Abstract
Schistosomiasis, one of the most important neglected tropical diseases worldwide, is caused by flatworms (blood flukes or schistosomes) that live in the bloodstream of humans. The hepatointestinal form of this debilitating disease results from a chronic infection with Schistosoma mansoni or Schistosoma japonicum. No vaccine is available to prevent schistosomiasis, and treatment relies predominantly on the use of a single drug, praziquantel. In spite of considerable research effort over the years, very little is known about the complex in vivo events that lead to granuloma formation and other pathological changes during infection. Here we use, for the first time, a lentivirus-based transduction system to deliver microRNA-adapted short hairpin RNAs (shRNAmirs) into the parasite to silence and explore selected protein-encoding genes of S. mansoni implicated in the disease process. This gene-silencing system has potential to be used for functional genomic–phenomic studies of a range of socioeconomically important pathogens.
Similar content being viewed by others
Introduction
Schistosomiasis is among the most important neglected tropical diseases worldwide, affecting ~200 million people globally and causing 300,000 deaths per annum1. The hepatointestinal form of this debilitating disease is usually caused by a chronic infection with S. mansoni or S. japonicum (blood flukes). No vaccine is available, and treatment relies on the use of one drug (praziquantel), to which resistance is emerging. Of the three main species of schistosome that infect people, S. mansoni is widespread throughout Africa, South America and the Caribbean1. Through a complex aquatic life cycle, S. mansoni is transmitted (via skin penetration) from an infected, aquatic snail (Biomphalaria spp.) to humans. Adult worms dwell in hepatic and intestinal vessels, where they release eggs that become embedded in the liver or intestinal wall, and trigger immune-mediated granuloma formation and associated clinical complications, such as periportal fibrosis and hypertension2. Although some immunopathological changes linked to the granulomata have been studied2, there is limited knowledge of the precise mechanisms underlying these alterations. This knowledge gap relates mainly to the complexity of the parasite life cycle and technical obstacles.
Underpinned by major advances in our understanding of schistosome genomes3,4, gradual progress in the development of functional genomic tools5,6,7,8 provides new opportunities to gain insights into the intricacies of the schistosome–host relationship. Various tools, such as RNA interference (RNAi) using double-stranded RNA or short interfering RNA (siRNA), have been used to investigate the functions of single genes; however, some techniques employed to date can have limitations, such as off-target (that is, nonspecific) effects and an inadequate persistence of gene knockdown for subsequent phenotypic assessment in vitro or in vivo in the host(s)9. These issues are compounded by the challenges of consistently producing sufficient amounts of appropriate parasite stages for functional genomic analyses10. Nonetheless, recent evidence has shown great promise for the use of a lentivirus-based transduction system for specific and persistent gene knockdown in mammalian cells11, with the potential of overcoming most of the disadvantages of previous knockdown methods.
Here we show how lentivirus transduction can be used to achieve specific and persistent knockdown of selected genes (omega-1, ipse and/or kappa-5)12 of S. mansoni, implicated in egg-induced granulomatous responses in the mammalian host13,14. This study allows, for the first time, the functional genomic–phenomic exploration of schistosomiasis mansoni in vivo.
Results
Lentivirus transduction
First, we transduced S. mansoni eggs with lentiviral constructs encoding microRNA-adapted short hairpin RNAs (shRNAmirs) (Fig. 1a). Ten days after transduction with lentiviral particles, both regions of the mCherry gene and regions encoding individual shRNAmirs were detected in the genomic DNAs of treated eggs, whereas neither of these regions was found in DNA from unexposed eggs (Fig. 1b), indicating successful delivery of viral DNA into the nucleus of the host cell. Southern blot hybridization confirmed proviral integration into the genome of S. mansoni (Fig. 1c). Moreover, mCherry was transcriptionally active under the control of the cytomegalovirus (CMV) promoter, as transcripts were detected in complementary DNA from virus exposed but not from unexposed eggs (Fig. 1b). Although attempts to detect fluorescence and assess transduction efficiency in transduced eggs were impaired by autofluorescence of the eggshell (not shown), results demonstrated successful lentiviral transduction and CMV-driven transgene expression in S. mansoni eggs.
(a) Lentivirus construct encoding shRNAmir/mCherry expression cassette under the control of a CMV promoter and containing a packaging signal (Ψ) as well as a long terminal repeat (5′-LTR) and a self-inactivating (3′-SIN) LTR sequence. The shRNAmir structure is indicated. (b) Genomic DNA (gDNA) and RNA were isolated from eggs 10 days after transduction. Lentiviral DNA was detected in gDNA by direct PCR amplification of transgene regions, using the gene ipse as a positive control; cDNA was produced by reverse transcription from total RNA from eggs. Transcription of the transgene was demonstrated by direct PCR of mCherry transgene regions from cDNA, employing the actin-β (act-b) coding region as a positive control (cf. Supplementary Fig. 1 for full gel images). (c) Representative Southern blot analysis of gDNA from eggs of S. mansoni (3 days after transduction) confirming genomic integration of the lentivirus by specific hybridization (vertical bar). For this analysis, gDNA was isolated from eggs 3 days after lentivirus transduction, digested with both restriction endonucleases SpeI and EcoRI (the plasmid vector contains only one site for each enzyme); 5 μg of digested DNA were resolved in an agarose gel (0.8%) and subjected to specific hybridization using a probe (see inset) directed to the transgene encoded by the lentivirus. The expected sizes of the empty vector (EV; plasmid) control and any non-integrated viral DNA were 3.5 and 5.5/2.1 kb, respectively. Lanes: M, molecular weight marker in bp; WT, wildtype; EV, EV control; I, shRNAmir-511+shRNAmir-557 to omega-1; II, shRNAmir-195+shRNAmir-384 to ipse; III, shRNAmir-616+shRNAmir-638 to kappa-5; shRNAmir, microRNA-adapted short hairpin RNA.
