Abstract
The 11-cis retinal chromophore is tightly packed within the interior of the visual receptor rhodopsin and isomerizes to the all-trans configuration following absorption of light. The mechanism by which this isomerization event drives the outward rotation of transmembrane helix H6, a hallmark of activated G protein-coupled receptors, is not well established. To address this question, we use solid-state NMR and FTIR spectroscopy to define the orientation and interactions of the retinal chromophore in the active metarhodopsin II intermediate. Here we show that isomerization of the 11-cis retinal chromophore generates strong steric interactions between its β-ionone ring and transmembrane helices H5 and H6, while deprotonation of its protonated Schiff’s base triggers the rearrangement of the hydrogen-bonding network involving residues on H6 and within the second extracellular loop. We integrate these observations with previous structural and functional studies to propose a two-stage mechanism for rhodopsin activation.
Similar content being viewed by others
Introduction
The visual receptor rhodopsin is a member of the family A G protein-coupled receptors (GPCRs)1,2. These receptors share a seven transmembrane (TM) helix architecture and the ability to activate heterotrimeric G proteins, yet they respond to a wide array of ligands ranging from small-molecule odorants in the olfactory receptors to peptide ligands in the hormone and chemokine receptors1,2. The diversity between receptor subfamilies largely lies in the extracellular, ligand-binding region, which has evolved to recognize and respond to different types of signals3,4. The extracellular loops and extracellular ends of the TM helices contain many subfamily-specific residues, while most of the sites with high sequence conservation across the family A GPCRs are found in the TM core and intracellular G protein-binding cavity1,5. A common feature of GPCR activation is the outward rotation of the intracellular end of TM helix H6, which serves to expose the G protein-binding site6,7,8. Nevertheless, the molecular mechanism by which this intracellular motion is achieved on extracellular binding of such diverse ligands remains largely unresolved.
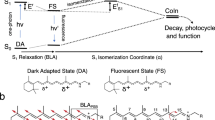
Rhodopsin provides an ideal model system for addressing the activation mechanism of family A GPCRs. Its light-sensitive retinal chromophore is covalently bound via a protonated Schiff’s base (PSB) linkage to Lys2967.43 (superscripts denote generic Ballesteros–Weinstein numbering of GPCRs9) in the interior of the protein. This covalent attachment ensures full ligand occupancy, a desirable property for structural studies. The retinal-binding site has evolved to accommodate the 11-cis isomer of the retinal PSB10, which acts as an inverse agonist and locks the receptor in a fully off-state. Photochemical isomerization and deprotonation of the PSB forms an all-trans retinal SB chromophore, which acts as a full agonist for receptor activation. That is, rhodopsin functions as a molecular off–on switch; it is designed to be fully inactive in the dark and to rapidly convert to a fully active structure in the light. This activation process differs from GPCRs that bind diffusible agonists where the signalling status is more complex and governed by an equilibrium between ligand-bound and ligand-free states11.
The crystal structure of the apoprotein opsin6,7 and constitutively active mutants of rhodopsin12,13 have provided several key insights into the conformational changes that occur on receptor activation. These structures confirm the large outward rotation of H6, the signature of an active receptor14. However, despite the large change in retinal configuration and orientation, the active-state crystal structures of rhodopsin show almost no change in structure on the extracellular side of the receptor when compared with the large changes observed on the intracellular side15. This observation is surprising as a substantial amount of absorbed light energy (∼35 kcal mol−1) is stored within retinal–protein interactions in the primary photoproduct bathorhodopsin16 and then released as the retinal and surrounding protein relax during the transition to the active metarhodopsin II (Meta-II) intermediate2. The lack of structural changes in the binding site surrounding the retinal raises the question of how retinal isomerization and PSB deprotonation generate the large helix rearrangements on the intracellular side to create the intracellular G protein-binding pocket.
We take advantage of solid-state nuclear magnetic resonance (NMR) and Fourier transform infrared (FTIR) spectroscopy of Meta-II to address how protein residues within the 11-cis retinal-binding site adapt to the constrained all-trans retinal configuration following retinal isomerization. We also gain insight into how hydrogen-bonding networks on the extracellular surface of rhodopsin rearrange in response to PSB deprotonation. Low temperature (below ∼0 °C) slows the thermal steps in the rhodopsin photoreaction and provides a way to trap the native light-activated Meta-II state17,18,19. In contrast, high-resolution crystal structures, which capture elements of the active state, rely on soaking crystals of the apoprotein opsin with all trans-retinal20 or have required stabilizing and/or activating mutations12,13. Crystals of native rhodopsin crack and no longer diffract following absorption of light and conversion to an active conformation21. Structural information from solid-state NMR experiments comes from distance measurements between specific 13C labels incorporated into the retinal and the protein, as well as from chemical shift measurements to assess changes in hydrogen bonding. Nevertheless, this approach relies on the crystal structures as a reference to establish how specific sites change position or orientation on activation. FTIR difference spectroscopy provides a complementary approach to NMR for assessing how specific residues in close association with the retinal control the equilibrium between Meta-I and Meta-II, the final step in the reaction pathway22,23,24.
