Abstract
Pathogen effector proteins are delivered to host cells to suppress plant immunity. However, the mechanisms by which effector proteins function are largely unknown. Here we show that expression of XopPXoo, an effector of rice pathogen Xanthomonas oryzae pv. oryzae, in rice strongly suppresses peptidoglycan (PGN)- and chitin-triggered immunity and resistance to X. oryzae. XopPXoo targets OsPUB44, a rice ubiquitin E3 ligase with a unique U-box domain. We find that XopPXoo directly interacts with the OsPUB44 U-box domain and inhibits ligase activity. Two amino-acid residues specific for the OsPUB44 U-box domain are identified, which are responsible for the interaction with XopPXoo. Silencing of OsPUB44 suppresses PGN- and chitin-triggered immunity and X. oryzae resistance, indicating that OsPUB44 positively regulates immune responses. Thus, it is likely that XopPXoo suppresses immune responses by directly interacting with and inhibiting a positive regulator of plant immunity.
Similar content being viewed by others
Introduction
Immunity in plants is generally initiated with the recognition of the pathogen- or microbe-associated molecular patterns (PAMPs/MAMPs), including bacterial flagellin or peptidoglycan (PGN), and fungal chitin1,2,3,4. The recognition of PAMPs by pattern-recognition receptors (PRRs) triggers PAMP-triggered immunity (PTI). PTI constitutes the first layer of plant immunity that restricts pathogen proliferation. To interfere with host PTI, plant pathogens secrete a variety of effector proteins into host cells, allowing the pathogen to manipulate host physiology to promote pathogen growth5. Some of the effectors interact either directly or indirectly with the intracellular immune receptors of the nucleotide-binding leucine-rich repeats, triggering a second layer of immune responses often accompanied with hypersensitive cell death, which is referred to as effector-triggered immunity (ETI)1.
Ubiquitination is an important post-translational modification in eukaryotic cells and is involved in a variety of plant processes, such as hormone signalling, immunity, flowering, cell cycle, circadian rhythm control, growth and development6. The ubiquitination process involves sequential enzymatic activities. Ubiquitin is activated by a ubiquitin-activating enzyme (E1). The activated ubiquitin is then transferred to a ubiquitin-conjugating enzyme (E2). Ubiquitin ligase (E3) mediates ubiquitin transfer from an E2 to a target protein. Eukaryotic E3 ligases are classified into two major classes, RING (really interesting new gene) finger/U-box and HECT (homologous to E6-associated protein C-terminus), according to the mechanism of ubiquitin transfer6. RING finger/U-box proteins transfer ubiquitin directly from the E2 to the target proteins, whereas HECT proteins form thioester intermediates with ubiquitin before ligating it to the target. The U-box and RING finger domains are structurally similar, although the U-box lacks the characteristic zinc-chelating cysteine and histidine residues7.
Recent studies revealed that RING finger/U-box ubiquitin ligases are implicated in many aspects of plant immune responses8. Tobacco U-box E3 ubiquitin ligase CMPG1 regulates programmed cell death in ETI9. Arabidopsis AtPUB22, AtPUB23 and AtPUB24, closely related to CMPG1, negatively regulate immune reactions in response to multiple PAMPs10. More recently, AtPUB12 and AtPUB13 were shown to ubiquitinate FLS2, an Arabidopsis receptor for bacterial flagellin (flg22, a conserved 22-amino-acid (aa) peptide from flagellin), resulting in the attenuation of immune signalling by degradation of FLS2 (ref. 11). AtPUB17 is implicated in ETI mediated by two nucleotide-binding leucine-rich repeats RPM1 and RPS4 (ref. 12). A rice U-box protein SPL11, the closest homologue of AtPUB12 and AtPUB13, is a negative regulator of plant cell death13. Rice RING finger protein APIP6 positively regulates resistance to Magnaporthe oryzae14. Together, the accumulating evidence indicates that plant U-box (PUB)/RING finger proteins play central roles in plant immune responses. Recently, these U-box/RING finger proteins were reported to be targeted by pathogen effectors. CMPG1 and APIP6 are targeted by Phytophthora infestans effector AVR3a and M. oryzae effector AvrPiz-t, respectively14,15,16. Nevertheless, the molecular mechanisms describing how the effectors inhibit these E3 ligases remain to be identified.
Xanthomonas oryzae pv. oryzae (Xoo) causes bacterial blight of rice, one of the most important rice diseases in much of Asia17. Xoo uses the type III secretion system to secrete the effectors into rice cells. Furutani et al.18 identified 16 effectors in Xoo. Among them, XopYXoo (Xoo1488) was shown to strongly suppress host PTI in rice19. Recently, we reported that XopYXoo targets OsRLCK185, a rice receptor-like cytoplasmic kinase that interacts with OsCERK1, a PRR for chitin and PGN19. In response to PAMP recognition, OsCERK1 directly phosphorylates OsRLCK185 to transmit the immune signals. The OsCERK1-mediated phosphorylation of OsRLCK185 is suppressed by XopYXoo, resulting in the inhibition of immune responses including MAP kinase activation19,20.
Expression of the Xoo effector XopAAXoo (Xoo2875) in rice cells also inhibits PTI21. XopAAXoo targets OsBAK1, a co-receptor for many PRRs, and OsBRI1, a brassinosteroid receptor, as supported by the observation that transgenic plants expressing XopAAXoo exhibit a brassinosteroid-insensitive phenotype21. In addition, XopN, XopR and XopX also have been shown to suppress host immune responses in plants22,23,24. The type III effectors from other Xanthomonas genera have been well documented to suppress host immune responses. AvrAC suppresses FLS2-mediated immune responses by transferring uridyl to BIK1, a RLCK family kinase and an essential immediate signalling partner of FLS2 (ref. 25). XopD represses host ethylene responses by desumoylating SIERF4, a tomato transcription factor26. XopD also targets Arabidopsis transcription factor MYB30 (ref. 27). XopN interacts with a tomato atypical receptor-like kinase and TFT1 (ref. 28). XopJ interacts with the proteasomal subunit RPT6 to inhibit proteasome activity29. Thus, these recent investigations have revealed that the effectors target key components controlling host immune responses in plants. Therefore, it is likely that identifying the host targets for the effectors will facilitate a more complete molecular understanding of how pathogens inhibit host immune responses.
In this study, we found that XopPXoo, one of the Xoo effectors, suppresses host resistance to Xoo, and PGN- and chitin-induced immune responses, and identified a U-box-type E3 ubiquitin ligase, OsPUB44, as an interacting protein with XopPXoo. Silencing of OsPUB44 reduces PGN- and chitin-induced immunity and resistance to Xoo, suggesting that OsPUB44 positively regulates immune responses in rice. XopPXoo interactes with the U-box domain of OsPUB44; however, XopPXoo is not able to interact with other PUB proteins closely related to OsPUB44. The limited interaction between XopPXoo and OsPUB44 is dependent on two amino-acid residues specific for the U-box domain of OsPUB44. The interaction of the U-box domain with XopPXoo results in suppression of the OsPUB44 ubiquitin ligase activity. These results indicate that XopPXoo modulates the OsPUB44 ligase activity by direct association with the U-box domain for Xoo virulence in rice.