shRNAmirs silence transcription in S. mansoni in vitro
Before testing in S. mansoni eggs, we assessed the functionality of seven shRNAmirs in mammalian (COS7) cells expressing cDNA of the target. Treatment with shRNAmir-511, shRNAmir-557 or shRNAmir-558, targeting the omega-1 gene, showed a 29–34% decrease in transcription compared with ‘empty-vector’ (EV) control samples; the transcription of the ipse and kappa-5 genes was consistently reduced by 33–42% (shRNAmir-195 or shRNAmir-384) and 28–34% (shRNAmir-616 or shRNAmir-638), respectively, following treatment (Fig. 2a). The subsequent transduction of S. mansoni eggs with lentivirus encoding shRNAmir-557 and targeting transcripts of omega-1 led to a 45–85% decrease in transcription compared with EV control samples in three independent experiments (Fig. 2b); transcription in EV-treated eggs did not differ from that of untreated control samples. This result shows, for the first time, that shRNAmirs are functional in S. mansoni and can be used for RNAi. Next, we assessed gene knockdown in S. mansoni eggs using a combination of two viruses encoding different shRNAmir sequences that targeted omega-1 (shRNAmir-511+shRNAmir-557), ipse (shRNAmir-195+shRNAmir-384) or kappa-5 (shRNAmir-616+shRNAmir-638). Three days after viral transduction, a 50–60% reduction in transcription was recorded for all shRNAmir treatment groups compared with EV controls (Fig. 2c); other results showed that the knockdown effect on target gene transcription persisted for at least 10 days following transduction (Fig. 2c). Importantly, lentiviral transduction and shRNAmir-induced knockdown of transcription had no effect on the maturation or vitality of miracidia in eggs (Supplementary Movie), providing the basis for an evaluation of transduced eggs in vivo in mice.
(a) Validation of shRNAmir lentivirus constructs in COS7 cells transfected with plasmids encoding omega-1, ipse or kappa-5. COS7 cells were either not transfected (C) or transfected with vector plasmids without the shRNAmir stem-loop sequence (that is, empty-vector (EV)) or with vector plasmids containing gene-specific shRNAmirs shRNAmir-511, shRNAmir-557 and shRNAmir-558 (omega-1); shRNAmir-195, shRNAmir-384 (ipse), shRNAmir-616 and shRNAmir-638 (kappa-5) (labelled with numbers only). Transcription was assessed by qPCR after 48 h, using act-b as a reference gene. Kruskal–Wallis and Dunns post-hoc tests were used to establish statistical significance compared with the EV control: *P<0.05; n=6. (b) Downregulation of omega-1 transcription in S. mansoni eggs 3 days after transduction with a single shRNAmir-containing lentivirus (shRNAmir-557) were compared with untreated eggs (wild type (WT) control) and eggs transduced with EV lentivirus. Relative transcription of omega-1 in eggs was assessed by qPCR 3 days after transduction; transcription was normalized against the geometric mean of Ct values for three reference genes (psdm4, ef1-a and cox-1). Bars show mean±s.e.m.-normalized transcription for untreated control eggs or omega-1-knockdown samples relative to the EV control eggs for three independent experiments. (c) Transcription of omega-1, ipse or kappa-5 was assessed by qPCR 3 or 10 days following transduction. Bars show the mean±s.e.m.-normalized transcription for biological replicates (n=3) of shRNAmir-treated eggs of the same experiment, relative to the EV control.
Effect of knockdown on egg–induced pathology
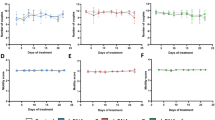
As omega-1, IPSE (egg-secreted cytotoxic or immunomodulatory glycoproteins) and kappa-5 (non-secreted protein) have been implicated in egg-induced responses12,14, we investigated whether knockdown of genes encoding each of these egg proteins had an effect on the development of granulomata associated with pulmonary schistosomiasis induced in BALB/c mice following the injection of S. mansoni eggs into the tail vein. In this established experimental mouse model, eggs are transported to the lungs via the bloodstream and pass into the lung tissues where they become embedded and induce immune responses, leading to subsequent granuloma formation that peaks ~2 weeks after injection15. In our investigation, there was a reduction in the numbers of dendritic cells (DCs), T-helper and B cells, as well as interstitial macrophages in the lungs of mice injected with omega-1 knockdown eggs compared with those injected with eggs carrying the EV (Fig. 3a); the numbers were consistent with those of naive mice. The number of alveolar macrophages in the omega-1 knockdown group was similar to that of the EV control mice, but elevated compared with naive mice (Fig. 3a). A slight decrease in infiltrating interstitial macrophages was also observed in ipse and kappa-5 knockdown mice, but was not significantly different from the EV control mice (Fig. 3a). Interestingly, the numbers of eosinophil granulocytes infiltrating the lung were not affected by knockdown of any of the target genes of S. mansoni (Fig. 3a). Moreover, lung-associated neutrophil numbers were not altered in any of the treatment groups compared with naive control mice (Fig. 3a). The numbers of leukocyte infiltrates in the lung tissue did not differ significantly between wild type (WT) and EV-transduced, egg-injected control mice. Serum samples from mice that had been injected with WT or control virus-transduced eggs had an expected, significant increase in IgE levels, typical of S. mansoni infection; an increase was also seen in all egg-protein-knockdown groups of mice (Fig. 3b). Following a soluble egg antigen (SEA)-specific re-stimulation of lung cell isolates (Fig. 4), all egg-injected mice responded with a characteristic, dominant Th2 cytokine pattern compared with the naive control group. However, no significant difference was observed in any of the knockdown groups compared with EV or WT egg-injected mice.