The first major observation to emerge from these studies is that the orientation of the retinal differs in Meta-II trapped along its reaction coordinate compared with Meta-II stabilized for crystallography. NMR constraints indicate that the β-ionone ring does not change orientation on activation, but rather the largest change in retinal structure involves rotation of the C20 methyl group toward extracellular loop-2 (EL2)18. In contrast, crystal structures obtained by diffusing all-trans retinal into opsin (Meta-II-opsin20) or using the constitutively active M257Y mutant25 (Meta-II-M257Y; ref. 13) suggest that the all-trans retinal SB is in an orientation roughly opposite to that observed by NMR (Supplementary Fig. 1). The second major observation is that there are larger changes in the position of key aromatic residues (Tyr191EL2 and Tyr2686.51) surrounding the retinal than observed in the crystal structures. These residues are highly conserved in the visual receptor subfamily. FTIR studies of the Y191F and Y268F mutants indicate that these tyrosines control the Meta-I ⇔ Meta-II equilibrium. We propose an activation mechanism that builds on these observations and previous studies by emphasizing the changes in extracellular residues in close proximity to the retinal. The mechanism that emerges from these studies implicates steric interactions as the dominant force driving structural changes in the light-triggered transition of rhodopsin to Meta-I, while electrostatic (and hydrogen bonding) interactions control the formation of the active Meta-II conformation.
Results
Retinal orientation in Meta-II
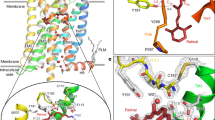
To establish the orientation of the retinal in the binding site of the active Meta-II intermediate, we focus on the positions of the retinal C18, C19 and C20 methyl groups relative to surrounding protein residues (Fig. 1 and Supplementary Fig. 1). The C19 and C20 methyl groups extend from the retinal polyene chain, while the C18 methyl group extends from the β-ionone ring. In the crystal structures of Meta-II obtained using the M257Y mutant13 or with opsin containing the bound retinal SB20, the β-ionone ring has rotated relative to its position in rhodopsin, with the C18 methyl group facing the extracellular surface. In these models, the closest tyrosine residue to the C18 methyl group is Tyr191EL2 (Supplementary Note 1 and Supplementary Tables 1–5). Below, we use NMR to measure a few specific distances between the retinal and surrounding protein to define the orientation of the retinal in Meta II. These constraints along with the results from previous biophysical and biochemical studies are used to suggest a general mechanism of receptor activation.
(a) Structure of the 11-cis retinal PSB and all-trans retinal SB in rhodopsin and Meta-II, respectively. (b) Crystal structure of rhodopsin (PDB-ID 1U19; ref. 26) showing interactions of the C18, C19 and C20 retinal methyl groups with surrounding residues. (c) Retinal C18 contacts with phenylalanine and tyrosine in rhodopsin and Meta-II. Rows through the 13C18 diagonal resonance are shown from 13C DARR NMR experiments of rhodopsin (black) and Meta-II (red) obtained with the receptor regenerated with 13C5, 13C18 retinal and incorporating 13C-ring-labelled phenylalanine and 13Cζ-labelled tyrosine. Since the chemical shifts can change between rhodopsin and Meta-II, the rows selected correspond to the diagonal chemical shifts of the 13C18 resonance in rhodopsin at 21.6 p.p.m. and Meta-II at 20.9 p.p.m. These rows maximize the crosspeak intensities. (d) Retinal C19 contacts with tyrosine. Rows through the 13C19 diagonal resonances in rhodopsin (black) at 14.7 p.p.m. and Meta-II (red) at 13.8 p.p.m. obtained with the receptor regenerated with 13C8, 13C19 retinal and incorporating 13C-ring Phe and 13Cζ-labelled Tyr. (e,f) Retinal C12, C20 contacts with tyrosine in rhodopsin and Meta-II. Rows through the 13C20 diagonal resonances in rhodopsin (black) at 16.4 p.p.m. and Meta-II (red) at 13.7 p.p.m. obtained with the receptor regenerated with 13C12, 13C20 retinal and incorporating 13Cζ-labelled Tyr. (g) Build-up curves for the observed retinal–tyrosine crosspeaks in rhodopsin (black) and Meta-II (red) scaled to the C12-C20 crosspeaks. The C12-C20 retinal distance is fixed at ∼2.4 Å. The retinal C12 and C20 distances to the Cζ carbon of Tyr2686.51 range from ∼4 to 6 Å (Supplementary Table 3). The retinal-Tyr2686.51 data are compared with build-up curves derived from model compounds and fixed distances in rhodopsin. The error in the measurements is ±0.3 Å on the basis of the signal-to-noise of the NMR spectra. (h) 13Cζ-Tyr2686.51 crosspeaks with the retinal 13C10,11 resonances (top) and the 13C14,15 resonances in rhodopsin (black) and Meta-II (red). Rows are taken through the 13Cζ-Tyr diagonal resonance at 155.2 p.p.m. in rhodopsin and 156.1 p.p.m. in Meta-II.