Results
XopPXoo suppresses the PTI response in rice
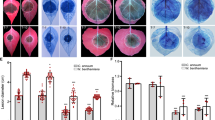
XopPXoo (Xoo3222), a X. oryzae pv. oryzae type III effector, belongs to a XopP family that is conserved in Xanthomonads18; however, the biochemical functions of the XopP family have not yet been identified. In fact, members of the XopP family do not contain any functional domains predicted from their amino-acid sequences. To investigate how XopPXoo functions in rice cells, we generated six independent transgenic rice lines expressing XopPXoo (XopPXoo-OX), and two of them were used for pathological experiments. The expression of XopPXoo mRNAs in rice plants was confirmed by semi-quantitative reverse transcription (RT)–PCR (Supplementary Fig. 1). The XopPXoo-OX plants did not exhibit any morphological phenotypes (Supplementary Fig. 2). We inoculated the XopPXoo-OX lines with the T3SS-deficient hrpX mutant of Xoo that is incapable of type III effector delivery18,19. Because inoculation of the Xoo hrpX mutant induces a strong PTI response in rice, the Xoo hrpX mutant did not cause any disease lesions in wild-type (non-transformed) plants (Fig. 1a,b). In contrast, the XopPXoo-OX plants had severe disease lesions following infection with the Xoo hrpX mutant. Bacterial populations of the Xoo hrpX mutant in the XopPXoo-OX leaves were significantly higher than in wild-type plants (Fig. 1c). In addition, we examined the resistance of XopPXoo-OX plants to wild-type Xoo by using Xoo T7174R, a spontaneous rifampicin-resistant mutant derived from the wild-type isolate Xoo MAFF311018. The XopPXoo-OX plants developed larger disease lesions than the wild type (Fig. 1d,e), and bacterial growth in the XopPXoo-OX plants was also slightly higher than the growth in the wild type (Fig. 1f). Thus, XopPXoo likely inhibited immunity induced by infection with both Xoo strains. Although the XopPXoo-knockout Xoo strain was generated and inoculated on wild-type rice plants, the strain had no defects in virulence (Supplementary Fig. 3).
(a) Transgenic plants expressing XopPXoo were more susceptible to the Xoo hrpX mutant. Disease lesions were photographed at 25 d.p.i. Scale bar, 10 mm. (b) The lesions induced by infection with the Xoo hrpX mutant were measured at 25 d.p.i. (c) The size of the Xoo hrpX mutant population in two independent XopPXoo-OX transgenic lines was determined at 5 d.p.i. CFU, colony-forming unit. (d) Transgenic plants expressing XopPXoo were more susceptible to Xoo T7174R, a spontaneous rifampicin-resistant mutant derived from the wild-type isolate Xoo MAFF311018. Disease lesions were photographed at 25 d.p.i. Scale bar, 10 mm. (e) The lesions induced by infection with the Xoo T7174R were measured at 25 d.p.i. (f) The size of the Xoo T7174R population was determined at 5 d.p.i. Data shown in b,c,e,f are the means±s.d. for three individual experiments. Asterisks indicate significant differences between wild-type and XopPXoo-OX plants (Student’s t-test P values, P<0.05). (g) Expression levels of PBZ1 and PAL1 in wild-type (WT) and XopPXoo-OX cells after treatments with 100 μg ml−1 PGN were analysed using quantitative RT–PCR. (h) Expression levels of PBZ1 and PAL1 in WT and XopPXoo-OX cells after treatments with 2 μg ml−1 chitin were analysed using quantitative RT–PCR. Data shown in g,h are the means±s.d. calculated using three biological replicates, where each biological replicate consists of two technical replicates. Asterisks indicate significant differences between WT and XopPXoo-OX cells (Student’s t-test P values, P<0.05).
To analyse the PTI-inhibitory activity of XopPXoo, we generated two suspension-cultured cell lines derived from callus lines independently transformed with the XopPXoo-OX construct. The expression of XopPXoo mRNA in rice cells was confirmed by semi-quantitative RT–PCR (Supplementary Fig. 4). We treated these XopPXoo-OX cells with a bacterial MAMP PGN. Quantitative RT–PCR indicated that the expression of two defence-related genes, Probenazole1 (PBZ1)30 and Phenylalanine ammonia-lyase1 (PAL1)31, was significantly suppressed in the XopPXoo-OX cells (Fig. 1g). In addition, we also treated these suspension-cultured cells with chitin, a fungal MAMP. The chitin-induced expression of PBZ1 and PAL1 was also reduced in the XopPXoo-OX cell lines (Fig. 1h). These data indicate that XopPXoo inhibits PTI induced by both PGN and chitin. However, it was noted that expression of XopPXoo in rice cells did not influence chitin-induced activation of MAP kinases (Supplementary Fig. 5).
Identification of OsPUB44 as a host target for XopPXoo
To elucidate how XopPXoo suppresses host immune responses, we screened host proteins capable of interacting with XopPXoo by a yeast two-hybrid assay using a rice complementary DNA library prepared from chitin-treated rice suspension cells32. We isolated five positive clones as potential candidates for host targets of XopPXoo from 8 × 105 cDNA clones. All five candidates corresponded to OsPUB44 (Os05g0439400) encoding a putative PUB E3 ubiquitin ligase with a U-box domain and a carboxy-terminal armadillo (ARM) repeat domain7. OsPUB44 belongs to the class III PUB family that includes CMPG1 and AtPUB22, AtPUB23 and AtPUB24, all known to be involved in immune responses9,10,33.
The deduced OsPUB44 protein is 452 amino-acid residues in length. Full-length OsPUB44 interacted with XopPXoo in yeast cells (Fig. 2a). We analysed the expression of OsPUB44 mRNA in response to PGN and chitin using wild-type rice suspension cells. The levels of OsPUB44 mRNA were markedly increased by both PGN and chitin treatments (Fig. 2b). The immunoblot treated with OsPUB44-specific antibody indicated that the levels of OsPUB44 protein were also increased with PGN and chitin treatments (Fig. 2c).
(a) Interaction between XopPXoo and OsPUB44 in a yeast two-hybrid assay. The growth of yeast colonies on the plate (–ULWH) lacking uracil (U), leucine (L), tryptophan (W) and histidine (H) indicates a positive interaction. (b) Transcript levels of OsPUB44 mRNA in rice suspension-cultured cells after treatments with 100 μg ml−1 PGN or 2 μg ml−1 chitin were analysed using quantitative RT–PCR. Data are the means±s.d. for three individual experiments. (c)The protein levels of OsPUB44 in rice suspension-cultured cells after treatments with 100 μg ml−1 PGN or 2 μg ml−1 chitin were determined by treating an immunoblot with an OsPUB44-specific antibody. Molecular weight of protein: OsPUB44=48 kDa. (d) Subcellular localization of OsPUB44-GFP, XopPXoo-GFP and GFP in transiently transfected rice protoplasts. Fluorescence was observed 18 h after transformation. Scale bar, 10 μm. (e) Visualization of the interaction between XopPXoo and OsPUB44 using a BiFC assay in transiently transfected rice protoplasts. Vn-XopPXoo and Vc-OsPUB44 contain the N-terminal and C-terminal fragments, respectively, of Venus. The interaction between XopPXoo and OsPUB44 was indicated by Venus fluorescence. Vn-GUS was used as a negative control. The fluorescence was observed 18 h after transformation. Scale bar, 10 μm. (f) In vitro interaction between XopPXoo and OsPUB44. OsPUB44 was incubated with GST or GST-XopPXoo and then pulled down with glutathione-Sepharose 4B beads. OsPUB44 was detected by immunoblotting using an anti-OsPUB44 antibody. GST-XopPXoo was detected by Coomassie brilliant blue (CBB) staining. Molecular weights of proteins: OsPUB44=48 kDa, GST-XopPXoo=107 kDa, GST=28 kDa.