Test mice were injected into the lateral tail vein with 1,000 eggs previously transduced with lentiviruses containing gene-specific shRNAmir stems (omega-1, n=5; ipse, n=6; kappa-5; n=7). Control groups of mice received PBS (naive; n=6), 1,000 untreated wild type eggs (WT; n=7) or 1,000 eggs previously transduced with a lentivirus lacking the shRNAmir stem (that is, empty-vector (EV); n=6). All mice were euthanized 15 days after injection. (a) Total leukocyte numbers in cell suspensions from the right lung; cell populations: eosinophil granulocytes (EG), neutrophil granulocytes (NG), dendritic cells (DC), T-helper cells (Th), B cells (BC), interstitial macrophages (IM) or alveolar macrophages (AM). The percentage of leukocyte populations was established by flow cytometry (see Supplementary Fig. 2 for cell gating) and calculated as the total cell number of live cells isolated from the right lung from individual mice. Box plot representing the median and upper/lower quartile. Whiskers indicate the highest or lowest value. Kruskal–Wallis and Dunns post-hoc tests were used to establish statistical significance compared with the EV control: **P<0.01. (b) Effect of omega-1, ipse or kappa-5 knockdown of S. mansoni eggs on total serum IgE levels in mice 15 days following injection of eggs. (c) Effect of egg knockdown (kd) on granuloma formation in the left lung. Granuloma sizes were determined as the ratio of the granuloma area to the egg area. Scatter plot with mean±s.e.m. of the data pool representing all granulomata from all lungs from each experimental group (WT: n=48; EV: n=82; omega-1 kd: n=77; ipse kd: n=74; kappa-5 kd: n=84). Kruskal–Wallis and Dunns post-hoc tests were used to establish statistical significance compared with the EV control: *P<0.05, **P<0.01, ***P<0.001; not significant (ns). Numbers of mice: n=4 (WT); n=5 (EV, omega-1 kd); n=6 (ipse kd) and n=7 (kappa-5 kd). (d) Mean granuloma size in left lung sections from mice examined microscopically (original magnification: × 200), following staining with Masson–Goldner Trichrome. Scale bars, 100 μM. Arrows indicate S. mansoni eggs.
Lung cells of BALB/c mice were analysed for cytokine secretion 15 days after injection, via the tail vein, with PBS (naive), eggs of S. mansoni treated with no virus (WT), empty-vector (EV), or lentiviruses containing shRNAmirs directed at omega-1, ipse or kappa-5 transcripts. Lung cell suspensions were restimulated with SEA (20 μg ml−1) for 72 h and culture supernatants (n=5–7) assayed by cytometric bead array. Box plot representing the median and upper/lower quartile. Whiskers indicate the highest or lowest value. IL, interleukin; IFN, interferon; kd, knockdown; nd, not detected.
Finally, we assessed whether gene knockdown for any of the three egg proteins had an effect on granuloma formation in the lungs of mice (Fig. 3c). Knockdown of omega-1 in S. mansoni eggs led to a highly significant (P<0.001) decrease in granuloma size compared with the EV and untreated, WT control groups, with mean±s.e. of granuloma/egg ratios of 6.505±0.963, 19.75±1.685 and 16.63±1.597, respectively. Furthermore, granulomata in the omega-1 knockdown group (6.505±0.963) were significantly (P<0.01; Kruskal–Wallis and Dunns post-hoc tests) smaller than those in ipse (10.31±1.162) and kappa-5 (12.77±1.648) knockdown groups. No significant differences in granuloma size were measured between mice injected with WT (19.75±1.685) or control virus-treated eggs (16.63±1.597; Fig. 3c). Granulomata found in the lungs of the kappa-5 knockdown group exhibited an increase in collagen deposition (Fig. 3d).
Discussion
Here we demonstrated lentiviral transduction of S. mansoni eggs for the delivery of a shRNAmir expression cassette. This approach probably overcomes some of the limitations of conventional RNAi (for example, by soaking or electroporation), including a lack of persistent knockdown over a longer period of time (which is required for in vivo application in animals) and possible off-target effects of RNAi triggers. Although double-stranded RNA and siRNA usually achieve knockdown for periods of up to 4 weeks in S. mansoni in vitro16,17, it has been shown that siRNA does not induce a long-term effect in vivo when schistosomules are re-introduced into the definitive host17. The findings of the present study show that the integration of a lentivirus encoding an RNAi-trigger expression cassette circumvents this limitation to achieve sustained knockdown in S. mansoni in vivo in mice. The results obtained using this method also indicate higher specificity of gene knockdown compared with conventional RNAi. In contrast to previous γ-retrovirus-based methods employed to deliver shRNAs to S. mansoni18,19,20, the lentiviral delivery system can transduce both arrested and dividing cells to maximize knockdown efficiency. Furthermore, the expression of shRNAs as artificial primary miRNAs with a Drosha and Dicer processing sites has been shown to diminish the cytotoxic effect of a given short hairpin RNA (shRNA)21. The thermodynamic design of hairpins allows for a biased incorporation of the antisense strand into the RNA-induced silencing complex22. This approach decreases the risk of off-target effects induced by the sense (miRNA*) strand and enables off-target gene predictions to be made based on the antisense (miRNA) strand sequence. Moreover, the expression of shRNAs as shRNAmirs can increase target gene knockdown by up to tenfold23. Therefore, the use of more efficient shRNAmirs might also allow the application of weak promoters to minimize possible cytotoxicity linked to a saturation of the RNAi pathway24,25, and of low virus copy numbers to reduce the risk of adverse effects due to viral integration and promoter/enhancer interactions with endogenous genes, as shown previously for γ-retroviruses26. To avoid adverse effects resulting from an abundance of the RNAi trigger, shRNAmir expression levels need to be controlled. Such controlled expression of the RNAi trigger can be realized by using pol II-driven shRNAmir expression cassettes (as shown here), as pol II promoter activity can be constrained by regulatory elements27 or tissue-specific endogenous miRNAs28.
