Abstract
Endonuclease V orthologues are highly conserved proteins found in all kingdoms of life. While the prokaryotic enzymes are DNA repair proteins for removal of deaminated adenosine (inosine) from the genome, no clear role for the eukaryotic counterparts has hitherto been described. Here we report that human endonuclease V (ENDOV) and also Escherichia coli endonuclease V are highly active ribonucleases specific for inosine in RNA. Inosines are normal residues in certain RNAs introduced by specific deaminases. Adenosine-to-inosine editing is essential for proper function of these transcripts and defects are linked to various human disease. Here we show that human ENDOV cleaves an RNA substrate containing inosine in a position corresponding to a biologically important site for deamination in the Gabra-3 transcript of the GABAA neurotransmitter. Further, human ENDOV specifically incises transfer RNAs with inosine in the wobble position. This previously unknown RNA incision activity may suggest a role for endonuclease V in normal RNA metabolism.
Similar content being viewed by others
Introduction
RNA editing covalently alter the nucleotide sequence of RNA transcripts relative to that of the encoding DNA, thereby contributing to gene diversity1. One of the most prevalent RNA modifications is adenosine (A) deamination, which results in conversion of the 6-aminopurine ring of A to the 6-oxopurine ring of inosine (I)2. Inosine has different base pairing properties to A and is interpreted as guanosine (G) in cells. Enzymes catalyzing this conversion are adenosine deaminases acting on RNA (ADARs)3, which are conserved enzymes found in most multicellular organisms4. Important ADAR targets are mRNAs for neurotransmitter receptors in mammals and editing is critical for normal brain development and behaviour5. In all cases, adenosine deamination results in a dramatic alteration of the receptor function5. Improper function of ADARs has been correlated with serious neurological and mental human disorders6. However, the vast majority of A to I conversions are found in non-coding regions where they are involved in controlling the activity of small RNAs such as short interfering (si)RNA and micro (mi)RNAs7,8.
Inosine is also a central component of transfer RNA (tRNA) where it is found in the wobble position (I34) in the anticodon loop of certain tRNAs9. In E. coli, only tRNAArg(ACG) undergoes A-to-I editing, whereas in eukaryotes seven to eight tRNAs contain I34 (ref. 10). Deamination of A to I is performed by adenosine-specific deaminases acting on tRNA (ADATs), which are homologues of the ADARs. I34 is crucial for decoding during protein synthesis as wobble I allow for base pairing with C, T and A9,11. Inosine in ribosomal RNA is unusual12.
While I is a normal and essential residue in RNA (rI), I in DNA (dI) is regarded as damage because of its miscoding properties. Inosine in DNA is a result of spontaneous or nitrosative stress-induced deamination of dA13. To counteract such threats, cells express DNA repair proteins specific for inosine. In E. coli, the primary enzyme for the repair of dI is endonuclease V (EndoV)14, which is encoded by the nfi gene15. EndoV initiates repair by Mg2+-dependent cleavage of the second phosphodiester bond 3′ to the lesion generating 3′-OH and 5′-P termini14,16,17. EndoV incises DNA without removing the inosine nucleotide (nt), thus completion of repair depends on additional proteins, a process that is currently poorly understood. Deaminated adenosines can also be repaired by the base excision repair pathway18,19. Homologues of EndoV are found in most species, and despite strong sequence conservation robust dI activity has only been demonstrated for the prokaryotic enzymes20,21,22,23,24,25,26.
Recently, we characterized human ENDOV without detecting any dI incision21. Interestingly, when fused to the green fluorescent protein, ENDOV was not found in the nucleus of HeLa cells as expected for a DNA repair protein. Rather, ENDOV localized to nucleoli and the cytoplasm, which are the compartments for RNA.
In the present study, we show that human ENDOV has a strong and specific incision activity on RNA substrates containing rI. Also EndoV from E. coli, which is an inosine-specific DNA repair enzyme, cleaves at inosines in RNA. Human ENDOV has a preference for single-stranded substrates, whereas E. coli EndoV is equally active on both single- and double-stranded RNA. These are the novel findings implying a role for endonuclease V in RNA metabolism.