To examine the subcellular localization of XopPXoo and OsPUB44 in planta, green fluorescent protein (GFP) was fused to the C termini of XopPXoo and OsPUB44, and constructs were expressed in rice protoplasts. Fluorescence for XopPXoo-GFP and OsPUB44-GFP was detected in the entire region (Fig. 2d). Expression of these GFP-fused proteins was confirmed by immunoblot analysis (Supplementary Fig. 6). In addition, the amino-terminal GFP fusions, GFP-XopPXoo and GFP-OsPUB44, were also localized in the entire region (Supplementary Fig. 7).
We performed bimolecular fluorescence complementation (BiFC) to analyse the interaction between OsPUB44 and XopPXoo in rice cells. OsPUB44 was tagged with the N-terminal domain (1–154 aa) of Venus (Vn-OsPUB44), and XopPXoo was tagged with the C-terminal domain (155–238 aa) of Venus (Vc-XopPXoo). Co-expression of Vc-XopPXoo with Vn-OsPUB44 in rice protoplasts resulted in fluorescence complementation in the entire region (Fig. 2e), suggesting the in vivo interaction of XopPXoo with OsPUB44. In addition, an in vitro pull-down assay was used to confirm the interaction between OsPUB44 and XopPXoo. We prepared OsPUB44 and glutathione S-transferase (GST)-fused XopPXoo using a protein expression system in Escherichia coli. OsPUB44 was pulled down with GST-XopPXoo, whereas GST alone did not interact with OsPUB44 (Fig. 2f). These data indicate that OsPUB44 directly interacts with XopPXoo.
OsPUB44 plays a role in PAMPs-triggered immunity
To determine whether OsPUB44 is involved in PTI, we produced transgenic rice suspension cells carrying the OsPUB44 RNAi construct (Supplementary Fig. 8). Both transcript and protein levels of OsPUB44 were significantly reduced in two independent OsPUB44 RNAi lines (Fig. 3a,b; Supplementary Fig. 9). PGN- and chitin-induced expressions of PBZ1 and PAL1 were significantly suppressed in both RNAi lines (Fig. 3c,d), whereas chitin-induced activation of MAP kinases was not altered in the OsPUB44 RNAi line (Supplementary Fig. 10). These data coincided with the results of the XopPXoo-OX lines. We also produced transgenic rice plants carrying the OsPUB44 RNAi construct (Supplementary Fig. 11) and tested whether silencing of OsPUB44 affects resistance to the Xoo hrpX mutant using two independent RNAi plant lines. The OsPUB44 RNAi plants developed larger disease lesions than the wild type (Fig. 3e; Supplementary Fig. 12a). The size of the bacterial population in the Xoo hrpX mutant in OsPUB44-RNAi leaves was significantly higher than in wild-type plants (Fig. 3f). We also analysed the disease resistance of the OsPUB44 RNAi plants to Xoo T7174R. Although we did not detect a significant difference in lesion lengths between the wild-type and the OsPUB44 RNAi plants (Fig. 3g; Supplementary Fig. 12b), the amount of bacterial growth indicated that the OsPUB44 plants were more susceptible to Xoo T7174R as compared with wild-type plants (Fig. 3h). Thus, it is likely that OsPUB44 positively regulates PTI and Xoo resistance in rice.
(a) Transcript levels of OsPUB44 in the OsPUB44 RNAi cells were analysed by RT–PCR. OsActin1 was used as a control. Molecular sizes of DNA fragments: OsPUB44=1,359 bp, OsActin1=387 bp. (b) The protein levels of OsPUB44 in the OsPUB44 RNAi cells were determined by immunoblot analysis with the OsPUB44-specific antibody. Molecular weight of protein: OsPUB44=48 kDa. (c) Expression levels of PBZ1 and PAL1 in wild-type and OsPUB44 RNAi cells after treatments with 100 μg ml−1 PGN were analysed using quantitative RT–PCR. (d) Expression levels of PBZ1 and PAL1 in wild-type and OsPUB44 RNAi cells after treatments with 2 μg ml−1 chitin were analysed using quantitative RT–PCR. Data shown in c,d are the means±s.d. for three individual experiments. Asterisks indicate significant differences between WT and OsPUB44 RNAi cells (Student’s t-test P values, P<0.05). (e) Plants transformed with the OsPUB44 RNAi construct were more susceptible to the Xoo hrpX mutant. The lesions induced by infection with the Xoo hrpX mutant were measured at 25 d.p.i. (f) The size of the Xoo hrpX mutant population in the OsPUB44 RNAi plants was determined at 5 d.p.i. CFU, colony-forming unit. (g) The disease lesions of the OsPUB44 RNAi plants induced by infection with Xoo T7174R were measured at 25 d.p.i. (h) The size of the Xoo T7174R population in the OsPUB44 RNAi plants was determined at 5 d.p.i. Data shown in e–h are the means±s.d. calculated using three biological replicates, where each biological replicate consists of two technical replicates. Asterisks indicate significant differences between wild type and RNAi line 1, or RNAi line 2 (Student’s t-test P values, P<0.05).
XopPXoo inhibits the ubiquitin ligase activity of OsPUB44
To identify the PTI-inhibitory mechanism mediated by XopPXoo, we analysed the domains of OsPUB44 that could interact with XopPXoo using yeast two-hybrid analysis. As shown in Fig. 4a, we generated four constructs containing the U-box and/or ARM domain with or without a linker domain (103–203 aa) that were designated as OsPUB441–452, OsPUB441–102, OsPUB441–203 and OsPUB44102–452. XopPXoo was shown to interact with OsPUB441–102 and OsPUB441–203, but not with OsPUB44102–452 (Fig. 4b). In addition, OsPUB441–203 interacted with XopPXoo more strongly than OsPUB441–102, which resulted from lower protein level of OsPUB441–102 in yeast (Supplementary Fig. 13).
(a) Schematic structures of full-length OsPUB441–452 and three truncated OsPUB44 fragments, OsPUB441–102 (1–102 aa), OsPUB441–203 (1–203 aa) and OsPUB44102–452 (102–452 aa). Black and grey boxes indicate the conserved U-box and ARM repeat domains, respectively. Numbers above the boxes indicate amino-acid residues. (b) Interaction of XopPXoo with OsPUB441–452, OsPUB441–102, OsPUB441–203 and OsPUB44102–452 in a yeast two-hybrid assay. (c) In vitro ubiquitination assays of the OsPUB44 protein. The GST-OsPUB441–203 protein was incubated with or without E1, E2 and ubiquitin (Ub)-FLAG. Ubiquitinated OsPUB441–203 proteins were detected with an anti-FLAG antibody (upper panel). The presence of GST-OsPUB441–203 protein was confirmed by immunoblotting using an anti-GST antibody (lower panel). Molecular weight of protein: GST-OsPUB441–203=48 kDa. Arrow indicates ubiquitin-loaded E2 proteins. Same results were obtained in more than three experiments. (d) The ubiquitin ligase activity of OsPUB44 was inhibited by XopPXoo. An in vitro ubiquitination assay of GST-OsPUB441–203 was performed with or without GST-XopPXoo. Ubiquitinated OsPUB441–203 was detected with an anti-FLAG antibody (upper panel). The presence of GST-OsPUB441–203 and GST-XopPXoo proteins was confirmed by immunoblotting using an anti-GST antibody (lower panel). The GST protein was detected by Coomassie brilliant blue staining. Molecular weights of proteins: GST-OsPUB441–203=48 kDa, GST-XopPXoo=107 kDa, GST=28 kDa. Arrow indicates ubiquitin-loaded E2 proteins. Same results were obtained in more than three experiments.