Using the present lentiviral transduction system, we demonstrated that omega-1 knockdown in S. mansoni eggs led to a significant decrease in granuloma size in mice. The increased infiltration of DCs, T-helper and B cells, and interstitial macrophages seen in the lungs of EV control mice was consistent with that expected in egg-induced disease29 and was clearly abrogated by omega-1 knockdown. Interestingly, although a decrease in B and T cells was detected in omega-1 knockdown groups, the cytokine profile remained unaltered following re-stimulation with SEA. This finding might relate to the fact that cell numbers in the re-stimulation assay were normalized against leukocyte numbers and not against total cell numbers in lung cell suspensions. Nonetheless, considering the largely diminished infiltration of effector cells in mice injected with omega-1 knockdown eggs, it is likely to be that the inflammatory milieu in the lungs is reduced compared with WT and EV control groups. However, the diminished infiltration of B and T cells, as well as macrophages, raises questions as to which cells are responsible for the cytokine secretion and how these cells are activated. Candidate innate immune cells are eosinophils (interleukin (IL)-4 and IL-13), mast cells (IL-1, IL-6 and IL-13) and basophils (IL-4 and IL-13). Moreover, natural killer T cells have been reported as a source of IL-4 and IL-13 (ref. 30). The data from the present and other studies of granuloma formation31,32,33 indicate that, although an anti-inflammatory environment can be established in the absence of CD4+T cells, Th2 effector cells are required to boost the IL-4 and IL-13 dominant cytokine milieu to induce granuloma formation.
The present study also suggests that the cytotoxicity of omega-1 (ref. 34) is a critical factor in the initiation of granuloma formation, leading to tissue destruction and activation of an innate immune response, which is maintained and amplified by Th2 cells. Indeed, Loke et al.33 showed that alternative activation is an innate response to injury that requires CD4+ T cells to be sustained during a chronic infection. Therefore, the change in macrophage numbers on knockdown suggests a new role for omega-1 relating to the recruitment of macrophages into tissues and involvement in fibrosis/granuloma formation35. Alternatively activated macrophages (AAMs) play a critical role in the prevention of severe disease development during S. mansoni infection, as they are essential for both wound-healing processes36 and the suppression of severe fibrosis37. Furthermore, macrophage-derived transforming growth factor-β1 has been linked to the activation of collagen production associated with healing and fibrosis38,39. Moreover, AAMs upregulate expression of the mannose receptor40, the receptor responsible for the uptake of omega-1 by DCs and its modulatory effect on DCs41. Thus, it would be informative to investigate whether omega-1 can modulate AAMs to promote their immunosuppressive effect(s) after binding to the mannose receptor.
Despite the diminished cell infiltration into lung tissues, granuloma formation was not entirely abolished in the omega-1 knockdown group. A possible explanation is an incomplete suppression of omega-1 expression in eggs linked to the formation of small granulomata. Interestingly, IPSE has also been shown to be cytotoxic42 and might be responsible for the induction of smaller granulomata in the omega-1 knockdown group. However, as IPSE is the most abundant egg-secreted protein43,44, it is conceivable that, in the case of an assumed IPSE-induced cytotoxicity, the impact on granuloma formation might have been more severe. A possible scenario is that, in the absence of omega-1, IPSE can activate basophils45 to secrete IL-4 and IL-13 at levels that are not excessive, because they are not amplified by Th2 effector cells. Therefore, IPSE might contribute to wound-healing processes to counter-regulate minor tissue damage caused by translocating eggs. The moderate knockdown of ipse transcripts and the observed phenotype might reflect the presence of multiple copies of the ipse gene in the genome of S. mansoni.
Interestingly, we found that large granulomata induced by kappa-5 knockdown eggs contained more collagen compared with the WT and EV control groups (cf. Fig. 3d). Although the collagen content in tissues was not quantified, the data suggest that kappa-5 has a modulatory, anti-fibrotic effect and is involved in the regulation of fibrosis. Owing to its molecular weight, kappa-5 is retained within schistosome eggs46 and might only become accessible to the immune system when eggs disintegrate. However, in the present study, both the eggshell and the miracidium appeared to be intact, contradicting this theory. As kappa-5 can bind to C-type lectin receptors47, it would be interesting to assess whether this protein has a modulatory effect on macrophages that regulates collagen synthesis. Although kappa-5 is the only molecule presently known to induce IgE responses in people with schistosomiasis46, kappa-5 knockdown did not lead to a reduction in total serum-IgE levels in mice in the present study (Fig. 3b). However, an effect on kappa-5-specific serum IgE levels cannot be excluded, such that further work is warranted to address this aspect.
In conclusion, using a functional genomic–phenomic approach, the present study provides evidence that a key mechanism underlying S. mansoni egg-induced pathological changes is tissue destruction caused by omega-1, facilitated by its immunomodulatory capacity. This study paves the way for future investigations of schistosomiasis using complementary post-genomic tools. Clearly, the lentivirus transduction system applied here, for the first time to any parasite, has major potential to be used broadly for functional genomic–phenomic investigations of socioeconomically important eukaryotic parasites.
Methods
Maintenance of S. mansoni
All experiments were approved by the Animal Ethics Committee of the University of Melbourne and performed in quarantine-approved facilities. The NMRI strain of S. mansoni was maintained in Biomphalaria glabrata (NMRI strain, NIH-NIAID Schistosomiasis Resource Center, USA; http://www.schisto-resource.org) and female, specific pathogen-free BALB/c mice (7–8 weeks of age)48.
