Abstract
Avirulence (Avr) genes of plant pathogens encode effector proteins that trigger immunity in plants carrying appropriate resistance (R) genes. The Phytophthora sojae Avr3a gene displays allelic variation in messenger RNA transcript levels. P. sojae strains with detectable Avr3a gene transcripts are avirulent on plants carrying the R-gene Rps3a, whereas strains lacking Avr3a mRNA escape detection by Rps3a and are virulent. Here we show non-Mendelian interactions between naturally occurring Avr3a alleles that result in transgenerational gene silencing, and we identify small RNA molecules of 25 nucleotides that are abundant in gene-silenced strains but not in strains with Avr3a mRNA. This example of transgenerational gene silencing is exceptional because it is naturally occurring and results in gain of virulence in a pathogenic organism.
Similar content being viewed by others
Introduction
Plant pathogens secrete effector proteins to enable disease, but evolution favors host plant immune systems to recognize these factors as cues for activating rapid defence responses that arrest the infection1. Effector-triggered immunity in plants is the consequence of signalling events that are determined by resistance (R) gene products interacting, directly or indirectly, with pathogen avirulence (Avr) effectors.
Phytophthora sojae causes stem and root rot of soybean and is a major disease problem that plagues this crop2. The P. sojae Avr3a gene encodes a predicted protein of 111 amino acids that includes a signal peptide, a host-targeting motif and a carboxy-terminal effector domain, which are common features of oomycete Avr effectors3,4,5,6. Avr3a resides in a 10.8 kilobase (kb) DNA segment that displays copy number variation among P. sojae strains5,6. Four haplotypes of Avr3a are known and gain of virulence on the R-gene Rps3a is caused by messenger RNA transcript differences between strains. In crosses between P. sojae strains P6497 (Avr3aP6497/Avr3aP6497) and P7064 (Avr3aP7064/Avr3aP7064), avirulence towards Rps3a plants segregates as a dominant Mendelian trait conditioned by the presence of Avr3aP6497. Transcripts of Avr3aP6497 but not Avr3aP7064 mRNA are detectable in each of the parental strains and their progeny5,6.
Sequencing of Avr3a alleles and analysis of mRNA transcripts in 17 different P. sojae strains revealed four different haplotypes6, but the inheritance behaviour of each of these haplotypes in segregating populations has not been investigated. The present study was intended to test the inheritance Avr3a in P. sojae strains with haplotypes that are different from strains P6497 and P7064. To our surprise, we discovered that genetic outcrosses between P. sojae strains ACR10 and P7076 resulted in transgenerational gene silencing of Avr3a and gain of virulence on soybean plants carrying the Rps3a gene.
Results
Transgenerational gene silencing of Avr3a
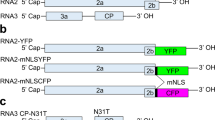
To determine the inheritance of other Avr3a alleles, we created crosses between P. sojae strains P7076 (Avr3aP7076/Avr3aP7076) and ACR10 (Avr3aACR10/Avr3aACR10). The strain P7076 is avirulent, whereas strain ACR10 is virulent on Rps3a plants, and mRNA transcripts are detectable for Avr3aP7076 but not Avr3aACR10, as we have previously shown5,6 (Fig. 1a). To select hybrid progeny and follow segregation of Avr3a alleles we used co-dominant DNA markers derived from cleaved amplified polymorphic (CAP) fragments, because Avr3aP7076 and Avr3aACR10 differ in sequence5,6. A total of 28 F1 hybrid (Avr3aP7076/Avr3aACR10) progeny were isolated, scored for virulence on Rps3a plants and assessed for the presence of Avr3a mRNA transcripts by reverse transcriptase polymerase chain reaction (RT–PCR) (Table 1, Fig. 2, Supplementary Fig. S1). All 28 F1 progeny were virulent on Rps3a plants and were lacking Avr3a mRNA transcripts. This result was unexpected because heretofore avirulent alleles of Avr3a that accumulate mRNA transcripts were dominant to virulent alleles that lack transcripts, such as in crosses between P. sojae strains P6497 and P7064. To further explore the segregation of Avr3a alleles, virulence and mRNA transcript accumulation in this cross, we generated three different F2 populations from separate F1 individuals. In all, 139 F2 progeny were tested for segregation of Avr3a alleles and mRNA transcripts and scored for virulence on Rps3a plants. The Avr3aP7076 and Avr3aACR10 alleles segregated normally in each F2 population but remarkably all 139 F2 progeny lacked Avr3a mRNA transcripts and were virulent on Rps3a plants (Table 1). A sampling of eight F3 individuals, derived from homozygous Avr3aP7076/Avr3aP7076 F2 cultures, indicates that gene silencing is maintained and meiotically stable in these inbred lines (Supplementary Fig. S2). These results demonstrate heritable transgenerational gene silencing of Avr3a in crosses between P. sojae strains ACR10 and P7076.