To address whether OsPUB44 possesses the E3 ubiquitin ligase activity, OsPUB441–203 was fused to GST and expressed in E. coli. The GST-fused OsPUB441–203 was incubated with E1 enzyme, E2 enzyme (UbcH6) and Flag-tagged ubiquitin. The ubiquitinated proteins were detected by immunoblot analysis using anti-FLAG antibody. As shown in Fig. 4c, OsPUB441–203 had significant ubiquitin ligase activity in the presence of all the essential reaction components. We also analysed the ubiquitin ligase activity of full-length OsPUB441–452. Full-length OsPUB441–452 had significant ubiquitin ligase activity, although much less activity than that of OsPUB441–203 (Supplementary Fig. 14).
The interaction of XopPXoo with the U-box domain of OsPUB44 raised a possibility that XopPXoo may inhibit the ubiquitin ligase activity. Therefore, we carried out an in vitro ubiquitin ligase assay of the U-box domain in the presence of XopPXoo. The ubiquitin ligase activity was significantly reduced by the addition of GST-XopPXoo in a dose dependent manner (Fig. 4d), whereas GST did not affect the activity. These data strongly suggest that the OsPUB44 ubiquitin ligase activity is suppressed by XopPXoo by direct binding.
The ubiquitin E3 ligases are known to regulate their own protein levels by autocatalytic ubiquitination-mediated protein turnover. In fact, the inactive mutants of AtPUB22 were found to be stable and could accumulate to high levels in the cells34. Therefore, if XopPXoo inhibits OsPUB44 activity in vivo, the inactivated OsPUB44 should accumulate in the XopPXoo-OX cells. The protein levels of OsPUB44 were compared between wild-type and the XopPXoo-OX cells after treatments with PGN or chitin. We found that the OsPUB44 proteins accumulated in XopPXoo-OX cells to higher levels than in wild type (Fig. 5a). The protein levels of OsPUB44 in the XopPXoo-OX cells were more than 15-fold higher as compared with those in wild type at 30 min after treatments with PGN and chitin. These data support the idea that XopPXoo interacts with the U-box domain in vivo and inhibits ubiquitin ligase activity, resulting in stabilization of OsPUB44 in the cells.
(a) OsPUB44 was stabilized in the presence of XopPXoo. The protein levels of OsPUB44 in wild-type (WT) and XopPXoo-OX cells after treatments with 100 μg ml−1 PGN or 2 μg ml−1 chitin were determined by immunoblot analysis with the OsPUB44-specific antibody. Molecular weight of protein: OsPUB44=48 kDa. (b) Transcript levels of OsPUB44 mRNA in WT and XopPXoo-OX cells after treatment with 100 μg ml−1 PGN or 2 μg ml−1 chitin were analysed using quantitative RT–PCR. Data are the means±s.d. for three individual experiments. Asterisks indicate significant differences between WT and XopPXoo-OX cells (Student’s t-test P values, P<0.05). (c) The protein levels of OsPUB44 in rice cells after treatment with the proteasomal inhibitor MG132 (30 μM) or an equivalent volume of dimethylsulphoxide (DMSO; control) were determined by immunoblot analysis with the OsPUB44-specific antibody. Molecular weight of protein: OsPUB44=48 kDa. (d) Transcript levels of OsPUB44 mRNA in rice cells after treatment with the proteasomal inhibitor MG132 (30 μM) or an equivalent volume of DMSO (control) were analysed using quantitative RT–PCR. Data are the means±s.d. for three individual experiments. Asterisks indicate significant differences between control and MG132-treated cells (Student’s t-test P values, P<0.05).
The PGN- and chitin-induced expression levels of OsPUB44 mRNA were slightly but significantly higher in the XopPXoo-OX cells as compared with wild type (Fig. 5b), although the increased levels of OsPUB44 mRNA in the XopPXoo-OX cells was much lower as compared with the increased levels of OsPUB44 proteins. It is possible that the slight increase in mRNA levels may be caused by the inhibition of OsPUB44 turnover. To test this possibility, rice cells were treated with a proteasome inhibitor MG132. Treatment with MG132 induced accumulation of the OsPUB44 protein (Fig. 5c), and also increased the transcript levels (Fig. 5d). These data suggest that inhibition of protein turnover of the OsPUB44 protein may enhance its transcription or stability of the transcript.
OsPUB44 possesses a unique U-box domain
A phylogenetic analysis of the U-box family indicated that the U-box domain of OsPUB44 is highly homologous to those of OsPUB45 (Os02g0540700) and OsPUB46 (Os04g0418500; Supplementary Fig. 15). We examined the interaction of XopPXoo with OsPUB45 and OsPUB46 (Fig. 6a); however, no interactions were detected, although these proteins were expressed in yeast (Supplementary Fig. 16). Therefore, we compared the amino-acid sequences of the OsPUB44 U-box domain with those of OsPUB45 and OsPUB46, and Arabidopsis orthologues AtPUB27, AtPUB28 and AtPUB29. Eight amino-acid residues within the U-box domain of OsPUB44 varied from the corresponding residues of the closely related PUB proteins including OsPUB45 and OsPUB46 (Fig. 6b), suggesting that these eight amino-acid residues may determine the specificity of interaction with XopPXoo. We produced OsPUB44 U-box mutants in which each of the OsPUB44-specific residues was substituted with the corresponding residues in OsPUB45 and OsPUB46. The interaction between XopPXoo and the OsPUB441–203 mutants was analysed by the yeast two-hybrid method. We found that XopPXoo did not interact with two mutants, OsPUB441–203, L86T and OsPUB441–203, H94W (Fig. 6c). Although OsPUB441–203, P48C also did not interact with XopPXoo, this failure was caused by undetectable expression of OsPUB441–203, P48C in yeast (Supplementary Fig. 16). To further examine whether L86 and H94 contribute to the interaction, we mutated two amino-acid residues (T133 and W141) of OsPUB45 with two residues corresponding to L86 and H94 of OsPUB44, and generated OsPUB451–234, T133L, W141H. The new construct, OsPUB451–234, T133L, W141H interacted with XopPXoo (Fig. 6d), demonstrating that leucine and histidine corresponding to L86 and H94 of OsPUB44 are essential residues for interaction with XopPXoo.
(a) Interaction of XopPXoo with OsPUB44, OsPUB45 and OsPUB46 in a yeast two-hybrid assay. (b) Eight amino-acid residues within the U-box domain of OsPUB44 are different from those of OsPUB44 homologues and Arabidopsis orthologues. A comparison of the amino-acid sequence of the U-box domain of OsPUB44 with those of OsPUB45, OsPUB46, AtPUB27, AtPUB28 and AtPUB29 was performed using ClustalW2 software. The conserved amino-acid residues are shaded. Asterisks indicate the eight amino-acid residues specific for OsPUB44. (c) Interaction of XopPXoo with the substituted OsPUB441–203 proteins in a yeast two-hybrid assay. (d) Interaction of XopPXoo with OsPUB451–234, T133L, W141H in a yeast two-hybrid assay.