NMR distance measurements between 13C sites in the retinal and surrounding protein are made using dipolar-assisted rotational resonance (DARR) of rhodopsin and Meta-II (Supplementary Fig. 2). To address the orientation of the retinal β-ionone ring, DARR NMR measurements were made using rhodopsin containing 13C-ring-labelled phenylalanine and 13Cζ-labelled tyrosine, and regenerated with 13C5, 13C18-labelled retinal. The diagonal 13C resonances in two-dimensional (2D) DARR NMR spectra arise from 13C-labelled amino acids and retinal. Crosspeaks appear between resonances when the distance between the 13C atoms is <6–6.5 Å.
Rows taken from 2D NMR spectra of rhodopsin (Fig. 1c, top panel, black) and Meta-II (Fig. 1c, bottom panel, red) correspond to the regions containing crosspeaks between 13C resonances. In rhodopsin, we observe an intense crosspeak between the two directly bonded retinal carbons, 13C5 at 131.0 p.p.m. and 13C18 at 21.6 p.p.m. There are also crosspeaks at ∼129 and 137 p.p.m. that correspond to the chemical shifts of the aromatic ring carbons of 13C-labelled phenylalanine. Phe2616.44 is the only phenylalanine within the upper distance limit (∼6.5 Å) of the DARR measurements (Supplementary Tables 1–5). The intensities of the crosspeaks are consistent with the short internuclear distances (4.9–5.4 Å) between the retinal C18 methyl and the ring carbons of Phe2616.44 observed in the crystal structure of rhodopsin (PDB-ID 1U19)26.
On conversion to Meta-II, the retinal 13C resonances change frequency due to changes in retinal structure, protonation state and retinal–protein interactions. For example, the crosspeak between the 13C5 and 13C18 resonances shifts from 131 to 126 p.p.m. (Fig. 1c), but the crosspeak intensity remains the same because of the fixed distance between these two carbons. The intra-retinal C5-C18 crosspeak provides an internal control for calibrating the intensity changes observed in protein–retinal crosspeaks. The intra-retinal 13C8-13C19 and 13C12-13C20 crosspeaks provided similar internal controls in Fig. 1d,e. The intensity of the crosspeaks between the retinal and Phe2616.44 ring-13C resonances remains approximately the same as in rhodopsin, indicating that there is little change in the distance between Phe2616.44 and the 13C18 methyl group. This observation argues that the β-ionone ring does not flip as suggested by the Meta-II crystal structures. The position of the β-ionone ring is additionally constrained by contacts between the 13C5, 13C18 and the 13C16, 13C17 retinal resonances and residues (Met2075.42 and His2115.46) on H5 (Supplementary Fig. 3 and Supplementary Note 2). Furthermore, in the Meta-II-opsin and Meta-II-M257Y crystal structures, the rotation of the β-ionone ring is predicted to bring the C18 methyl group to within the DARR distance limit (∼6 Å) of 13Cζ-Tyr191EL2, which is not observed in the present studies (Fig. 1c).
The C19 methyl group provides another probe of the retinal orientation. In rhodopsin, the C19 methyl is oriented towards EL2 and packed between Thr1183.33, Tyr191EL2, Ile189EL2 and Tyr2686.51 (Fig. 1b). We observe two C19-13Cζ-tyrosine crosspeaks at 155.2 and 156.6 p.p.m. (C19-Tyr2686.51, 4.3 Å; and C19-Tyr191EL2, 4.7 Å) (Fig. 1d). In Meta-II, the 13C19-13Cζ-tyrosine crosspeak intensity is lost, consistent with an increased separation between these tyrosines and the retinal C19 methyl group to >∼6–6.5 Å. We had previously attributed the loss of C19-tyrosine crosspeak intensity to translational motion of the retinal towards H5 and motion of EL2 (ref. 27). Below, we now show that the chemical shifts of both Tyr191EL2 and Tyr2686.51 change in Meta-II, suggesting that they move as well. This contributes to the loss of crosspeak intensity between the retinal C19 methyl resonance and the overlapping 13Cζ-Tyr191EL2 and Tyr2686.51 resonances. Moreover, we show that the alternative explanation, a flip of the β-ionone ring, does not occur until Meta-II decays (Supplementary Fig. 4 and Supplementary Note 3).