Lentivirus vector and production
The lentivirus vector pGIPZ_mCherry was based on pGIPZ (Open Biosystems) in which shRNA constructs are expressed as human microRNA-30 (miR30) primary transcripts (shRNAmir). The CMV turboGFP was substituted (via the XbaI and NotI sites) with the mCherry reporter gene sequence from pFPVmCherry49. The CMV promoter was inserted into the pGIPZmCherry construct (via XbaI and SpeI) to yield pGIPZ_CMVmCherry. The IRES/puromycin resistance region was extracted from the plasmid (via NotI and MluI) and substituted with the miR30 flanks produced by conventional PCR. The EcoRI restriction site encoded in the miR30 flank sequence was deleted by PCR. To do this, the 3′-miR30 region was amplified without the EcoRI sequence, and the original 3′-miR30 region was substituted with the ΔEcoRI amplicon (via BamHI and MluI restriction sites). All plasmid constructs were maintained in Escherichia coli (strain Sfbl3; Invitrogen). Virus-cassette sequences were verified by automated, capillary (Sanger) sequencing.
Virus was produced in TLA-HEK293 cells using the Trans-Lenti shRNA Packaging Kit (Open Biosystems) with calcium phosphate. Virus titres were estimated by quantitative PCR50 using primers F (5′- TGGACAGGGGCTCGGCTGTT -3′) and R (5′- TCCGCTGGATTGAGGGCCGA -3′), following the removal of residual plasmid DNA by treatment with DNaseI (NEB). The functional virus titre of the calibrator (that is, EV control virus) was determined by calculating fluorescent cell colonies according to the manufacturer’s instructions (Open Biosystems). Resultant regression fit equations were used to estimate unknown titres; generally, virus titres of 1–5 × 106 transducing units per ml were used.
Cloning and validation of shRNAmir constructs
Novel, mature, artificial miRNA sequences encoded in the antisense strand of the shRNAmir stem and targeting coding regions of relevant messenger RNAs were inferred using the algorithm DSIR51. Each shRNAmir was designated according to the binding site in the mature sequence of the respective mRNA target. For each target gene, various shRNAmir sequences were initially selected and individually cloned into the lentiviral shRNAmir expression vector27,52 via XhoI and BamHI sites. The sequence of each cloned insert was verified by Sanger sequencing before testing in COS7 cells.
The full-length cDNA sequences of omega-1, ipse and kappa-5, spanning the signal peptide-coding region and 3′-untranslated region (3′-UTR), were individually PCR-amplified from cDNA and introduced into the mammalian expression vector pcDNA3.1 myc-His (Invitrogen) via HindIII and XbaI. Insertions were each verified by sequencing. Protein-expressing cell lines were established by transfecting COS7 cells with linearized pcDNA3.1 plasmids using the Optifect reagent (Invitrogen) and selected using G418 (Invitrogen) over 7–10 days. Clones were analysed for transgene transcription by conventional PCR from cDNA. Next, the functionality of individual shRNAmirs was assessed by transfecting COS7 cell lines with lentiviral vector plasmids encoding shRNAmirs or the empty (control) plasmid using Lipofectamine LTX PLUS reagent (Invitrogen). Target gene transcription was assessed 48 h after transfection by qPCR53.
Inference of shRNAmir target sequence specificity
Genes in the S. mansoni genome (http://www.genedb.org/Homepage/Smansoni) were screened for potential off-target sites for the six mature artificial miRNAs used in this study (that is, omega-1 shRNAmir-511 and shRNAmir-557; ipse shRNAmir-195 and shRNAmir-384; kappa-5 shRNAmir-616 and shRNAmir-638) based on homology to annotated 3′-UTRs; 3′-UTR sequences sharing complementarity to the mature artificial miRNA ‘seed’ region (nt 2–8) were predicted to be potential off-target sites54,55,56. The sequences for the mature artificial miRNAs (5′–3′) used here were as follows: UUGAUUGUAUCAUAUUGUCUG (shRNAmir-511), UGCAGAACCAUUGUAACCGAA (shRNAmir-557), UAUGGACGCUCUCUCUUCGUG (shRNAmir-195), UAGAAUCGACAGUAUGUCCUU (shRNAmir-384), UUUAGUUCGGUGUAUCUUCAU (shRNAmir-616) and UUGACCUACAGUCAACCUCGG (shRNAmir-638).
Transduction of S. mansoni eggs
Following their isolation from mouse livers48, schistosome eggs were washed extensively in sterile PBS; individual batches of 3,000 eggs were then exposed to lentivirus particles at a multiplicity of infection of 10, or were not exposed (WT control) in 500 μl of serum-free DMEM (Gibco) and 8 μg ml−1 of polybrene (Sigma). Virus without shRNAmir stem (that is, EV) or viruses each containing the gene-specific shRNAmir stems omega-1 shRNAmir-511 or shRNAmir-557; ipse shRNAmir-195 or shRNAmir-384; and kappa-5 shRNAmir-616 or shRNAmir-638 were used. After an incubation for 24 h at 37 °C in 5% CO2, individual egg batches were washed extensively in PBS (37 °C) to remove virus particles and polybrene. Then, individual egg batches (n=3,000) were cultured (for 2–9 more days) in DMEM with 10% FCS and 2 mM L-glutamine and egg genomic DNA isolated with TRIzol (Invitrogen). Subsequently, the presence of viral DNA was verified by conventional PCR using specific primer sets and Southern blot analysis57 of genomic DNA of S. mansoni. The maturity and viability of eggs was monitored daily by microscopic examination. Hatching of eggs was induced 10 days after virus exposure and monitored microscopically.
Assessing gene knockdown following transduction
The knockdown of target gene transcription in cultured S. mansoni eggs was assessed by qPCR. To do this, total RNA was isolated from individual batches of 3,000 eggs using TRIzol (Invitrogen) and residual DNA removed by DNase I treatment; cDNA was synthesized from 250 ng of RNA by reverse transcription using random primers and MMLV reverse transcriptase (Bioline). Following the calculation of PCR efficiencies58, which ranged between 1.9 and 2.1, the relative transcription (R) of individual target genes was related to EV control samples (that is, mean Ct value), and transcription was normalized against those of cox-1, ef1-a and psdm-4 after 3 days of in vitro culture; or act-b59 after 10 days of in vitro culture, according to the efficiency-corrected calculation model for multiple samples60. The suitability of cox-1, ef1-a and psdm-4 was verified using the geNorm tool of the qbasePLUS software (Biogazelle), employing established principles61.