Results
Human ENDOV is an inosine-specific ribonuclease
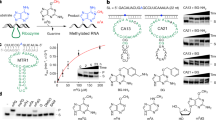
ENDOV proteins are highly conserved16, a feature that normally reflects conserved function—in this case incision at dI residues in DNA. However, human ENDOV appears to be inactive towards dI21. As inosines are not only found in DNA, but are also abundant in RNA, we hypothesized that inosines in RNA could be the substrate for ENDOV. Furthermore, the protein structure of endonuclease V from Thermotoga maritima reveals an ‘RNase H-like motif’ supporting link to RNA16. Finally, RNA as a substrate for ENDOV is also consistent with the observation of ENDOV–green fluorescent protein fusion proteins in the cytoplasm and nucleoli of HeLa cells21. Therefore, human ENDOV was purified as described and analyzed for activity on single- and double-stranded RNA oligonucleotide substrates with centrally located rI residues (Supplementary Table S1). Interestingly, human, as well as E. coli endonuclease V, efficiently cleaved both the single- (rI) and double-stranded (rI:rU) RNA substrates (Fig. 1a,b). Both endonucleases were most effective at the highest pH tested (9.5) when Mg2+ (5 mM) was used in the reaction, however, with Mn2+ as the divalent ion, both enzymes were most active at pH 7.5 (Supplementary Fig. S1a). At pH 7.5, both enzymes were active over a broad range of Mn2+ concentrations (0.25–5 mM) (Supplementary Fig. S1b). Neither of the enzymes incised an RNA substrate with a cognate rA instead of rI (Fig. 1c). Single- and double-stranded DNA substrates with inosine were cleaved by the E. coli enzyme, but not by the human ENDOV (Fig. 1d,e) as previously described21.
The substrates (a) single-stranded (ss) RNA rI, (b) double-stranded (ds) RNA rI:rU, (c) ss RNA rA, (d) ssDNA dI, (e) ds DNA dI:dT were incubated with the wild-type enzymes (1–25 nM, as indicated) or the two site-specific mutants E. coli D35A and human D52A (25 nM) and reaction products analyzed by PAGE. RNA sizes (in nt) are indicated, –=no enzyme added, r=ribonucleotide and d=deoxynucleotide. (f) Single-turnover kinetic analysis with enzyme and substrates as indicated.
Mapping of the exact position of cleavage revealed that both enzymes nicked the RNA substrate at the second phosphodiester bond 3′ to the deaminated base as expected (Supplementary Fig. S2a). Single-turnover kinetic analyses revealed a 1.3-fold higher turnover rate (kobs) for human ENDOV on single-stranded than on double-stranded RNA, whereas E. coli EndoV showed the same turnover rate for incision of both the substrates (Fig. 1f and Supplementary Table S2). Inosine-containing DNA appears to be the preferred substrate for E. coli EndoV, as the turnover rate on inosine-containing single-stranded DNA was twice that of single-stranded RNA and five times higher for double-stranded DNA versus double-stranded RNA (Supplementary Table S2).
RNAs are transcribed as single-stranded molecules but fold spontaneously under physiological conditions to adopt a secondary structure27. Thus, all four combinations of base pairs with rI may form, of which rI:rC is the most stable pair28. As double-stranded RNA with a rI:rU pair had already been tested (Fig. 1b), the three remaining base pair combinations (rI:rG, rI:rA, rI:rC) were used as substrates in activity assays. E. coli EndoV cleaved all double-stranded RNA substrates with lowest affinity for the rI:rC substrate (Supplementary Fig. S2b–d). As is the case for the rI:rU substrate (Fig. 1b), human ENDOV also incised the three other double-stranded RNA substrates (Supplementary Fig. S2b–d).
Site-specific ENDOV mutants were tested for ribonuclease activity. Mutants of the conserved catalytic aspartates (E. coli: D35A and human: D52A) could not incise the rI substrates (Fig. 1a–e, Supplementary Fig. S2b–d). Tyrosine 91 of human ENDOV corresponds to Y80 of T. maritima EndoV, which is a key residue for dI recognition16. The ENDOV Y91A mutant displayed wild-type affinity for branched DNA substrates21, however, rI incision was totally abolished (Supplementary Fig. S2e). The human mutants RK (R248A, K249A) and Wedge (the four residues PYVS[90–93] mutated to four glycines) previously shown to have defective DNA binding21 were also inactive on RNA (Supplementary Fig. S2e). These results confirm that endonuclease V enzymes indeed are inosine-specific enzymes, yet the affinity for DNA versus RNA differs.