The retinal C20 methyl group is twisted out of the plane of the retinal polyene chain and primed to rotate in the clockwise direction when viewed from the Schiff’s base end of the retinal28,29,30. We previously concluded that the C20 methyl rotates towards EL2 on the basis of a C20-Tyr2686.51 crosspeak18,19. Here NMR measurements on the orientation of the C20 methyl group were undertaken with rhodopsin 13C-labelled at the ζ-carbon of tyrosine and regenerated with 13C12, 13C20 retinal (Fig. 1e,f). The row from the 2D DARR NMR spectrum of rhodopsin (black) through the C20 diagonal resonance yields an intense C12-C20 crosspeak at 132.2 p.p.m. (internal control) and a strong crosspeak to 13Cζ-Tyr2686.51 at 155.2 p.p.m. (refs 18, 19). On conversion to Meta-II (red), the C20-Tyr2686.51 crosspeak is only modestly weaker relative to the C12-C20 intra-retinal crosspeak, whereas the C12-Tyr2686.51 crosspeak becomes markedly weaker.
The crosspeak intensities between the 13Cζ-Tyr2686.51 and the retinal 13C12,13C20 resonances constrain the position of the tyrosine side chain relative to the retinal. The large difference in intensity in Meta-II between the C12 and C20 crosspeaks with Tyr2686.51 indicate that the C20 methyl group is closer to the tyrosine-Cζ carbon than the C12 methyl group. Molecular dynamics simulations performed on the basis of previous NMR constraints28 generated similar distances (∼4.3 Å) between Tyr2686.51 and the C12 and C20 carbons in Meta-II (Supplementary Table 3). The current NMR measurements indicate that the C20-Tyr2686.51 separation (∼4.4 Å) is shorter than the C12-Tyr2686.51 separation (∼5.6 Å). In contrast, the X-ray structures of Meta-II show the reverse, namely, the C12-Tyr2686.51 separation is shorter (∼3.9 Å) than the C20-Tyr2686.51 separation (∼5.9 Å). To accommodate the NMR distances, we propose (see below, Supplementary Fig. 5 and Supplementary Note 4) that H6 tilts inward on activation, a motion that is not captured in the molecular dynamics simulations.
Figure 1g presents build-up curves of crosspeak intensity for the retinal C12 and C20 carbons and Tyr2686.51. These curves provide a more accurate measure of internuclear distance than measurements at a single mixing time31,32. The distances in rhodopsin (black symbols) derived from this experiment are consistent with crystal structure distances (C12-Tyr2686.51, 4.5 Å; C20-Tyr2686.51, 4.1 Å; PDB-ID 1GZM (ref. 33)). In contrast, in Meta-II (red symbols) there is a marked difference between the distances estimated on the basis of the NMR build-up curves and the distances taken from the Meta-II crystal structures (Supplementary Table 3). For example, in the Meta-II-opsin and Meta-II-M257Y structures, the C12 carbon is much closer to the extracellular surface and consequently is in closer proximity to Tyr2686.51 than C20. The crystal structures place C12 at a distance of 3.8–4.0 Å from Tyr2686.51 (Supplementary Table 3), which should give rise to a stronger NMR crosspeak than observed. The intensity of the C12-Tyr2686.51 crosspeak drops considerably between rhodopsin and Meta-II indicating an increase in distance of C12 from Tyr2686.51. The NMR-derived distance is estimated to be ∼5.6±0.3 Å. Together, the NMR distance constraints presented above indicate that the orientation of the retinal in Meta-II trapped along its reaction coordinate is different than in the crystal structures.
