al. Enhancing chemotherapy efficacy in pten-deficient prostate tumors by activating the senescence-associated antitumor immunity. Cell Rep. 9, 75–89 (2014) .
Filhol, O. & Cochet, C. Protein kinase CK2 in health and disease: Cellular functions of protein kinase CK2: a dynamic affair. Cell. Mol. Life Sci. 66, 1830–1839 (2009) .
Kim, J. & Hwan Kim, S. CK2 inhibitor CX-4945 blocks TGF-beta1-induced epithelial-to-mesenchymal transition in A549 human lung adenocarcinoma cells. PLoS ONE 8, e74342 (2013) .
Cozza, G., Pinna, L. A. & Moro, S. Protein kinase CK2 inhibitors: a patent review. Expert Opin. Ther. Pat. 22, 1081–1097 (2012) .
Wallace, T. A. et al. Tumor immunobiological differences in prostate cancer between African-American and European-American men. Cancer Res. 68, 927–936 (2008) .
Singh, D. et al. Gene expression correlates of clinical prostate cancer behavior. Cancer Cell 1, 203–209 (2002) .
Rhodes, D. R. et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 6, 1–6 (2004) .
Setlur, S. R. et al. Estrogen-dependent signaling in a molecularly distinct subclass of aggressive prostate cancer. J. Natl Cancer Inst. 100, 815–825 (2008) .
Glinsky, G. V., Glinskii, A. B., Stephenson, A. J., Hoffman, R. M. & Gerald, W. L. Gene expression profiling predicts clinical outcome of prostate cancer. J. Clin. Invest. 113, 913–923 (2004) .
Carver, B. S. et al. Reciprocal feedback regulation of PI3K and androgen receptor signaling in PTEN-deficient prostate cancer. Cancer Cell 19, 575–586 (2011) .
Bindea, G., Galon, J. & Mlecnik, B. CluePedia Cytoscape plugin: pathway insights using integrated experimental and in silico data. Bioinformatics 29, 661–663 (2013) .
Smoot, M. E., Ono, K., Ruscheinski, J., Wang, P. L. & Ideker, T. Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27, 431–432 (2011) .
Acknowledgements
We thank members of the IRB animal core facility for technical assistance and the animal work. We are grateful to Erica Montani for her help with microscopy. We thank all the members of Dr A. Alimonti laboratory and Dr Carlo Catapano for their scientific discussions. We thank Dr Ramon Parsons for being generous in providing the reagents. This work was supported by Swiss National Science Foundation (SNF) grant Ambizione (PZ00P3_136612/1), European Society for Medical Oncology (ESMO) translational research award, Swiss Bridge Award and European research council starting grant to A.A. (ERCsg 261342). I.J.F. was supported by (ERCsg 260316). The work of L.A.P. was supported by AIRC grant IG-14180.
Author information
Authors and Affiliations
Contributions
A.A., A.T., M.K. and I.F. originally developed the concept, further elaborated on it and designed the experiments. A.P., Le.M., La.M., F.M. and Al.T. performed selection of compounds for the screening campaign and analysed screening data. M.K., A.T., J.C., A.R. and C.D. performed experiments. R.G.E and R.K.R.K. performed bioinformatics analysis. M.S. and S.P. performed immunohistochemistry. L.A.P., M.R. and G.C. provided valuable reagents and technical advice. M.K., A.T., I.F. and A.A. interpreted the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-7, Supplementary Tables 1-5 and Supplementary Methods. (PDF 841 kb)
Rights and permissions
About this article
Cite this article
Kalathur, M., Toso, A., Chen, J. et al. A chemogenomic screening identifies CK2 as a target for pro-senescence therapy in PTEN-deficient tumours. Nat Commun 6, 7227 (2015). https://doi.org/10.1038/ncomms8227
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms8227
This article is cited by
-
Protein kinase CK2 – diverse roles in cancer cell biology and therapeutic promise
Molecular and Cellular Biochemistry (2023)
-
Bidirectional effects of intestinal microbiota and antibiotics: a new strategy for colorectal cancer treatment and prevention
Journal of Cancer Research and Clinical Oncology (2022)
-
Protein kinase CK2: a potential therapeutic target for diverse human diseases
Signal Transduction and Targeted Therapy (2021)
-
A new biological triangle in cancer: intestinal microbiota, immune checkpoint inhibitors and antibiotics
Clinical and Translational Oncology (2021)
-
Regulation of the oncogenic phenotype by the nuclear body protein ZC3H8
BMC Cancer (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.