DNA with a ribonucleotide 3′ to dI is incised by ENDOV
Endonuclease V belongs to the same structural family as RNase H enzymes, which specifically cleaves the RNA strand of RNA:DNA hybrids16,29. RNA:DNA hybrids are formed during DNA replication, RNA transcription and reverse transcription, and adopt a different conformation to double-stranded DNA30. To test whether ENDOV has activity for mixed RNA:DNA substrates, two hybrid substrates RNA:DNA- and DNA:RNA-containing rI or dI, respectively, were designed. Human ENDOV cleaved only the hybrid with rI in the RNA strand, whereas E. coli EndoV cleaved both the substrates (Fig. 2a,b). From these data, we conclude that human ENDOV cleavage is strictly dependent on ribonucleotides in the inosine containing strand. Specifically, we find that ENDOV is highly active on single-stranded DNA when a single ribonucleotide is present directly following the dI residue (Fig. 2c). Hence, the 2′ OH group in this particular ribose seems to be critical and sufficient for incision activity. A model of human ENDOV with rG in this position reveals that the 2′ OH group may interact with the conserved catalytic glutamate (E100) and possibly replace an active site water molecule (Fig. 2d). This assumption is based on comparison with the corresponding coordination sphere around Mg2+ in the active site of T. maritima EndoV in complex with the cleavage product16. The difference in activity of human ENDOV between single- and double-stranded DNA with rG next to dI (Figs 1f and 2c,e), suggests that the nucleic acid helical structure is also critical for substrate processing; incision activity is only observed for double-stranded inosine substrates having A-form helices, like double-stranded RNA and RNA:DNA hybrids31, but not for double-stranded B-form DNA. A B-form DNA may have steric conflicts with surface-exposed residues in human ENDOV.
The substrates (a) ds DNA:RNA dI:rU, (b) ds RNA:DNA rI:dT, (c) ss DNA dI rG were incubated with the wild-type enzymes (0.1–25 nM, as indicated) or the two site-specific mutants E. coli D35A and human D52A (25 nM) and reaction products analyzed by PAGE. (d) Structural model of the active site of human ENDOV with a ribonucleotide. The 2′ OH group is close to the conserved glutamate (E100), as well as the active site water molecule (Wat) coordinating the Mg2+ cofactor, which bridges the 3′- and 5′-ends in the incised product. (e) The endonuclease V enzymes were tested for activity against the ds DNA dI:dT rG:dC substrate as in a–c). RNA sizes (in nt) are indicated, –=no enzyme added, r=ribonucleotide and d=deoxynucleotide.
In vivo and in vivo-like RNAs as substrates for ENDOV
The substrates used above are all random oligonucleotide sequences, and to test for activity towards an in vivo deamination target, an oligonucleotide corresponding to a part of the mouse (m)Gabra-3 transcript of the neurotransmitter GABAA was synthesized. This 38 nt RNA has an A in position 27, known to be deaminated by adenosine deaminases ADAR1 and ADAR2, and is referred to as the I/M site32. Upon folding of this RNA, rI will form a stable base pair with cytosine (C), whereas an unedited A will form a mismatch with C32. Human ENDOV, as well as the E. coli enzyme, generated an incision product of the expected size of 28 nt when assayed with the rI-containing mGabra3 substrate, whereas no activity was seen with the unedited variant (Fig. 3a,b). This result suggests that ENDOV could be involved in antagonizing the effect of the ADAR enzymes by destruction of rI-containing transcripts.
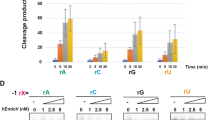
The substrates (a) mGabra3 rI, (b) mGabra3 rA, (c) tRNAArg rI, (d) tRNAArg rA were incubated with the endonuclease V enzymes (1–25 nM, as indicated) and reaction products analyzed by PAGE. RNA sizes (in nt) are indicated, –=no enzyme added, r=ribonucleotide and glyphs illustrate the different substrates.