We investigated whether the U-box proteins containing these two OsPUB44-specific residues generally exist in the plant kingdom. A database search did not find any PUB proteins with two residues corresponding to L86 and H94 of OsPUB44, except for OsPUB44.
XopPXoo targets the unique U-box domain of OsPUB44
Because OsPUB45 did not interact with XopPXoo, we hypothesized that XopPXoo could not inhibit the ubiquitin ligase activity of OsPUB45. The U-box domain of OsPUB45 had ubiquitin ligase activity in vitro (Fig. 7a). However, the ligase activity of OsPUB45 was not suppressed in the presence of XopPXoo, whereas the same amount of XopPXoo inhibited the OsPUB44 activity (Fig. 7b). To investigate whether the two amino-acid residues (L86 and H94) responsible for interaction with XopPXoo affect the inhibition of the OsPUB44 ligase activity by XopPXoo, we prepared the recombinant OsPUB44 U-box protein with L86T and H94W in E. coli. The OsPUB441–203, L86T, H94W protein possessed E3 ligase activity that was not inhibited by XopPXoo (Fig. 7c). In contrast, OsPUB45 U-box protein with T133L and W141H was inhibited by XopPXoo (Fig. 7d). These data indicated that leucine and histidine corresponding to L86 and H94 of OsPUB44 are required for inhibition of the ligase activities by XopPXoo.
The same amounts (10 nM) of GST-OsPUB451–234 (a), GST-OsPUB441–203 (b), OsPUB441–203, L86T, H94W (c) and OsPUB451–234, T133L, W141H (d) were incubated in a reaction mixture containing E1, E2 and ubiquitin (Ub)-FLAG with or without GST-XopPXoo (10 nM). Ubiquitinated proteins were detected with an anti-FLAG antibody (upper panels). The presence of GST-OsPUB441–203, GST-OsPUB451–234, GST-OsPUB441–203, L86T, H94W, OsPUB451–234, T133L, W141H and GST-XopPXoo was confirmed by immunoblot analysis using anti-GST antibody (lower panels). Molecular weights of proteins: GST-OsPUB441–203=48 kDa, GST-OsPUB451–234=52 kDa, GST-XopPXoo=107 kDa. Similar results were obtained in more than three experiments.
Discussion
U-box proteins are implicated in immunity and most function as negative regulators10,11,13. To date, only one U-box protein, CMPG1, has been identified as a positive immune regulator. CMPG1, originally identified as a gene rapidly induced after Cladosporum fulvum Avr9 elicitation in Cf-9 tobacco cells35, positively regulates cell death and disease resistance in several types of plant–microbe interactions9. In this study, we found that silencing OsPUB44 resulted in reduced expression of PGN- and chitin-induced defence genes and suppressed resistance to Xoo. These data strongly suggest that OsPUB44 is another positive regulator of immune responses.
Since OsPUB44 and CMPG1 positively regulate immune responses, these proteins are likely important targets for pathogens to suppress host defence. CMPG1 is known to be targeted by P. infestans effector AVR3a15,16. AVR3a interacts with and stabilizes CMPG1, resulting in the inhibition of CMPG1. Nevertheless, how AVR3a stabilizes CMPG1 in plant cells and why CMPG1 function is lost by the stabilization remain to be identified. In this study, we discovered that XopPXoo directly interacts with the U-box domain of OsPUB44, thereby reducing the E3 ligase activity. As found in the interaction between CMPG1 and AVR3a, the expression of XopPXoo in host cells also stabilizes OsPUB44. The phenomenon may be explained by the hypothesis that XopPXoo inhibits autocatalytic ubiquitination-mediated turnover of OsPUB44. This possibility is also supported by the observation that MG132-mediated inhibition of 26S proteasomes stabilized OsPUB44. These results indicate that stabilization of OsPUB44 occurs by inactivation of ligase activity. Although many pathogen effectors are known to target host proteins involved in the ubiquitin-proteasome pathway8,14,15,16, this is the first case documenting that pathogen effectors interact with the E3 ligase domain and modulate its activity.
Other U-box proteins involved in immunity function as negative regulators. For example, Arabidopsis AtPUB12 and AtPUB13 ubiquitinate FLS2, a flagellin receptor, and then FLS2 is degraded in the proteasome pathway11, leading to attenuation of immune signalling. Arabidopsis AtPUB22, AtPUB23 and AtPUB24 redundantly and negatively regulate PTI10. In addition, AtPUB22 and AtPUB23 also function as negative regulators in the water-stress response36. Recently, Exo70B2, a component of the exocytic machinery, was identified as a substrate for AtPUB22 (ref. 34). AtPUB22 is proposed to control PTI by degradation of Exo70B2.
Although a limited number of the U-box E3 ligases function as positive regulators, there are several RING-finger-type E3 ligases that positively regulate immune responses8. Recently, Park et al.14 reported that M. oryzae effector AvrPiz-t targets rice RING finger-type E3 ubiqutin ligase APIP6, a positive regulator for PTI. Interestingly, APIP6 ubiquitinates AvrPiz-t in vitro. In fact, when AvrPiz-t and APIP6 are ectopically co-expressed in tobacco cells, AvrPiz-t is degraded in the presence of APIP6. In addition, the E3 ligase activity of APIP6 is reduced in the presence of AvrPiz-t; however, the molecular details of how AvrPiz-t reduces the ligase activity of APIP6 remain to be determined.
In this study, we found that two amino-acid residues in the OsPUB44 U-box domain are responsible for interaction with XopPXoo. Among 78 U-box proteins in rice, only OsPUB44 contains two unusual amino-acid residues. Although Arabidopsis contains 63 U-box proteins, there are no OsPUB44-type U-box proteins encoded in the Arabidopsis genome. How these unusual U-box proteins evolved is an intriguing question.
Another important question is why XopPXoo targets the unusual U-box domain. If XopPXoo could interact with typical U-box domains, many U-box proteins would be influenced by XopPXoo, which is an unsuitable condition for pathogen proliferation. For example, as mentioned above, AtPUB12, AtPUB13, AtPUB22, AtPUB23 and AtPUB24 negatively regulate immunity in Arabidopsis10,11. If XopPXoo interacted with these negative regulators and inhibited their functions, enhanced immune responses would result. If XopPXoo targeted the U-box proteins that play essential roles in host physiological processes, XopPXoo would damage the host cells. Thus, it is likely that pathogen effectors require a high degree of specificity for interaction with host targets.
Although XopPXoo is classified into the XopP family, XopPXoo has only 40% identity with the XopP homologue of X. campestris pv. campestris18. Therefore, Xcc XopP may possibly target other factors involved in immunity. The biochemical function of XopPXoo cannot be predicted due to the lack of any known domains. Therefore, it is unclear whether the E3 ligase activity of OsPUB44 is inhibited by an enzymatic reaction of XopPXoo or a physical interaction with XopPXoo. Another possibility is that XopPXoo may inhibit an interaction between OsPUB44 and E2 enzyme by competing for the binding site with E2 enzyme. Although we tested this hypothesis, the data were inconclusive from the in vitro interaction experiment. In some cases, the enzymatic properties of pathogen effectors have been discovered by determining the tertiary structures of effector proteins37. Crystal structures of XopPXoo and/or OsPUB44 will help to reveal how XopPXoo inhibits OsPUB44 activity at the molecular level.