Mouse experimentation and pathological changes
Following the verification of gene knockdown, virus-treated and untreated egg batches were introduced into mice under sterile condition to then assess alterations in their lungs and sera. To do this, 1,000 eggs in 100 μl of sterile PBS were injected into the lateral tail vein of female BALB/c mice (7–8 weeks of age)15. Individual mice in treatment groups were injected with eggs previously transduced with lentiviruses containing gene-specific shRNAmir stems (omega-1, ipse or kappa-5). Mice in control groups were each injected with PBS (naive), untreated eggs (WT) or with eggs previously transduced with a lentivirus lacking the shRNAmir stem (that is, EV). Mice were euthanized 15 days after injection; immediately, individual lungs were perfused with 10 ml of sterile PBS (4 °C) via the heart, after which they were extracted from the thoracic cavity, and connective tissue and mediastinal lymph nodes trimmed off. Then, four investigatory components were conducted:
First, from individual right-lung lobes, single-cell suspensions were prepared62. Residual erythrocytes were lysed using ACK solution (Gibco) and cells resuspended in PBS containing 1% w/v of BSA. Total leukocyte counts were established by flow cytometry (FACS Calibur, BD Bioscience, USA) using Countbright absolute counting beads (Invitrogen); dead cells were excluded using 7-amino-actinomycin D (Sigma). Lung cell suspensions were incubated in FACS-staining buffer (PBS containing 1% of BSA and 5 mM of EDTA) with LEAF-purified anti-mouse CD16/32 (clone 93; BioLegend) and the following antibodies (BioLegend): allophycocyanin-conjugated anti-CD11c (clone N418, eBioscience), -CD3e (clone 145-2C11); R-phycoerythrin-conjugated anti-CD4 (clone RM4-5, Invitrogen), -F4/80 (clone BM8), -Siglec-F (clone E50-2440, BD Bioscience); fluorescein isothiocyanate-conjugated anti-B220 (clone RA3-6B2), -Ly-6G/Ly-6C (RB6-8C5). All antibodies were used at a 1:200 dilution, except anti-mouse F4/80 (1:100). Pooled cells from all mice of individual groups were also stained with antibody mixes for the respective isotypes. The LIVE/DEAD fixable Red stain kit (Invitrogen) was used to exclude dead cells. Flow cytometric acquisitions were carried out with a FACS Calibur and CellQuest software (BD) and analysed with FlowJo (Tree Star) (Cell gating; see Supplementary Fig. 2). Second, left-lung lobes were subjected to histopathological examination. Individual lobes were fixed in 10% normal-buffered formalin (pH 7.4) or embedded in Tissue-Tek OCT compound (Sakura Finetek), serially sectioned (5–8 μm), stained with haematoxylin–eosin or Masson’s Trichrome stain and then mounted in DPX (Sigma). Samples were examined for granulomata containing a centrally sectioned egg using a BX60 microscope (Olympus) equipped with a SPOT camera (Diagnostic Instruments) and Image-Pro software (MediaCybernetics). Subsequently, for individual granulomata, the granuloma area and the egg area were measured by computerized morphometry and the ratio between the two areas calculated. Third, total IgE levels in sera from individual mice were measured using the Ready-Set-Go! mouse IgE ELISA kit (eBioscience), according to the manufacturer’s instructions, using a Synergy H1 Hybrid Reader (BioTek). Fourth, lung cell suspensions were restimulated with SEA (20 μg ml−1) for 72 h, and culture supernatants assayed using the Mouse Th1/Th2/Th17/Th22 13-plex Kit FlowCytomix (eBioscience).
Statistical analyses
Treatment groups were analysed for significant differences using the Kruskal–Wallis one-way analysis of variance (P<0.05) and Dunn post-hoc tests63 in relation to the EV control group (n=5–7) using Prism software (Graph-Pad Software Inc.).
Additional information
How to cite this article: Hagen, J. et al. Omega-1 knockdown in Schistosoma mansoni eggs by lentivirus transduction reduces granuloma size in vivo. Nat. Commun. 5:5375 doi: 10.1038/ncomms6375 (2014).
References
Rollinson, D. A wake up call for urinary schistosomiasis: reconciling research effort with public health importance. Parasitology 136, 1593–1610 (2009).
Burke, M. L. et al. Immunopathogenesis of human schistosomiasis. Parasite Immunol. 31, 163–176 (2009).
Berriman, M. et al. The genome of the blood fluke Schistosoma mansoni. Nature 460, 352–358 (2009).
Young, N. D. et al. Whole-genome sequence of Schistosoma haematobium. Nat. Genet. 44, 221–225 (2012).
Hagen, J., Lee, E. F., Fairlie, W. D. & Kalinna, B. H. Functional genomics approaches in parasitic helminths. Parasite Immunol. 34, 163–182 (2012).
Rinaldi, G. et al. Development of functional genomic tools in trematodes: RNA interference and luciferase reporter gene activity in Fasciola hepatica. PLoS Negl. Trop. Dis. 2, e260 (2008).
Rinaldi, G. et al. Germline transgenesis and insertional mutagenesis in Schistosoma mansoni mediated by murine leukemia virus. PLoS Pathog. 8, e1002820 (2012).
Mann, V. H., Morales, M. E., Kines, K. J. & Brindley, P. J. Transgenesis of schistosomes: approaches employing mobile genetic elements. Parasitology 135, 141–153 (2008).