(a) A schematic illustration of Avr3a alleles from four different strains of P. sojae. The segment of DNA harbouring Avr3a and four other predicted genes is shown. Predicted open reading frames are indicated by bold arrows. Shown for each allele is the copy number of the DNA segment, presence (+) or absence (−) of Avr3a mRNA transcripts and disease outcome on Rps3a plants as virulent (V) or avirulent (A). (b) Profiles of sRNA from P. sojae strains ACR10 and P7076, and from a pool of F2 cultures selected for their Avr3aP7076/Avr3aP7076 genotype. Average depth of coverage across the 10.8-kb genome segment containing Avr3a was normalized to the total number of trimmed reads for each sample. (c) Size of sRNA aligning to the 10.8-kb DNA segment. The total number of counts of RNA sequence reads and the size in nucleotides (nt) is shown for P. sojae strain ACR10.
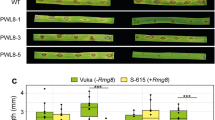
Virulence assays of P. sojae cultures were performed on greenhouse-grown seedlings of soybean cultivar Williams (rps3a) and the isoline L83-570 (Rps3a) by hypocotyl inoculation with P. sojae mycelia. Shown are disease outcomes 6 days after inoculation of plants with the parental strains, the F1 hybrids used to construct each of the three F2 populations, or with representative F2 individuals illustrating each Avr3a genotype from the three F2 populations tested. Pots are 10 cm in diameter.
Distinctive profile of small RNA (sRNA) in silenced strains
To explore the cause of transgenerational silencing of Avr3a, we performed deep sequencing of sRNA from each parental P. sojae strain ACR10 and P7076, and from a sample of F2 cultures with the genotype Avr3aP7076/Avr3aP7076. Results from this analysis show that the profile of sRNA coverage across the Avr3a region differs between ACR10 and P7076 (Fig. 1b). The sRNA sequences matching to the Avr3a region are far more abundant in strain ACR10 compared with P7076 (Table 2). The most prevalent of these sRNA molecules are 24–26 nucleotides in length (Fig. 1c). In strain ACR10, there is a nearly continuous coverage of sRNA sequences that align to a 3.7 kb span of the Avr3a interval including the 5′ and 3′ intergenic segments but not extending into flanking genes. In contrast, in strain P7076 coverage of matching sRNA is limited to a 0.3-kb segment occurring in the 5′ intergenic region. Sequencing of sRNA from homozygous Avr3aP7076/Avr3aP7076 F2 cultures resulted in a profile of coverage that is nearly identical to that from strain ACR10 (Avr3aACR10/Avr3aACR10) rather than that from P7076 (Avr3aP7076/Avr3aP7076). The sRNA profiles were replicated from independent samples of ACR10, P7076, F1 and F2 cultures, and extended to include samples from additional P. sojae strains P6497 and ACR16 (Table 2; Supplementary Fig. S3). The results consistently show a pattern of extensive sRNA coverage of Avr3a occurs in gene-silenced strains.