In Arabidopsis, AtPUB12 and AtPUB13 ubiquitinate FLS2, the extracellular leucine-rich repeats-containing receptor-like kinase11. Thus, it is possible that OsPUB44 may be associated with OsCERK1, a rice extracellular LysM domain-containing receptor-like kinase responsible for recognizing chitin and PGN. However, chitin-induced activation of MAP kinases was not altered in OsPUB44 RNAi and XopPXoo-OX cells, suggesting that OsPUB44 as well as XopPXoo may not regulate the protein level and the activity of OsCERK1.
Recently, we identified rice receptor-like cytoplasmic kinase OsRLCK185 as a target for X. oryzae effector XopYXoo and found that OsRLCK185 transmits the immune signal from OsCERK1 to the downstream MAP kinase cascade19. Since MAP kinase activation was not altered in the OsPUB44 RNAi cells, it is likely that OsPUB44 functions in a different pathway than OsRLCK185 or downstream of the MAPK cascade. Identification of the host proteins ubiquitinated by OsPUB44 will be a key factor in understanding how OsPUB44 regulates PAMPs-triggered immunity.
Methods
Plasmid constructs
Full-length cDNAs for OsPUB44 (Os05g0439400), OsPUB45 (Os02g0540700) and OsPUB46 (Os04g0418500) were amplified by PCR from cDNA pools prepared from leaves of rice cultivars Nipponbare or Kinmaze (primers are described in Supplementary Table 1). The coding region of XopPXoo was amplified from genomic DNA prepared from X. oryzae pv. oryzae. Forward and reverse primers were designed on the basis of sequence information in the Rice Annotation Project Database ( http://rapdb.dna.affrc.go.jp/) or the Xanthomonas oryzae pv. oryzae Genome Database ( http://microbe.dna.affrc.go.jp/Xanthomonas/). The coding region of each gene was amplified by PCR and ligated into the pENTR/D-TOPO cloning vector (Invitrogen). To produce XopPXoo-OX cells and plants, the XopPXoo coding region was introduced into the p2K1 vector to control expression with the maize ubiquitin 1 promoter38. For RNAi vectors, a 514-bp (base pair) fragment of OsPUB44 was amplified, combined by PCR and cloned into pENTR/D-TOPO (Supplementary Fig. 8). The 514-bp fragment was subcloned into the pANDA vector in the inverse orientation by one LR clonase reaction39. Point mutations of OsPUB44 were generated by PCR using primers designed to substitute the corresponding amino acids (primers are described in Supplementary Table 1). OsPUB44A31F, E39D, OsPUB44H66N, R73M, OsPUB44P48C, OsPUB44A51V, OsPUB44L86T and OsPUB44H94W were made by replacing Ala-31and Glu-39, His-66 and Arg-73, Pro-48, Ala-51, Leu-86 and His-94 in OsPUB44 with Phe and Glu, Asn and Met, Cys, Val, Thr and Trp, respectively. OsPUB45T133L, W141H was made by replacing Thr-133 and Trp-141 in OsPUB45 with Leu and His, respectively. OsPUB44, OsPUB441–102, OsPUB441–203, OsPUB44102–452, OsPUB44C34A, OsPUB44V45I, OsPUB44A31F, E39D, OsPUB44H66N, R73M, OsPUB44P48C, OsPUB44A51V, OsPUB44L86T, OsPUB44H94W, OsPUB45, OsPUB45T133L, W141H, OsPUB46 and XopPXoo clones were transferred by the Gateway system using an LR clonase reaction into pBTM116-GW and pVP16-GW for the yeast two-hybrid assays40, into p35S-GFP-GW for subcellular localization assays, into p35S-Vn-GW and p35S-Vc-GW for the BiFC assay and into the p2K-GW binary vector to produce transgenic plants expressing trans genes under the control of the maize ubiquitin promoter19.
Rice transformation
Calli generated from embryos of rice cultivars Nipponbare or Kinmaze, which were used for XopPXoo-OX or OsPUB44 RNAi, respectively, were transformed by infection of Agrobacterium tumefaciens EHA101 carrying corresponding constructs as described41. The transformed calli were selected by resistance to hygromycin and used for generation of T0 plants. Suspension-cultured cells were also prepared from transformed calli or from calli generated from T0 plant seeds.
Yeast two-hybrid assays
A yeast two-hybrid library was prepared using total RNA extracted from rice suspension cells that had been treated with chitin32. To construct the bait vector, the coding region of XopPXoo was cloned into pBTM116. Screening was performed on synthetic complete (SC) medium lacking histidine. Combinations of the bait and prey vectors were introduced into cells of Saccharomyces cerevisiae L40 (MATatrp1 leu2 his3 LYS2::lexA-HIS3 URA3::lexA-lacZ). Transformants were selected on minimal medium lacking tryptophan and leucine. The two-hybrid interaction was analysed based on the histidine requirement for yeast growth as described previously32,40.
PAMP treatment
Rice suspension-cultured cells were subcultured for 3 days in fresh medium, divided into 12-well plates (150 mg cells, 2 ml fresh medium per well) and treated with 2 μg ml−1 hexa-N-acetylchitohexaose (Seikagaku Biobusiness) or 100 μg ml−1 PGN from Bacillus subtilis (Sigma).
RNA isolation and RT–PCR analysis
Total RNA was isolated from rice suspension-cultured cells using TRIzol reagent (Invitrogen) and then treated with RNase-free DNase I (Roche). First-strand cDNA was synthesized from 1 μg total RNA with an oligo-dT primer and ReverTra Ace (Toyobo). Expression levels were quantified by qRT–PCR using SYBR Green master mix (Applied Biosystems) in a Step-One Plus Real-Time PCR system (Applied Biosystems), and normalized against a ubiquitin reference gene. Three biological replicates were used for each experiment, and two quantitative iterations were performed for each biological replicate.
Protein extraction and immunoblotting
Total proteins were extracted in a buffer including 100 mM Tris–HCl, pH 7.5, 20% (v/v) glycerine and protease inhibitor cocktail (Roche) and analysed by protein immunoblotting with anti-OsPUB44 antibody (1:2,000 dilution), anti-GFP antibody (ab6556; Abcam, 1:2,000 dilution) or anti-pMAPK (4370; Cell Signaling, 1:2,000 dilution). Polyclonal antibody (prepared by MBL) for OsPUB44 was raised in rabbits using a polypeptide fragment (aa 330–452) from OsPUB44 as the antigen. The uncropped immunoblots are shown in Supplementary Fig. 17).
Transient assays using rice protoplasts
Protoplasts were isolated from cultured rice cells by digestion of cell wall with Cellulase RS (Yakult) and Macerozyme R-10 (Yakult) as previously described42. Plasmid transformations were performed using the polyethylene glycol method43. Aliquots (100 μl) of protoplasts prepared from suspension-cultured cells (2.5 × 106 cells ml−1) were transformed by mixing with 5 μg plasmid DNA. For localization analysis and BiFC assays, transfected protoplasts were observed using confocal microscopy (BZ-9000, Keyence, Japan).