Dalzell, J. J. et al. Considering RNAi experimental design in parasitic helminths. Parasitology 139, 589–604 (2012).
Geldhof, P. et al. RNA interference in parasitic helminths: current situation, potential pitfalls and future prospects. Parasitology 134, 609–619 (2007).
Manjunath, N., Wu, H., Subramanya, S. & Shankar, P. Lentiviral delivery of short hairpin RNAs. Adv. Drug Deliv. Rev. 61, 732–745 (2009).
Schramm, G. & Haas, H. Th2 immune response against Schistosoma mansoni infection. Microbes Infect. 12, 881–888 (2010).
Fairfax, K., Nascimento, M., Huang, S. C., Everts, B. & Pearce, E. J. Th2 responses in schistosomiasis. Semin. Immunopathol. 34, 863–871 (2012).
Hams, E., Aviello, G. & Fallon, P. G. The Schistosoma granuloma: friend or foe? Front. Immunol. 4, 89 (2013).
Boros, D. L. & Warren, K. S. Delayed hypersensitivity-type granuloma formation and dermal reaction induced and elicited by a soluble factor isolated from Schistosoma mansoni eggs. J. Exp. Med. 132, 488–507 (1970).
Correnti, J. M., Brindley, P. J. & Pearce, E. J. Long-term suppression of cathepsin B levels by RNA interference retards schistosome growth. Mol. Biochem. Parasitol. 143, 209–215 (2005).
Krautz-Peterson, G. et al. Suppressing glucose transporter gene expression in schistosomes impairs parasite feeding and decreases survival in the mammalian host. PLoS Pathog. 6, e1000932 (2010).
Kines, K. J. et al. Electroporation facilitates introduction of reporter transgenes and virions into schistosome eggs. PLoS Negl. Trop. Dis. 4, e593 (2010).
Tchoubrieva, E. B., Ong, P. C., Pike, R. N., Brindley, P. J. & Kalinna, B. H. Vector-based RNA interference of cathepsin B1 in Schistosoma mansoni. Cell Mol. Life Sci. 67, 3739–3748 (2010).
Duvoisin, R. et al. Human U6 promoter drives stronger shRNA activity than its schistosome orthologue in Schistosoma mansoni and human fibrosarcoma cells. Transgenic Res. 21, 511–521 (2012).
McBride, J. L. et al. Artificial miRNAs mitigate shRNA-mediated toxicity in the brain: implications for the therapeutic development of RNAi. Proc. Natl Acad. Sci. USA 105, 5868–5873 (2008).
Khvorova, A., Reynolds, A. & Jayasena, S. D. Functional siRNAs and miRNAs exhibit strand bias. Cell 115, 209–216 (2003).
Silva, J. M. et al. Second-generation shRNA libraries covering the mouse and human genomes. Nat. Genet. 37, 1281–1288 (2005).
Grimm, D. The dose can make the poison: lessons learned from adverse in vivo toxicities caused by RNAi overexpression. Silence 2, 8 (2011).
Zychlinski, D. et al. Physiological promoters reduce the genotoxic risk of integrating gene vectors. Mol. Ther. 16, 718–725 (2008).
Nienhuis, A. W., Dunbar, C. E. & Sorrentino, B. P. Genotoxicity of retroviral integration in hematopoietic cells. Mol. Ther. 13, 1031–1049 (2006).
Dickins, R. A. et al. Tissue-specific and reversible RNA interference in transgenic mice. Nat. Genet. 39, 914–921 (2007).
Liu, Y. P. & Berkhout, B. miRNA cassettes in viral vectors: problems and solutions. Biochim. Biophys. Acta 1809, 732–745 (2011).
Pearce, E. J. & MacDonald, A. S. The immunobiology of schistosomiasis. Nat. Rev. Immunol. 2, 499–511 (2002).
Akbari, O. et al. Essential role of NKT cells producing IL-4 and IL-13 in the development of allergen-induced airway hyperreactivity. Nat. Med. 9, 582–588 (2003).
Doenhoff, M. J. et al. Immunological control of hepatotoxicity and parasite egg excretion in Schistosoma mansoni infections: stage specificity of the reactivity of immune serum in T-cell deprived mice. Trans. R. Soc. Trop. Med. Hyg. 75, 41–53 (1981).
Kaplan, M. H., Whitfield, J. R., Boros, D. L. & Grusby, M. J. Th2 cells are required for the Schistosoma mansoni egg-induced granulomatous response. J. Immunol. 160, 1850–1856 (1998).
Loke, P. et al. Alternative activation is an innate response to injury that requires CD4+ T cells to be sustained during chronic infection. J. Immunol. 179, 3926–3936 (2007).
Dunne, D. W. et al. Identification and partial purification of an antigen (omega 1) from Schistosoma mansoni eggs which is putatively hepatotoxic in T-cell deprived mice. Trans. R. Soc. Trop. Med. Hyg. 75, 54–71 (1981).
Wynn, T. A. & Ramalingam, T. R. Mechanisms of fibrosis: therapeutic translation for fibrotic disease. Nat. Med. 18, 1028–1040 (2012).
Herbert, D. R. et al. Alternative macrophage activation is essential for survival during schistosomiasis and downmodulates T helper 1 responses and immunopathology. Immunity 20, 623–635 (2004).
Pesce, J. T. et al. Arginase-1-expressing macrophages suppress Th2 cytokine-driven inflammation and fibrosis. PLoS Pathog. 5, e1000371 (2009).
Lee, C. G. et al. Interleukin-13 induces tissue fibrosis by selectively stimulating and activating transforming growth factor beta(1). J. Exp. Med. 194, 809–821 (2001).
Wynn, T. A. Cellular and molecular mechanisms of fibrosis. J. Pathol. 214, 199–210 (2008).
Stein, M., Keshav, S., Harris, N. & Gordon, S. Interleukin 4 potently enhances murine macrophage mannose receptor activity: a marker of alternative immunologic macrophage activation. J. Exp. Med. 176, 287–292 (1992).