Next, a substrate corresponding to the anticodon loop in E. coli tRNAArg(ACG), wherein the wobble base A34 is the only known A-to-I deaminated tRNA position in prokaryotes33, was tested. Cleavage within the anticodon loop of tRNAs is a well-documented response to different stress conditions, such as starvation and oxidation, in different organisms34. The rI-containing anticodon substrate, but not the unedited A-containing control substrate, was incised by both enzymes, though most efficiently by human ENDOV (Fig. 3c,d). Further, total tRNA isolated from human U373 cells, was incubated with ENDOV and examined for cleavage. Individual tRNAs were detected by Northern blot hybridization using labelled oligonucleotides complementary to the 5′-fragment of the different tRNAs as probes. Full-length processed tRNAs are 71–82 nt long and cleavage next to the wobble A/I would result in products of 35 (5′-fragment) and 36–47 (3′-fragment) nt. Specific cleavage in the anticodon loop of tRNASer(AGA), tRNALeu(AAG) and tRNAArg(ACG), all known to have rI in the anticodon loop, was demonstrated (Fig. 4a–c). In contrast, probes for tRNAAsp(GTC), tRNAGlu(CTC) and tRNALys(CTT) lacking A/I34 editing, revealed no incision products (Fig. 4d–f). No general degradation of the tRNA was seen as total tRNA remained intact after incubation with ENDOV (Supplementary Fig. S2f). These data could imply a role for ENDOV in fragmentation of tRNAs containing rI.
Northern blots of tRNA isolated from human U373 cells incubated with human wild-type (0.3 and 0.6 μM) or mutant D52A (0.6 μM) ENDOV enzymes, hybridized with DNA probes complementary to the 5′-termini of (a) tRNASer(AGA), (b) tRNALeu(AAG), (c) tRNAArg(ACG), (d) tRNAAsp(GTC), (e) tRNAGlu(CTC) and (f) tRNALys(CTT). Size markers (80, 38 and 25 nt) are shown and glyphs indicate full-length and fragment tRNA species.
Discussion
In this report, we demonstrate efficient and specific cleavage at single rI residues in RNA by human ENDOV. ENDOV has activity for both single- and double-stranded RNAs, as well as for edited tRNA anticodon loops containing rI. To our knowledge, ribonuclease activity has not been previously described for endonuclease V, and we propose that ENDOV may have an important role in RNA metabolism.
We have previously reported that human ENDOV binds to DNA with higher affinity for branched DNA structures than linear DNA21. Here we find that the nucleic acid helical structure is also of importance for ENDOV activity (Fig. 2c–e). It appears that the helical conformation (A- or B-form) is more important for binding than the type of nucleic acid (DNA or RNA), whereas catalytic activity is critically dependent on a ribonucletide in the correct position with a 2′ hydroxyl group close to the active site metal cofactor. At present, we do not know whether ENDOV binding to DNA is biologically important or whether it simply reflects a general affinity of ENDOV for nucleic acids.
Several reports demonstrate that deamination of adenosine to inosine by the ADAR enzymes has a central function in vivo5,6. We may speculate that the ribonuclease activity of ENDOV could antagonize the effect of the ADAR enzymes by specific cleavage and destruction of edited transcripts. As many ADAR targets are neurotransmitters, fine-tuning of receptor activity by regulation of the level of edited versus non-edited forms could be of major importance for optimal brain function. It should be mentioned that the Tudor-SN nuclease, which is part of the RISC complex, has also been coupled to cleavage of inosines in RNA35,36. It is proposed that Tudor-SN cleaves or promotes the cleavage of hyperedited double-stranded RNA by acting as an activator, however, its specific enzymatic properties have not been thoroughly investigated35,36.
Cleavage of tRNAs within the anticodon loop is part of the cellular response to stress conditions such as oxidation and starvation in many organisms37. Nucleases responsible for this cleavage have been identified, and defects in tRNA fragmentation are associated with apoptosis, cancer and disease progression34. As far as we know, specific cleavage of tRNAs at inosines has not previously been described, but an analogous system is the killer toxin zymocin from Kluyveromyces lactis, which depends on the wobble uridine modification 5-methoxy-carbonyl-methyl for tRNA cleavage38.
Interestingly, E. coli EndoV was also active on inosines in RNA, demonstrating that EndoV is more than a DNA repair enzyme. Enzymatic deamination of adenosines in RNA appears to be uncommon in prokaryotes, and we may speculate that a more general role exists for endonuclease V enzymes in removal of damaged/deaminated RNA transcripts. Recently, several DNA repair proteins including AlkB/ABH2, SMUG1, APE1 and TDP2 have been shown to possess activity on damaged RNA39,40,41,42,43 and this is suggested to function as a quality control for RNA. In any case, tight regulation of the ENDOV ribonuclease activity to prevent aberrant cleavage, for example, by compartmentalization, post-translational modification or interacting partners, will be of great importance.