Discussion
Previous studies have shown that transformation of organisms, including species of Phytophthora, with recombinant DNA or RNA molecules can cause internuclear or trangenerational gene silencing that resembles our observations of naturally occurring silencing of Avr3a (refs 7,8). Moreover, results from comparative genome sequencing of species within the Phytophthora infestans clade suggest a role for epigenetic mechanisms in the rapid evolution of Phytophthora species9. Thus, we propose that transgenerational gene silencing of Avr3a in P. sojae is the result of epigenetic changes that modulate the expression state of this gene, and that are transmitted to progeny in a non-Mendelian fashion.
Given the emerging functions of sRNAs in controlling epigenetic marks and gene expression states in numerous experimental systems, it is noteworthy that we found an association between sRNAs and gene silencing at the Avr3a locus. In plants, sRNA of 24–26 nucleotides are involved in directing chromatin changes including DNA methylation, whereas shorter ones of 21–22 nucleotides participate in mRNA degradation10. Heterochromatic silencing in yeast is initiated by an interaction of sRNA and RNA polymerase II, which guides histone modifications that control chromatin structure and gene expression11. Thus, the evidence suggests a role for sRNA in directing the transgenerational gene expression changes of the P. sojae Avr3a gene.
Our observations of transgenerational gene silencing of Avr3a provide a remarkable example of pathogen adaptation to host immune surveillance. The interactions between pathogens and their hosts are often interpreted as an evolutionary race, where pathogen virulence and host resistance are at the forefront12. Transgenerational gene silencing of an Avr effector locus represents a novel method of host immune evasion that offers the ability to spread quickly in pathogen populations, but yet is potentially reversible as the sequence of the Avr gene itself remains unchanged. We predict that other eukaryotic pathogens of plants, humans and animals will employ similar mechanisms to escape detection by host immune systems.
Methods
Phytophthora sojae strains and crosses
The origin of P. sojae strains and methods of culture, propagation and oospore isolation have been described5,6,13,14,15. P. sojae is an oomycete and diploid microorganism that reproduces sexually through the development of oogonia (female gametophyte) and antheridia (male gametophyte). P. sojae is homothallic and is not known to have mating types, therefore oospores can develop from self-fertilization or from out-crossing when different strains of the organism are grown together2. To perform sexual crosses of P. sojae, parental strains are co-cultivated and oospores are isolated from culture homogenates by a series of purification steps13,14,15. Purified oospores are diluted and plated out on water agar to germinate. Individual germinating oospores are picked under a dissecting microscope and transferred to rich growth medium. Samples of purified DNA from the resulting cultures are tested using co-dominant DNA markers polymorphic between the parents, to determine whether the individuals result from self-fertilization or from out-crossing events between the parental strains. Such markers designed to distinguish Avr3aACR10 and Avr3aP7076 alleles (described below) were used to identify F1 progeny in crosses between strain ACR10 and P7076. The F2 populations were generated by isolating oospores from self-fertilized F1 individuals.