Tyr268 movement suggests an inward tilt of H6 in Meta-II
Tyr2686.51 has the highest subfamily conservation (97%) in the visual GPCRs after Lys2967.43, the site of retinal attachment, indicating that its position and interactions are critically important within the visual receptors. A change in the position of Tyr2686.51 in Meta-II is detected by monitoring tyrosine crosspeaks to the retinal 13C10, 13C11 and the 13C14, 13C15 carbons. Tyr2686.51 exhibits crosspeaks to both the retinal 13C10 and 13C11 resonances in rhodopsin (Fig. 1h). These carbons are ∼4 Å from the Cζ position of Tyr2686.51 in the rhodopsin crystal structures. In contrast, no crosspeak intensity is observed between the retinal 13C14, 13C15 resonances and the 13Cζ resonance of Tyr2686.51 in rhodopsin. On conversion to Meta-II, the intensities of both the 13C10 and 13C11 crosspeaks to Tyr2686.51 decrease, while the 13C14 and 13C15 crosspeaks increase. Together, these changes indicate a shift in the position of Tyr2686.51 towards the Schiff base linkage in Meta-II.
The shift of Tyr2686.51 towards the retinal C14 and C15 carbons argues for a location between the retinal and EL2. We also observe strong chemical shift changes in the region of the SB consistent with a change in its electrostatic environment34. The retinal C15=N bond is strongly polarized; a large downfield chemical shift of C15 on conversion to Meta-II indicates that this carbon bears a substantial partial positive charge. The C=N polarization suggests close interactions with Glu1133.28, Glu181EL2 and/or Tyr2686.51.
The movement of Tyr2686.51 towards EL2 is supported by changes in the intensity of crosspeaks to Gly188EL2 and Cys187EL2 in Meta-II (Fig. 2). We have previously shown that Tyr2686.51 exhibits a weak crosspeak with Gly188EL2 in rhodopsin27. 2D NMR spectra using rhodopsin containing 13Cα-Gly, 13Cζ-Tyr and 13C=O-Cys allow us to highlight the Cys187EL2-Gly188EL2 pair since the Cys-13C=O to Gly-13Cα distance is short (2.4 Å) and consequently yields an intense crosspeak. As with the double 13C-labelled retinals, the Cys187EL2-Gly188EL2 pair serves as an internal control since the distance between these labelled 13C sites does not change between rhodopsin and Meta-II (Fig. 2b). On conversion to Meta-II, we observe two 13Cα-Gly-13Cζ-Tyr crosspeaks (Fig. 2b). The weaker crosspeak at 44.5 p.p.m. in Meta-II is assigned to Gly1143.29-Tyr178EL2 on the basis of the Y178F mutant (Fig. 2f). The stronger crosspeak at 43.1 p.p.m. is assigned to an overlap of the Gly188EL2-Tyr2686.51 crosspeak and at least one other Tyr-Gly contact. There is considerable loss of intensity at this position in the G188A mutant, and complete loss of the Cys187EL2-Gly188EL2 crosspeak (Fig. 2e).
(a) 2D DARR spectrum highlighting Tyr-Gly contacts in rhodopsin using rhodopsin containing 13Cζ-Tyr, 13Cα-Gly and 13C=O-Cys. In the rhodopsin crystal structure (PDB-ID 1U19; ref. 26), there are six Tyr(Cζ)-Gly(Cα) contacts, which are all located in the extracellular region of rhodopsin. These contacts involve five tyrosines and five glycines: Tyr10N−term-Gly3N−term, 3.9 Å; Tyr10N−term-Gly280EL3, 4.4 Å; Tyr29N−term-Gly101EL1, 4.0 Å; Tyr178EL2-Gly1143.29, 4.5 Å; Tyr191EL2-Gly188EL2, 5.2 Å; and Tyr2686.51-Gly188EL2, 5.3 Å. Above a are shown rows through the Tyr-Gly crosspeaks. The rows better illustrate the intensity change occurring in the Tyr2686.51-Gly188EL2 peak on activation. The observation that the Tyr178EL2-Gly1143.29 crosspeak does not change intensity is consistent with the lack of influence of the Y178F mutation on the Meta-I–Meta-II transition (Fig. 4g). (b) Wild-type Meta-II spectrum using rhodopsin containing 13Cζ-Tyr, 13Cα-Gly and 13C=O-Cys. (c,d) Region of the 2D DARR spectra of rhodopsin (black) and Meta-II (red) corresponding to 13Cζ-Tyr to 13C=O-Cys crosspeaks. Above c are shown rows through the 13Cζ-Tyr diagonal of rhodopsin and Meta-II. (e,f) G188A Meta-II and Y178F Meta-II spectra, respectively, using the same 13C-labelling scheme as above. Rows are shown through the Tyr-Gly crosspeaks in the G188A Meta-II and Y178F Meta-II spectra above e.