Everts, B. et al. Schistosome-derived omega-1 drives Th2 polarization by suppressing protein synthesis following internalization by the mannose receptor. J. Exp. Med. 209, 1753–1767 S1751 (2012).
Abdulla, M. H., Lim, K. C., McKerrow, J. H. & Caffrey, C. R. Proteomic identification of IPSE/alpha-1 as a major hepatotoxin secreted by Schistosoma mansoni eggs. PLoS Negl. Trop. Dis. 5, e1368 (2011).
Cass, C. L. et al. Proteomic analysis of Schistosoma mansoni egg secretions. Mol. Biochem. Parasitol. 155, 84–93 (2007).
Mathieson, W. & Wilson, R. A. A comparative proteomic study of the undeveloped and developed Schistosoma mansoni egg and its contents: the miracidium, hatch fluid and secretions. Int. J. Parasitol. 40, 617–628 (2010).
Schramm, G. et al. Molecular characterization of an interleukin-4-inducing factor from Schistosoma mansoni eggs. J. Biol. Chem. 278, 18384–18392 (2003).
Schramm, G. et al. Molecular characterisation of kappa-5, a major antigenic glycoprotein from Schistosoma mansoni eggs. Mol. Biochem. Parasitol. 166, 4–14 (2009).
Meevissen, M. H. et al. Specific glycan elements determine differential binding of individual egg glycoproteins of the human parasite Schistosoma mansoni by host C-type lectin receptors. Int. J. Parasitol. 42, 269–277 (2012).
Mann, V. H., Morales, M. E., Rinaldi, G. & Brindley, P. J. Culture for genetic manipulation of developmental stages of Schistosoma mansoni. Parasitology 137, 451–462 (2010).
Drecktrah, D. et al. Dynamic behavior of Salmonella-induced membrane tubules in epithelial cells. Traffic 9, 2117–2129 (2008).
Mann, V. H., Suttiprapa, S., Rinaldi, G. & Brindley, P. J. Establishing transgenic schistosomes. PLoS Negl. Trop. Dis. 5, e1230 (2011).
Vert, J.-P., Foveau, N., Lajaunie, C. & Vandenbrouck, Y. An accurate and interpretable model for siRNA efficacy prediction. BMC Bioinformatics 7, 520 (2006).
Paddison, P. J. et al. Cloning of short hairpin RNAs for gene knockdown in mammalian cells. Nat. Methods 1, 163–167 (2004).
Liu, S. et al. Genome-wide identification and characterization of a panel of house-keeping genes in Schistosoma japonicum. Mol. Biochem. Parasitol. 182, 75–82 (2012).
Anderson, E. M. et al. Experimental validation of the importance of seed complement frequency to siRNA specificity. RNA 14, 853–861 (2008).
Birmingham, A. et al. 3' UTR seed matches, but not overall identity, are associated with RNAi off-targets. Nat. Methods 3, 199–204 (2006).
Jackson, A. L. et al. Widespread siRNA "off-target" transcript silencing mediated by seed region sequence complementarity. RNA 12, 1179–1187 (2006).
Southern, E. M. Southern blotting. Nat. Protoc. 1, 518–525 (2006).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, e45 (2001).
Rinaldi, G. et al. RNA interference targeting leucine aminopeptidase blocks hatching of Schistosoma mansoni eggs. Mol. Biochem. Parasitol. 167, 118–126 (2009).
Pfaffl, M. W. in:A-Z of Quantitative PCR 1st edition ed. Bustin S. A. International University Line (2004).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, RESEARCH0034 (2002).
Loebbermann, J. et al. IL-10 regulates viral lung immunopathology during acute respiratory syncytial virus infection in mice. PLoS ONE 7, e32371 (2012).
Quinn, G. P. & Keough, M. J. Experimental Design and Data Analysis for Biologists Cambridge University Press (2002).
Acknowledgements
Funding from the National Health and Medical Research Council (NHMRC) and the Australian Research Council (ARC) is gratefully acknowledged. This project was also partially supported by a Victorian Life Sciences Computation Initiative (VLSCI; grant number VR0007) on its Peak Computing Facility at the University of Melbourne, an initiative of the Victorian Government. We thank Dr Ross Dickins (Walter and Eliza Hall Institute of Medical Research) for discussion and support to J.H. early in her study. J.H. was the recipient of MIRS and MIFRS scholarships from the University of Melbourne. N.D.Y. is an NHMRC Early Career Research Fellow (ECRF).
Author information
Authors and Affiliations
Contributions
J.H. conceived and conducted the study, under the supervision of R.B.G. and B.H.K., with support from N.D.Y., A.L.E., C.N.P., C.S. and J.-P.Y.S.; J.H. and R.B.G. drafted the text and J.-P.Y.S., N.D.Y. and R.B.G. edited the manuscript, with inputs from other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
Supplementary Figures 1-2. (PDF 202 kb)
Supplementary Movie
Vitality of miracidia within eggs. (AVI 31507 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Hagen, J., Young, N., Every, A. et al. Omega-1 knockdown in Schistosoma mansoni eggs by lentivirus transduction reduces granuloma size in vivo. Nat Commun 5, 5375 (2014). https://doi.org/10.1038/ncomms6375
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms6375
This article is cited by
-
Genetic manipulations in helminth parasites
Journal of Parasitic Diseases (2023)
-
Transgenesis in parasitic helminths: a brief history and prospects for the future
Parasites & Vectors (2022)
-
Granulomas in parasitic diseases: the good and the bad
Parasitology Research (2020)
-
Investigating immune responses to parasites using transgenesis
Parasites & Vectors (2019)
-
Profiling microRNAs through development of the parasitic nematode Haemonchus identifies nematode-specific miRNAs that suppress larval development
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.