Purification of nucleic acids and scoring of markers and transcripts for Avr3a
Mycelia cultures of P. sojae for nucleic acid isolation were grown on vegetable juice (V8) agar media overlaid with cellophane (BioRad). After growth in darkness for 7 days at 25 °C, mycelia-grown cellophane disks were peeled off the media, frozen at −80 °C and ground to a fine powder in liquid N2 using a mortar and pestle. Genomic DNA was purified using conventional phenol–chloroform extraction procedures followed by precipitation with isopropanol16. Purfied DNA pellets were washed with 70% (v/v) ethanol, dried and resuspended in a solution of 10 mM Tris HCl, pH 7.4, 0.1 mM EDTA. CAP markers were used to track segregation of the Avr3aACR10 and Avr3aP7076 alleles. For CAPs analysis, PCR amplification was performed with 5 ng of genomic DNA as template in a volume of 25 μl under the following conditions: 0.2 mM dNTPs, 1.5 mM MgCl2, 0.4 μM of each primer (forward primer 5′-GCTGCTTCCTTCCTGGTTGC-3′ and reverse primer 5′-GCTGCTGCCTTTTGCTTCTC-3′), 1.25 U Taq DNA polymerase and 1 × PCR buffer as recommended by the supplier (Invitrogen, Life Technologies). The amplification was carried out in a 96-well plate (GeneAmp PCR System 9700, Applied Biosystems) under the following conditions: 94 °C for 2 min; 40 cycles of 94 °C for 40 s, 58 °C for 40 s, 72 °C for 2 min; followed by an extension step of 10 min at 72 °C. For restriction analysis of the PCR fragments, 10 μl of the amplified products were incubated overnight at 37 °C in a volume of 20 μl using 3 U of AluI (New England Biolabs). Digestion products were resolved by agarose gel electrophoresis and visualized by ethidium bromide staining according to standard methods16. The Avr3aACR10 sequence includes an AluI site, whereas the Avr3aP7076 sequence lacks the restriction site and is not digested5,6.
Total RNA was extracted from frozen, pulverized mycelia samples in a solution of phenol-guanidine isothiocyanate (TRIzol, Invitrogen) according to instructions provided by the manufacturer. Approximately 1 μg total RNA was subject to DNaseI (Invitrogen) treatment before RT–PCR, as previously described5,6. Cycling conditions for detecting transcripts of Avr3a as well as for the P. sojae actin gene (forward primer 5′-CGAAATTGTGCGCGACATCAAG-3′ and reverse primer 5′-GGTACCGCCCGACAGCACGAT-3′), used as a control, were described as above with the exception that 25 cycles were used for transcript analysis.
Plant inoculation and virulence assays
Virulence assays of P. sojae cultures were performed on greenhouse-grown seedlings of soybean cultivar Williams (rps) and the isoline L83-570 (Rps3a), according to standard methods5,6. Soybean seeds (20–30 seeds per pot) were sown in 10 cm pots and grown for 7 days before disease testing. For inoculation, P. sojae cultures were grown for 5–7 days on 0.9% (v/v) V8 agar plates. Inoculum was prepared by excising 1 cm segments of agar cut from the growing edge of mycelia colonies and macerating the cultures through a 3-ml syringe attached to an 18-gauge needle. Soybean plants were inoculated in the mid-section of each hypocotyl by making a small incision for injection of the mycelial slurry. Inoculated plants were covered with plastic bags to maintain humidity for 2 days. Disease symptoms were allowed to develop for an additional 4 days before phenotypes were scored. A minimum of three independent replicates of the disease assay were performed for each P. sojae culture tested.
Deep sequencing of P. sojae sRNA
To profile sRNA molecules of P. sojae cultures, samples of RNA were purified from mycelia cultures as described above. The size selection of sRNA, attachment of adaptor molecules and library construction was performed using prepared supplies and reagents (Illumina small RNA v1.5), following instructions provided by the manufacturer. Sequencing was performed on a flow cell instrument (Illumina GAII). After trimming to remove adaptor sequences and low-quality sequences, the depth of sRNA coverage of the Avr3a region was determined by alignment, and normalized to the number of reads per library. Average depth of coverage across the 10.8-kb genome segment containing Avr3a was determined using a 50-base pair moving interval and values normalized to the total number of trimmed reads for each sample. The number of trimmed reads obtained for each of the sequenced sRNA libraries is proved in Table 2.
Additional information
How to cite this article: Qutob, D. et al. Transgenerational gene silencing causes gain of virulence in a plant pathogen. Nat. Commun. 4:1349 doi: 10.1038/ncomms2354 (2013).
References
Jones J. D. & Dangl J. L. The plant immune system. Nature 444, 323–329 (2006).
Erwin D. C. & Ribeiro O. K. Phytophthora Diseases Worldwide (APS Press, 1997).