The Cys187EL2 13C=O label provides an additional probe for the location of Tyr2686.51 in Meta-II. There are two Cys-Tyr pairs in rhodopsin that are <6 Å apart (Tyr1363.51-Cys1403.55, 5.1 Å; and Tyr2065.41-Cys1674.56, 5.9 Å; PDB-ID 1U19 (ref. 26)). Both Tyr-Cys distances increase to >6 Å in the opsin crystal structure6. We observe a weak crosspeak in rhodopsin between a Cys-13C=O resonance (168–172 p.p.m.) and a Tyr-13Cζ resonance (∼156 p.p.m.; Fig. 2c). We observe two stronger crosspeaks in Meta-II at 172.5 p.p.m. (Fig. 2d), the same shift as the Cys187EL2 13C=O defined in Fig. 2b. One of the tyrosine contacts can be assigned to Tyr191EL2 on the basis of its unique 13Cζ chemical shift at 159.3 p.p.m. (ref. 18). We assign the other crosspeak to a Cys187EL2-Tyr2686.51 interaction.
The position of the 13Cζ-Tyr2686.51 relative to 13C15-labelled retinal, as well as to 13Cα-Gly188 and 1-13C-Cys187, indicates that Tyr2686.51 has shifted in the retinal-binding site. The separation between the 13Cζ Tyr2686.51 carbon and these sites is >6 Å in the Meta-II crystal structures, but much shorter in Meta-II trapped in the NMR experiments. The shorter distance is consistent with a small inward tilt of the extracellular end of H6. In addition to the crosspeak of retinal C20 with Tyr2686.51 in Meta-II (Fig. 1e), the C20 methyl group also exhibits a crosspeak with a glycine having a chemical shift of 46.5 p.p.m., which is assigned to Gly1213.36 on the basis of mutational studies (Supplementary Fig. 5 and Supplementary Note 4). Together, the retinal C20-Tyr2686.51 and -Gly1213.36 contacts indicate that the orientation of the C20 methyl group is roughly halfway between the orientation found in the Meta-II crystal structures and the orientation that previously resulted from molecular dynamics simulations (Supplementary Fig. 1).
Extracellular tyrosine switch releases constraints on H6
Tyr2686.51 on TM helix H6 is strongly hydrogen-bonded to Glu181EL2 on the β3 strand of EL2 and to Tyr191EL2 on the β4 strand of EL2 (Fig. 3d and Supplementary Table 5). EL2 caps the extracellular side of the retinal-binding site and Glu181EL2 has previously been identified as a second counter-ion to the retinal PSB, shifting position in the transition of rhodopsin to the Meta-I intermediate35. As a result, Tyr2686.51 is in a key location to couple hydrogen-bonding changes involving EL2 and the retinal PSB with motion of H6. The NMR data discussed above show that Tyr2686.51 has shifted relative to the retinal in Meta-II, while the large downfield chemical shift of Tyr191EL2 suggests that it has shifted relative to Glu181EL2.
1D NMR difference spectra are shown of rhodopsin minus Meta-I (orange) and rhodopsin minus Meta-II (black) using rhodopsin containing labelled 13Cɛ-Met (a), 13Cβ-Ser (b) or 13Cζ-Tyr (c). The rhodopsin spectra correspond to positive peaks, while the Meta-I and Meta-II spectra correspond to negative peaks. The assignments of the resonances have previously been reported18,19,38,64. (d) Crystal structure of rhodopsin in the region of Tyr2686.51 (PDB-ID 1U19; ref. 26).
To confirm that the changes in Tyr191EL2 and Tyr2686.51 are triggered by the shift in the position of Glu181EL2, we obtained both one-dimensional (1D) difference NMR spectra and 2D DARR NMR spectra (Supplementary Fig. 6 and Supplementary Note 5) focusing on residues in close proximity to Glu181EL2. 1D NMR difference spectra of 13Cɛ-Met-labelled rhodopsin show that the side-chain methyl group of Met2887.35 has an unusual downfield chemical shift in rhodopsin, which moves upfield in both Meta-I and Meta-II (Fig. 3a). Met2887.35 represents a contact point between H7 and EL2, and is in close proximity to Glu181EL2 in the rhodopsin structure. These chemical shift changes are consistent with the shift of the negatively charged Glu181EL2 side chain away from the methyl group of Met2887.35. Motion of Glu181EL2 towards the retinal PSB suggests that it comes into close proximity with Ser186EL2. 1D difference spectra of 13Cβ-labelled-serine reveal a downfield shift in Meta-I and upfield shift in Meta-II (Fig. 3b). The chemical shift of the 13Cβ carbon of serine is sensitive to Cβ-OH hydrogen bonding, and the observed chemical shift changes are consistent with increased hydrogen bonding in Meta-I (or earlier photointermediate)36 and decreased hydrogen bonding in Meta-II.