Methods
Expression and purification of endonuclease V enzymes
Wild-type and mutants human ENDOV proteins were expressed in E. coli BL21-Codon Plus (DE3)-RIPL cells (Agilent Technologies) as fusion proteins with an N-terminal His-MBP tag21. After induction of protein expression with isopropyl β-D-1 thiogalactopyranoside (0.25 mM), bacteria were grown at 18 °C over night. Cells were harvested by centrifugation and the pellet resuspended in 50 mM Tris HCl pH 8.0, 300 mM NaCl, 10 mM imidazole and 10 mM β-mercaptoethanol (β-ME) (buffer A). Cells were lysed by sonication and the cleared lysate applied to Ni-nitrilotriacetic acid affinity chromatography. Recombinant His-MBP-ENDOV was eluted with 300 mM imidazole in buffer A. Peak fractions were pooled and dialyzed at 4 °C in 50 mM Tris HCl pH 8.0, 0.5 mM EDTA, 1 mM dithiothreitol (TEV buffer). TEV protease was added (ratio 1:100) and dialysis continued at 12 °C over night. After proteolysis, the protein mixtures were dialysed against buffer A and the free His-MBP and TEV proteins were separated from ENDOV by a second Ni-NTA purification step. The untagged ENDOV proteins were collected in the flow-through and wash fractions, concentrated and applied to a Superdex 75 size-exclusion chromatography column (GE Healthcare) equilibrated with 50 mM Tris HCl pH 8.0, 50 mM NaCl and 10 mM β-ME. Purified ENDOV was concentrated and stored at −20 °C. The two inactive endonuclease V mutants E. coli D35A and human D52A were made by site-specific mutagenesis using the forward primers 5′-ACCGGATCTGATCGCCGGAGCCGCTGTCGGGTTTGAGCAGGGC-3′ for D35A and 5′-CAGCGTGTGGGCGGTGTGGCTGTTAGTTTCGTGAAAGGTG-3′ for D52A with their corresponding reverse and complementary oligonucleotides (mutated codons are underlined).
DNA/RNA substrates
All DNA/RNA substrates were oligonucleotides synthesized by The Midland Certified Reagent Company, and are listed in Supplementary Table S1. The oligonucleotides were 5′-end labelled with [γ-32P]ATP (Perkin Elmer) using T4 polynucleotide kinase (New England BioLabs). Polynucleotide kinase was inactivated by the addition of 10 mM EDTA. Double-stranded substrates were made by adding the complementary strand and heating to 55 °C followed by slowly cooling to room temperature. The DNA and RNA substrates were separated by 20% native polyacrylamide gel electrophoresis (PAGE; LongRanger Gel Solution, Lonza, 0.5xTBE), excised from the gel, eluted by diffusion in H2O and stored at 4 °C.
DNA and RNA incision activities and single-turnover analysis
The DNA and RNA nicking assays were performed with 1 nM substrate, 0.1–25 nM of enzyme (as indicated in the figures) and standard reaction buffer (10 mM Tris-HCl pH 7.5, 0.5 mM MnCl2, 50 mM KCl, 1 mM dithiothreitol and 5% glycerol) in a total volume of 10 μl or with changes are indicated. To adjust for pH, Tris-HCl, pH 7.5, 8.5 or 9.5 was used and divalent ions kept at 5 mM. When testing Mn2+ concentrations, Tris-HCl pH 7.5 was used and different amounts of MnCl2 added (0.25–5 mM). The samples were incubated at 37 °C for 30 min and formamide loading buffer (90% formamide, 0.1% xylene xyanol and 0.1% bromphenol blue) was added to terminate the reactions. Samples were heated to 50 °C for 5 min before separation of substrate and product by 15% denaturating PAGE (Long Ranger, 7 M urea and 1x taurine). To map ENDOV cleavage position, samples were run on a 20% sequencing gel together with 32P-labelled RNA oligonucleotides of defined lengths (13–16 nt; Supplementary Table S1, primers 16–19). The results were visualized by phosphorimaging (Typhon 9410, Amersham Biosciences) and quantified with ImageQuant TL (Molecular Dynamics). Full size images of all panels presented are found in Supplementary Fig. S3 in the order they appear in the manuscript.
For the single-turnover assays, 5 nM of endonuclease V was incubated with 1 nM substrate in 100-μl reaction volume using the same buffer as above. Samples were withdrawn after 3, 8, 15, 30, 60, 120, 300, 900, 1,800, 3,600 and 7,200 s and reactions stopped and analyzed as described above. Quenched samples were kept on ice until all time points were collected. Results were visualized and quantified as above. For the calculation of the catalytic turnover rate kobs (s−1), a one phase association model was fitted to three parallel data sets.