In vitro ubiquitination assays
The OsPUB441–203, OsPUB451–234, OsPUB441–203, L86T, H94W, OsPUB451–234, L133T, H141W and XopPXoo fragments were subcloned into a pCold GST vector44. These proteins were expressed as fusion proteins having a GST tag using a Cold-Shock Bacterial Expression System (Takara Bio) following the supplier’s instructions. Each protein was purified using glutathione-Sepharose 4B (GE Healthcare). Ubiquitination assays were performed as described previously10 with minor modification. Protein samples (500 ng) of purified GST-OsPUB441–203, OsPUB451–234, and OsPUB441–203, L86T, H94W, OsPUB451–234, L133T, H141W and 0–20 nM purified GST-XopPXoo were incubated in 30 μl reaction buffer containing 50 mM Tris–HCl, pH 7.5, 5 mM MgCl2, 4 mM ATP, 2 mM dithiothreitol (DTT), 30 ng yeast E1 (Wako), 200 ng UbcH6 for E2 (Wako) and 5 μg FLAG-ubiquitin (Sigma-Aldrich). Reactions were stopped by adding sample buffer (125 mM Tris–HCl, pH 6.8, 20% (v/v) glycerine, 4% (w/v) SDS and 0.4 M DTT) and were then analysed by SDS–PAGE followed by protein immunoblot analysis using an anti-FLAG antibody (F3165; Sigma-Aldrich (1:2,000 dilution)) or an anti-GST antibody (04435-26; Nacalai Tesque (1:5,000 dilution)).
In vitro pull-down assays
The full-length OsPUB44 protein was purified by removing the GST tag through digestion with Turbo 3C protease (Nacalai Tesque). OsPUB44 was incubated with GST-XopPXoo or GST at 30 °C for 15 min in a reaction buffer (50 mM Tris–HCl, pH 7.5, 5 mM MgCl2, 4 mM ATP and 2 mM DTT). Glutathione-Sepharose 4B beads were added and the mixture was incubated at 4 °C for 15 min, and then washed five times with a washing buffer (50 mM Tris–HCl, pH 7.5, 300 mM NaCl and 1 mM DTT). After the final wash, bound proteins were eluted with SDS sample buffer and subjected to protein immunoblot analysis.
Pathology assays
X. oryzae pv. oryzae (Xoo) strains were cultured on NBY plates containing either rifampicin (20 μg ml−1) for Xoo T7174R, a spontaneous rifampicin-resistant mutant derived from wild-type MAFF311018, or rifampicin/kanamycin (25 μg ml−1) for the Xoo hrpX strain for 3 days at 28 °C. Fully expanded rice leaves were inoculated by clipping leaf tips with scissors that had been immersed in bacterial suspensions (OD600=0.3)45. Symptoms were scored by measuring lesion lengths 25 days post inoculation (d.p.i.). Bacterial growth in planta was measured as reported previously18. The inoculated rice leaves (1 cm long) were collected at 5 d.p.i. Leaves were ground in and diluted with sterile water and then plated on the peptone-sucrose medium containing rifampicin or rifampicin and kanamycin for direct colony counts.
Additional information
How to cite this article: Ishikawa, K. et al. Bacterial effector modulation of host E3 ligase activity suppresses PAMP-triggered immunity in rice. Nat. Commun. 5:5430 doi: 10.1038/ncomms6430 (2014).
References
Jones, J. D. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Miya, A. et al. CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc. Natl Acad. Sci. USA 104, 19613–19618 (2007).
Wan, J. et al. A LysM receptor-like kinase plays a critical role in chitin signaling and fungal resistance in Arabidopsis. Plant Cell 20, 471–481 (2008).
Willmann, R. et al. Arabidopsis lysin-motif proteins LYM1 LYM3 CERK1 mediate bacterial peptidoglycan sensing and immunity to bacterial infection. Proc. Natl Acad. Sci. USA 108, 19824–19829 (2011).
Block, A. & Alfano, J. R. Plant targets for Pseudomonas syringae type III effectors: virulence targets or guarded decoys? Curr. Opin. Microbiol. 14, 39–46 (2011).
Sadanandom, A., Bailey, M., Ewan, R., Lee, J. & Nelis, S. The ubiquitin-proteasome system: central modifier of plant signalling. New Phytol. 196, 13–28 (2012).
Zeng, L. R., Park, C. H., Venu, R. C., Gough, J. & Wang, G. L. Classification, expression pattern, and E3 ligase activity assay of rice U-box-containing proteins. Mol. Plant 1, 800–815 (2008).
Trujillo, M. & Shirasu, K. Ubiquitination in plant immunity. Curr. Opin. Plant Biol. 13, 402–408 (2010).
Gonzalez-Lamothe, R. et al. The U-box protein CMPG1 is required for efficient activation of defense mechanisms triggered by multiple resistance genes in tobacco and tomato. Plant Cell 18, 1067–1083 (2006).
Trujillo, M., Ichimura, K., Casais, C. & Shirasu, K. Negative regulation of PAMP-triggered immunity by an E3 ubiquitin ligase triplet in Arabidopsis. Curr. Biol. 18, 1396–1401 (2008).
Lu, D. et al. Direct ubiquitination of pattern recognition receptor FLS2 attenuates plant innate immunity. Science 332, 1439–1442 (2011).
Yang, C. W. et al. The E3 ubiquitin ligase activity of Arabidopsis PLANT U-BOX17 and its functional tobacco homolog ACRE276 are required for cell death and defense. Plant Cell 18, 1084–1098 (2006).
Zeng, L. R. et al. Spotted leaf11, a negative regulator of plant cell death and defense, encodes a U-box/armadillo repeat protein endowed with E3 ubiquitin ligase activity. Plant Cell 16, 2795–2808 (2004).
Park, C. H. et al. The Magnaporthe oryzae effector AvrPiz-t targets the RING E3 ubiquitin ligase APIP6 to suppress pathogen-associated molecular pattern-triggered immunity in rice. Plant Cell 24, 4748–4762 (2012).
Bos, J. I. et al. Phytophthora infestans effector AVR3a is essential for virulence and manipulates plant immunity by stabilizing host E3 ligase CMPG1. Proc. Natl Acad. Sci. USA 107, 9909–9914 (2010).
Yaeno, T. et al. Phosphatidylinositol monophosphate-binding interface in the oomycete RXLR effector AVR3a is required for its stability in host cells to modulate plant immunity. Proc. Natl Acad. Sci. USA 108, 14682–14687 (2011).
Nino-Liu, D. O., Ronald, P. C. & Bogdanove, A. J. Xanthomonas oryzae pathovars: model pathogens of a model crop. Mol. Plant Pathol. 7, 303–324 (2006).
Furutani, A. et al. Identification of novel type III secretion effectors in Xanthomonas oryzae pv. oryzae. Mol. Plant Microbe Interact. 22, 96–106 (2009).
Yamaguchi, K. et al. A receptor-like cytoplasmic kinase targeted by a plant pathogen effector is directly phosphorylated by the chitin receptor and mediates rice immunity. Cell Host Microbe 13, 347–357 (2013).
Yamaguchi, K., Yamada, K. & Kawasaki, T. Receptor-like cytoplasmic kinases are pivotal components in pattern recognition receptor-mediated signaling in plant immunity. Plant Signal. Behav. 8, e25662 (2013).