In rhodopsin, the chemical shifts of both 13Cζ-Tyr191EL2 and 13Cζ-Tyr2686.51 are at ∼156 p.p.m. (Fig. 1d). There is a positive peak in the rhodopsin–Meta-I difference spectrum at this position, but no corresponding negative peaks (Fig. 3c). In contrast, we have previously assigned18,19 the downfield resonance at 159.3 p.p.m. in the rhodopsin–Meta-II difference spectrum to Tyr191EL2 and have suggested that this unusual chemical shift is due to direct interaction with Glu181EL2.
Together, the chemical shift changes of Met2887.35 and Ser186EL2 indicate that the counter-ion shift of Glu181EL2 in Meta-I precedes the motion of Tyr191EL2 in Meta-II. Tyr191EL2 is tightly packed against Ala2726.55 on H6, one helical turn from Tyr2686.51, and its motion towards Glu181EL2 would release steric constraints hindering the inward tilt of the extracellular end of H6 in the dark state.
Conserved extracellular tyrosine residues stabilize Meta-I
The interactions involving Tyr191EL2, Tyr2686.51 and Met2887.35 indicate that these residues mediate the transition between Meta-I and Meta-II. To further explore their influence on activation, we undertook FTIR studies on the Y191F, Y268F and M288L mutants alongside wild-type rhodopsin. We previously showed that the amide I infrared band along with the C=O stretching modes of the protonated Glu1223.37 and Asp832.50 carboxylic groups provide conformationally sensitive markers and can be used to deconvolute the Meta-I ⇔ Meta-II equilibrium into the inactive Meta-I state and several Meta-II substates (Meta-IIa, Meta-IIb and Meta-IIbH+; Supplementary Fig. 7 and Supplementary Note 6)22.
A two-state equilibrium is observed at 0 °C between Meta-I and Meta-IIbH+ using wild-type rhodopsin reconstituted into egg phosphocholine membranes, where the transition between Meta-I and Meta-IIbH+ has an apparent pK of ∼6.8 and is similar for each of these FTIR markers (Fig. 4a). At higher temperature, the titration curves become more complex with stabilization of the Meta-IIb substate (Supplementary Fig. 7). The Y268F, Y191F and M288L mutants exhibit complex titration curves (Fig. 4b,c,e) even at 0 °C due to a strong forward shift of the Meta-I ⇔ Meta-IIb equilibrium to Meta-IIb. For wild-type rhodopsin, an increase in temperature will lead to a shift of the equilibrium as the stabilization of Meta-IIb has an entropic component22. In contrast for the mutants, the forward shift reflects the loss of stabilizing interactions in Meta-I.
Wild-type (a), and the Y268F (b), Y191F (c), Y192F (d), M288L (e), M288A (f), Y178F (g) and W175F (h) mutants. FTIR difference spectra were obtained at 0 °C as a function of pH, and the relative contributions of Meta-I and Meta-IIbH+ were determined on the basis of reference spectra (Supplementary Fig. 7). Spectral decomposition was performed between 1,800 and 1,600 cm−1. This range is the most diagnostic of the Meta-I–Meta-II transition and comprises the amide I vibrations of the protein backbone and the C=O stretch of protonated carboxylic acids Glu1223.37 and Asp832.50. The vertical dashed lines are at the positions of the inflection points.
The Y192F mutant (Fig. 4d) exhibits a forward shift of the Meta-I ⇔ Meta-IIb equilibrium as in Y191F, but the effect is not as large. The M288A mutant (Fig. 4f) reveals a downshift of the apparent pKa, indicating stabilization of the Meta-I intermediate on mutation. Thus, depending on the specific mutation at Met2887.35, the Meta-I ⇔ Meta-II equilibrium can be shifted in either direction. Finally, the equilibrium is not affected by the W175F and Y178F mutants (Fig. 4g,h) indicating that there are conserved aromatic residues in EL2 that do not have the effects observed for Tyr191EL2 and Tyr2686.51.
Discussion
Activation of the visual receptors is triggered by changes in both steric and electrostatic interactions caused by 11-cis to all-trans isomerization of the retinal and deprotonation of the retinal PSB. Retinal isomerization occurs in the first step of the rhodopsin photoreaction, while deprotonation occurs on formation of the Meta-II intermediate37. The specific retinal–protein interactions observed by NMR and FTIR provide insights into how these two components of the activation mechanism work together to drive the outward tilt of TM helix H6. We liken these connected activation events to the firing mechanism of a two-stage trigger.