Modelling
A model of human ENDOV in complex with DNA dI rG was constructed by superposing the homology model of human ENDOV21 with the T. maritima EndoV in complex with incised DNA44 (PDB id 2W35) and subsequently replacing the dG 3′ to the inosine with an rG nt retrieved from a 0.95-Å resolution X-ray structure of a double-helix RNA fragment45 (PDB id 3NJ6).
Cell culture and Northern blot analysis
Human epithelial glioblastoma cells U373-MG (American Type Culture Collection) were cultured in Dulbecco’s modified Eagle’s medium (Gibco) supplemented with 10% fetal calf serum (PAA lab), 1x GlutaMAX (200 mM, Gibco) and 1 × penicillin–streptomycin (Lonza) at 37 °C in 5% CO2 atmosphere. Total tRNAs were isolated using RNAzol RT (Molecular Research Center) according to the manufacturer’s recommendations. Gel electrophoresis showed that the isolates contained mainly tRNA. Total tRNA (4.5 nM) was incubated with ENDOV (0.3 and 0.6 μM) as described under activity assays. After heat denaturation at 55 °C for 5 min, samples were separated by 15% PAGE in 1x taurine at 200 V for 50 min. The tRNA was transferred to a nylon membrane (Hybond XL, GE Healthcare) by electroblotting in 1x taurine at 4 °C, 5 V and over night. RNA was UV-crosslinked to the membranes (120 mJ cm−2 in a CL-1000 UV-Crosslinker, UVP). The Northern Max kit (Ambion, Applied Biosystems) was used for prehybridization, hybridization and washing steps as described by the manufacturer. 32P 5′-end-labelled oligonucleotides (Eurofins; Supplementary Table S3) complementary to the 5′-ends of tRNASer(AGA), tRNALeu(AAG), tRNAArg(ACG), tRNAAsp(GTC), tRNAGlu(CTC) and tRNALys(CTT) were used as probes. Hybridization signals were analyzed by phosphorimaging and ImageQuant TL software. If endonuclease V incises the tRNA at the wobble I, the expected size of the 5′-cleavage product is 35 nt. Markers used were 32P labelled RNA oligonucleotides (25 and 38 nt), which will migrate slightly faster than unlabelled fragments due to the negative charge of the phosphate label. Hybridized probes were removed from the filters by boiling in 0.1% SDS. Full size images of the Northern blots presented are found in Supplementary Fig. S3n–s.
Additional information
How to cite this article: Vik, E. S. et al. Endonuclease V cleaves at inosines in RNA. Nat. Commun. 4:2271 doi: 10.1038/ncomms3271 (2013).
References
Gray, M. W. Evolutionary origin of RNA editing. Biochemistry 51, 5235–5242 (2012).
Keegan, L. P., Leroy, A., Sproul, D. & O'Connell, M. A. Adenosine deaminases acting on RNA (ADARs): RNA-editing enzymes. Genome Biol. 5, 209 (2004).
Bass, B. L. RNA editing by adenosine deaminases that act on RNA. Annu. Rev. Biochem. 71, 817–846 (2002).
Jin, Y., Zhang, W. & Li, Q. Origins and evolution of ADAR-mediated RNA editing. IUBMB. Life 61, 572–578 (2009).
Rosenthal, J. J. & Seeburg, P. H. A-to-I RNA editing: effects on proteins key to neural excitability. Neuron 74, 432–439 (2012).
Maas, S., Kawahara, Y., Tamburro, K. M. & Nishikura, K. A-to-I RNA editing and human disease. RNA. Biol. 3, 1–9 (2006).
Nishikura, K. Editor meets silencer: crosstalk between RNA editing and RNA interference. Nat. Rev. Mol. Cell Biol. 7, 919–931 (2006).
Orlandi, C., Barbon, A. & Barlati, S. Activity regulation of adenosine deaminases acting on RNA (ADARs). Mol. Neurobiol. 45, 61–75 (2012).
Agris, P. F., Vendeix, F. A. & Graham, W. D. tRNA's wobble decoding of the genome: 40 years of modification. J. Mol. Biol. 366, 1–13 (2007).
Su, A. A. & Randau, L. A-to-I and C-to-U editing within transfer RNAs. Biochemistry (Mosc.) 76, 932–937 (2011).