Yamaguchi, K. et al. Suppression of rice immunity by Xanthomonas oryzae type III effector Xoo2875. Biosci. Biotechnol. Biochem. 77, 796–801 (2013).
Sinha, D., Gupta, M. K., Patel, H. K., Ranjan, A. & Sonti, R. V. Cell wall degrading enzyme induced rice innate immune responses are suppressed by the type 3 secretion system effectors XopN, XopQ, XopX and XopZ of Xanthomonas oryzae pv. oryzae. PLoS ONE 8, e75867 (2013).
Cheong, H. et al. Xanthomonas oryzae pv. oryzae type III effector XopN targets OsVOZ2 and a putative thiamine synthase as a virulence factor in rice. PLoS ONE 8, e73346 (2013).
Akimoto-Tomiyama, C. et al. XopR, a type III effector secreted by Xanthomonas oryzae pv. oryzae, suppresses microbe-associated molecular pattern-triggered immunity in Arabidopsis thaliana. Mol. Plant Microbe. Interact. 25, 505–514 (2012).
Feng, F. et al. A Xanthomonas uridine 5'-monophosphate transferase inhibits plant immune kinases. Nature 485, 114–118 (2012).
Kim, J. G., Stork, W. & Mudgett, M. B. Xanthomonas type III effector XopD desumoylates tomato transcription factor SlERF4 to suppress ethylene responses and promote pathogen growth. Cell Host Microbe. 13, 143–154 (2013).
Canonne, J. et al. The Xanthomonas type III effector XopD targets the Arabidopsis transcription factor MYB30 to suppress plant defense. Plant Cell 23, 3498–3511 (2011).
Kim, J. G. et al. Xanthomonas T3S effector XopN suppresses PAMP-triggered immunity and interacts with a tomato atypical receptor-like kinase and TFT1. Plant Cell 21, 1305–1323 (2009).
Ustun, S., Bartetzko, V. & Bornke, F. The Xanthomonas campestris type III effector XopJ targets the host cell proteasome to suppress salicylic-acid mediated plant defence. PLoS Pathog. 9, e1003427 (2013).
Midoh, N. & Iwata, M. Cloning and characterization of a probenazole-inducible gene for an intracellular pathogenesis-related protein in rice. Plant Cell Physiol. 37, 9–18 (1996).
Zhu, Q. et al. Cloning and properties of a rice gene encoidng phenylalanine ammonia-lyase. Plant Mol. Biol. 29, 535–550 (1995).
Kawasaki, T. et al. Cinnamoyl-CoA reductase, a key enzyme in lignin biosynthesis, is an effector of small GTPase Rac in defense signaling in rice. Proc. Natl Acad. Sci. USA 103, 230–235 (2006).
Azevedo, C., Santos-Rosa, M. J. & Shirasu, K. The U-box protein family in plants. Trends Plant Sci. 6, 354–358 (2001).
Stegmann, M. et al. The ubiquitin ligase PUB22 targets a subunit of the exocyst complex required for PAMP-triggered responses in Arabidopsis. Plant Cell 24, 4703–4716 (2012).
Durrant, W. E., Rowland, O., Piedras, P., Hammond-Kosack, K. E. & Jones, J. D. cDNA-AFLP reveals a striking overlap in race-specific resistance and wound response gene expression profiles. Plant Cell 12, 963–977 (2000).
Cho, S. K., Ryu, M. Y., Song, C., Kwak, J. M. & Kim, W. T. Arabidopsis PUB22 and PUB23 are homologous U-Box E3 ubiquitin ligases that play combinatory roles in response to drought stress. Plant Cell 20, 1899–1914 (2008).
Janjusevic, R., Abramovitch, R. B., Martin, G. B. & Stebbins, C. E. A bacterial inhibitor of host programmed cell death defenses is an E3 ubiquitin ligase. Science 311, 222–226 (2006).
Lieberherr, D. et al. A sphingolipid elicitor-inducible mitogen-activated protein kinase is regulated by the small GTPase OsRac1 and heterotrimeric G-protein in rice. Plant Physiol. 138, 1644–1652 (2005).
Miki, D. & Shimamoto, K. Simple RNAi vectors for stable and transient suppression of gene function in rice. Plant Cell Physiol. 45, 490–495 (2004).
Yamaguchi, K. et al. SWAP70 functions as a Rac/Rop guanine nucleotide-exchange factor in rice. Plant J. 70, 389–397 (2012).
Hiei, Y., Ohta, S., Komari, T. & Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 6, 271–282 (1994).
Kawano, Y. et al. Activation of a Rac GTPase by the NLR family disease resistance protein Pit plays a critical role in rice innate immunity. Cell Host Microbe. 7, 362–375 (2010).
Chen, L. et al. The Hop/Sti1-Hsp90 chaperone complex facilitates the maturation and transport of a PAMP receptor in rice innate immunity. Cell Host Microbe. 7, 185–196 (2010).
Hayashi, K. & Kojima, C. pCold-GST vector: a novel cold-shock vector containing GST tag for soluble protein production. Protein Expr. Purif. 62, 120–127 (2008).
Kauffman, H. E., Reddy, A. P. K., Hsieh, S. P. Y. & Merca, S. D. An improved technique for evaluatinng resistance to rice varieties of Xanthomonas oryzae. Plant Dis. Rep. 57, 537–541 (1973).
Acknowledgements
We thank Dr Ayako Furutani (Ibaraki University) for technical advice, Dr Seiji Takayama (NAIST) for his kind gift of the BiFC vectors and members of the Kawasaki Lab for technical assistance and participation in discussions. This research was supported by Grants-in-Aid for Scientific Research (B)(23380028), for Scientific Research on Innovative Areas (24113519 and 25114517), and by a Strategic Project to Support the Formation of Research Bases at Private Universities: Matching Fund Subsidy from the Ministry of Education, Culture, Sports, Science and Technology, 2011-2015 (S1101035) to T.K., by a Platform for Drug Discovery, Informatics, and Structural Life Science from the Ministry of Education, Culture, Sports, Science and Technology to C.K. and a Grant-in-Aid for JSPS Fellows to K. Ishikawa.
Author information
Authors and Affiliations
Contributions
K. Ishikawa. and T.K. designed the project, analysed the results and wrote the paper. K. Ishikawa, K.Y., K.S., S.Y., K. Inoue, S.T. and C.K. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-17 and Supplementary Table 1. (PDF 527 kb)
Rights and permissions
About this article
Cite this article
Ishikawa, K., Yamaguchi, K., Sakamoto, K. et al. Bacterial effector modulation of host E3 ligase activity suppresses PAMP-triggered immunity in rice. Nat Commun 5, 5430 (2014). https://doi.org/10.1038/ncomms6430
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms6430
This article is cited by
-
Development and characterization of a new sunflower source of resistance to race G of Orobanche cumana Wallr. derived from Helianthus anomalus
Theoretical and Applied Genetics (2024)
-
TALE-induced immunity against the bacterial blight pathogen Xanthomonas oryzae pv. oryzae in rice
Phytopathology Research (2022)
-
Cooperative regulation of PBI1 and MAPKs controls WRKY45 transcription factor in rice immunity
Nature Communications (2022)
-
Arms and ammunitions: effectors at the interface of rice and it’s pathogens and pests
Rice (2021)
-
Tomato SlPUB24 enhances resistance to Xanthomonas euvesicatoria pv. perforans race T3
Horticulture Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.