Tyler B. M. Entering and breaking: virulence effector proteins of oomycete plant pathogens. Cell Microbiol. 11, 13–20 (2009).
Oliva R. et al. Recent developments in effector biology of filamentous plant pathogens. Cell Microbiol. 12, 705–715 (2010).
Qutob D. et al. Copy number variation and transcriptional polymorphisms of Phytophthora sojae RXLR effector genes Avr1a and Avr3a. PLoS ONE e5066 (2009).
Dong S. et al. Sequence variants of the Phytophthora sojae RXLR effector Avr3a/5 are differentially recognized by Rps3a and Rps5 in soybean. PLoS ONE e20172 (2011).
van West P., Kamoun S., van’t Klooster J. W. & Govers F. Internuclear gene silencing in Phytophthora infestans. Mol. Cell. 3, 339–348 (1999).
van West P. et al. Internuclear gene silencing in Phytophthora infestans is established through chromatin remodelling. Microbiology 154, 1482–1490 (2008).
Raffaele S. et al. Genome evolution following host jumps in the Irish potato famine pathogen lineage. Science 330, 1540–1543 (2010).
Molnar A., Melnyk C. & Baulcombe D. C. Silencing signals in plants: a long journey for small RNAs. Genome Biol. 12, 215 (2011).
Zaratiegui M. et al. RNAi promotes heterochromatic silencing through replication-coupled release of RNA Pol II. Nature 479, 135–137 (2011).
Woolhouse M. E., Webster J. P., Domingo E., Charlesworth B. & Levin B. R. Biological and biomedical implications of the co-evolution of pathogens and their hosts. Nat. Genet. 32, 569–577 (2002).
MacGregor T. et al. Genetic and physical mapping of Avr1a in Phytophthora sojae. Genetics 160, 949–959 (2002).
Tyler B. M., Forster H. & Coffey M. D. Inheritance of avirulence factors and restriction fragment length polymorphism markers in outcrosses of the oomycete Phytophthora sojae. Mol. Plant-Microbe Interact. 8, 515–523 (1995).
Gijzen M. & Qutob D. in Oomycete Genetics and Genomics: Diversity, Interactions and Research Tools eds Lamour K., Kamoun S. 303–329 (Wiley-Blackwell, 2009).
Sambrook J., Fritsch E. F. & Maniatis T. Molecular Cloning: A Laboratory Manual Cold Spring Harbor Press (1989).
Acknowledgements
We thank The Centre for Applied Genomics (Toronto) and the BC Cancer Agency (Vancouver) for sRNA sequencing; Aldona Gaidauskas-Scott, Irina Mutiu, Kuflom Kuflu and Alex Molnar for contributions to the work; and Brett Tyler and Krzysztof Szczyglowski for comments on the manuscript. This work was funded by the Agriculture and Agri-Food Canada Crop Genomics Initiative.
Author information
Authors and Affiliations
Contributions
D.Q. and M.G. conceived and designed the experiments. D.Q. performed the experiments. D.Q., B.P.C. and M.G. analysed the data. M.G. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1-S3 (PDF 1069 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Qutob, D., Patrick Chapman, B. & Gijzen, M. Transgenerational gene silencing causes gain of virulence in a plant pathogen. Nat Commun 4, 1349 (2013). https://doi.org/10.1038/ncomms2354
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms2354
This article is cited by
-
A global-temporal analysis on Phytophthora sojae resistance-gene efficacy
Nature Communications (2023)
-
Evasion of plant immunity by microbial pathogens
Nature Reviews Microbiology (2022)
-
Epigenetics of wheat–rust interaction: an update
Planta (2022)
-
Small RNAs generated by bidirectional transcription mediate silencing of RXLR effector genes in the oomycete Phytophthora sojae
Phytopathology Research (2019)
-
Small RNA discovery in the interaction between barley and the powdery mildew pathogen
BMC Genomics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.