Basilio, C., Wahba, A. J., Lengyel, P., Speyer, J. F. & Ochoa, S. Synthetic polynucleotides and the amino acid code. V. Proc. Natl Acad. Sci. USA 48, 613–616 (1962).
Cantara, W. A. et al. The RNA modification database, RNAMDB: 2011 update. Nucleic Acids Res. 39, D195–D201 (2011).
Lindahl, T. Instability and decay of the primary structure of DNA. Nature 362, 709–715 (1993).
Yao, M., Hatahet, Z., Melamede, R. J. & Kow, Y. W. Purification and characterization of a novel deoxyinosine-specific enzyme, deoxyinosine 3' endonuclease, from Escherichia coli. J. Biol. Chem. 269, 16260–16268 (1994).
Guo, G., Ding, Y. & Weiss, B. nfi, the gene for endonuclease V in Escherichia coli K-12. J. Bacteriol. 179, 310–316 (1997).
Dalhus, B. et al. Structures of endonuclease V with DNA reveal initiation of deaminated adenine repair. Nat. Struct. Mol. Biol. 16, 138–143 (2009).
Gates, F. T. & Linn, S. Endonuclease from Escherichia coli that acts specifically upon duplex DNA damaged by ultraviolet light, osmium tetroxide, acid, or x-rays. J. Biol. Chem. 252, 2802–2807 (1977).
Hardeland, U., Bentele, M., Jiricny, J. & Schar, P. The versatile thymine DNA-glycosylase: a comparative characterization of the human, Drosophila and fission yeast orthologs. Nucleic Acids Res. 31, 2261–2271 (2003).
Saparbaev, M., Mani, J. C. & Laval, J. Interactions of the human, rat, Saccharomyces cerevisiae and Escherichia coli 3-methyladenine-DNA glycosylases with DNA containing dIMP residues. Nucleic Acids Res. 28, 1332–1339 (2000).
Feng, H., Klutz, A. M. & Cao, W. Active site plasticity of endonuclease V from Salmonella Typhimurium. Biochemistry 44, 675–683 (2005).
Fladeby, C. et al. The human homolog of Escherichia coli endonuclease V is a nucleolar protein with affinity for branched DNA structures. PLoS ONE 7, e47466 (2012).
Huang, J., Lu, J., Barany, F. & Cao, W. Multiple cleavage activities of endonuclease V from Thermotoga Maritima: recognition and strand nicking mechanism. Biochemistry 40, 8738–8748 (2001).
Kanugula, S., Pauly, G. T., Moschel, R. C. & Pegg, A. E. A bifunctional DNA repair protein from Ferroplasma acidarmanus exhibits O6-alkylguanine-DNA alkyltransferase and endonuclease V activities. Proc. Natl Acad. Sci. USA 102, 3617–3622 (2005).
Liu, J., He, B., Qing, H. & Kow, Y. W. A deoxyinosine specific endonuclease from hyperthermophile, Archaeoglobus fulgidus: a homolog of Escherichia coli endonuclease V. Mutat. Res. 461, 169–177 (2000).
Mi, R., ford-Zappala, M., Kow, Y. W., Cunningham, R. P. & Cao, W. Human endonuclease V as a repair enzyme for DNA deamination. Mutat. Res. 735, 12–18 (2012).
Moe, A. et al. Incision at hypoxanthine residues in DNA by a mammalian homologue of the Escherichia coli antimutator enzyme endonuclease V. Nucleic Acids Res. 31, 3893–3900 (2003).
Onoa, B. & Tinoco, I. Jr RNA folding and unfolding. Curr. Opin. Struct. Biol. 14, 374–379 (2004).
Vendeix, F. A., Munoz, A. M. & Agris, P. F. Free energy calculation of modified base-pair formation in explicit solvent: a predictive model. RNA. 15, 2278–2287 (2009).
Cerritelli, S. M. & Crouch, R. J. Ribonuclease H: the enzymes in eukaryotes. FEBS J. 276, 1494–1505 (2009).
Shaw, N. N. & Arya, D. P. Recognition of the unique structure of DNA:RNA hybrids. Biochimie 90, 1026–1039 (2008).
Noy, A., Perez, A., Marquez, M., Luque, F. J. & Orozco, M. Structure, recognition properties, and flexibility of the DNA:RNA hybrid. J. Am. Chem. Soc. 127, 4910–4920 (2005).
Daniel, C., Veno, M. T., Ekdahl, Y., Kjems, J. & Ohman, M. A distant cis acting intronic element induces site-selective RNA editing. Nucleic Acids Res. 40, 9876–9886 (2012).
Grosjean, H., de Crecy-Lagard, V. & Marck, C. Deciphering synonymous codons in the three domains of life: co-evolution with specific tRNA modification enzymes. FEBS Lett. 584, 252–264 (2010).
Thompson, D. M. & Parker, R. Stressing out over tRNA cleavage. Cell 138, 215–219 (2009).
Scadden, A. D. The RISC subunit Tudor-SN binds to hyper-edited double-stranded RNA and promotes its cleavage. Nat. Struct. Mol. Biol. 12, 489–496 (2005).
Yang, W. et al. Modulation of microRNA processing and expression through RNA editing by ADAR deaminases. Nat. Struct. Mol. Biol. 13, 13–21 (2006).
Phizicky, E. M. & Hopper, A. K. tRNA biology charges to the front. Genes Dev. 24, 1832–1860 (2010).
Lu, J., Huang, B., Esberg, A., Johansson, M. J. & Bystrom, A. S. The Kluyveromyces lactis gamma-toxin targets tRNA anticodons. RNA 11, 1648–1654 (2005).
Aas, P. A. et al. Human and bacterial oxidative demethylases repair alkylation damage in both RNA and DNA. Nature 421, 859–863 (2003).
Berquist, B. R., McNeill, D. R. & Wilson, D. M. III Characterization of abasic endonuclease activity of human Ape1 on alternative substrates, as well as effects of ATP and sequence context on AP site incision. J. Mol. Biol. 379, 17–27 (2008).
Jobert, L. et al. The human base excision repair enzyme SMUG1 directly interacts with DKC1 and contributes to RNA quality control. Mol. Cell 49, 339–345 (2013).
Vascotto, C. et al. APE1/Ref-1 interacts with NPM1 within nucleoli and plays a role in the rRNA quality control process. Mol. Cell Biol. 29, 1834–1854 (2009).
Virgen-Slane, R. et al. An RNA virus hijacks an incognito function of a DNA repair enzyme. Proc. Natl Acad. Sci. USA 109, 14634–14639 (2012).
Dalhus, B. et al. Structural insight into repair of alkylated DNA by a new superfamily of DNA glycosylases comprising HEAT-like repeats. Nucleic Acids Res. 35, 2451–2459 (2007).
Kiliszek, A., Kierzek, R., Krzyzosiak, W. J. & Rypniewski, W. Atomic resolution structure of CAG RNA repeats: structural insights and implications for the trinucleotide repeat expansion diseases. Nucleic Acids Res. 38, 8370–8376 (2010).
Acknowledgements
We acknowledge financial support from the Norwegian Research Council and the Norwegian Cancer Society. E.S.V., B.D. and I.A. have additionally received support from the MLSUiO program for Molecular Life Science research at the University of Oslo. M.B. and B.D. have received support from the South-Eastern Norway Regional Health Authority (grants no. 2009100, 2011040 and 2012085) for establishing the Regional Core Facility for Structural Biology and Bioinformatics. We thank Rune J. Forstrøm for technical assistance and Alexander Rowe for editing the manuscript.
Author information
Authors and Affiliations
Contributions
B.D., M.B. and I.A. conceived and planned the study. P.S.A. purified proteins, E.S.V. and M.S.N. performed the activity assays, B.D. made the 3D homology model. E.S.V., B.D., M.B., C.F. and I.A. wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1-S3 and Supplementary Tables S1-S3 (PDF 1359 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Sebastian Vik, E., Sameen Nawaz, M., Strøm Andersen, P. et al. Endonuclease V cleaves at inosines in RNA. Nat Commun 4, 2271 (2013). https://doi.org/10.1038/ncomms3271
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms3271
This article is cited by
-
RNA Editing Therapeutics: Advances, Challenges and Perspectives on Combating Heart Disease
Cardiovascular Drugs and Therapy (2023)
-
Crystal structure of the yeast heterodimeric ADAT2/3 deaminase
BMC Biology (2020)
-
In vivo diversification of target genomic sites using processive base deaminase fusions blocked by dCas9
Nature Communications (2020)
-
Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage
Nature (2017)
-
Deoxyinosine triphosphate induces MLH1/PMS2- and p53-dependent cell growth arrest and DNA instability in mammalian cells
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.