The first stage of the trigger results from isomerization-induced steric contact between the β-ionone ring and residues on helices H3, H5 and H6 (Fig. 5). NMR38 and fluorescence39 studies both reveal that the first major change in receptor conformation is rotation of H6 in the formation of Meta-I, which breaks the intracellular Arg1353.50-Glu2476.30 ionic lock40 and results in Trp2656.48 displacement38,41. These rearrangements are accompanied by small changes at the cytoplasmic end of H5 (refs 39, 42). Our current results indicate direct contact of the β-ionone ring with His2115.46 and Phe2616.44 in Meta-II (Fig. 5a). Previously, we reported that activation results in a loss of the retinal C18-Trp2656.48 contact19. Mutation of either Phe2616.44 or Trp2656.48 lowers the initial rate of G protein transducin activation, ∼20–60% for F261A rhodopsin43 and ∼90% in W265F rhodopsin44, consistent with their roles in the activation mechanism. The steric contact of the β-ionone ring with Phe2616.44 observed in Meta-II suggests that the phenylalanine ring acts a lever for rotation of H6. This residue is part of a transmission switch in the conserved TM core of GPCRs15,45.
(a) Residues interacting with the β-ionone ring in the dark state of rhodopsin (PDB-ID 1U19; ref. 26). The first stage of the two-stage trigger involves isomerization of the 11-cis-retinal chromophore within a tightly packed retinal-binding pocket. Steric interactions arise because the shape of the retinal-binding site fits the 11-cis isomer but does not accommodate the longer all-trans form10. The position of the β-ionone ring defined by distance measurements with Met2075.42 and His2115.46 on H5 (Supplementary Fig. 3 and refs 18, 19), as well as Phe2616.44 and Trp2656.48 on H6 (Fig. 1 and refs 18, 19, 63) are consistent with strong steric interactions between the β-ionone ring in this region of the retinal-binding site. Motion of H5 and H6 is present at the Meta I stage as a result of isomerization38,39,42. (b) Cartoon representation of the structural changes occurring on the extracellular side of the retinal-binding site in rhodopsin comprising the second stage of the two-stage trigger. Retinal isomerization leads to a change in the position of Glu181EL2 in Meta-I (ref. 35). Deprotonation of the retinal PSB and protonation of Glu1133.28 leads to a further rearrangement of the hydrogen-bonding network connecting Glu1133.28 to Tyr191EL2. Both Tyr2686.51 and Tyr191EL2 shift towards Glu181EL2 to maintain hydrogen-bonding interactions. The proposed motion of the extracellular side of H6 due to Tyr191EL2 motion is coupled to the outward pivot of the intracellular side of H6. (c) Schematic representation of the two-stage trigger mechanism.
The conserved proline (Pro2155.50) on H5 is also a component of the transmission switch and results in a free backbone carbonyl at His2115.46. This carbonyl forms an interhelical hydrogen bond with Glu1223.37 in rhodopsin, which is replaced in Meta-II by a direct interaction between the His2115.46 and Glu1223.37 side chains20. The strong steric contacts of the β-ionone ring with His2115.46 and Glu1223.37 that drive this transition allow H5 to reorient in Meta-II (ref. 46). Mutation of Glu1223.37 breaks their direct interaction and destabilizes Meta-II (ref. 47). Retinal analogues in which the ring is truncated shift the conformational equilibrium between the Meta-I and Meta-II intermediates towards the inactive Meta-I state24,48. Thus, the β-ionone ring is a key element in both the first stage of the activation trigger and in stabilizing the Meta-II intermediate.
In Supplementary Fig. 4, we follow the change in orientation of the C18 methyl group of the β-ionone ring during the decay of Meta-II and observe a gain in contact with Tyr191EL2 consistent with a flip of the β-ionone ring and suggesting that the crystal structures are capturing a relaxed, low-energy conformation that has retained the capacity to bind all-trans retinal. The final conformation observed by NMR is possibly Meta-III, in which the SB is reprotonated49. The observation of an all-trans retinal PSB following Meta-II suggests that the reprotonation event is the driving force for rotation along the long axis of the retinal during Meta-II decay. Together, our NMR results are consistent with the low temperature trapped Meta-II intermediate being a transient, high-energy state that is formed along the reaction coordinate and consequently captures the conformation triggering receptor activation.
The second stage in the two-stage trigger mechanism involves deprotonation of the retinal PSB along with steric interactions of the retinal C19 and C20 methyl groups with Tyr191EL2 and Tyr2686.51, respectively. Tyr191EL2 and Tyr2686.51 are part of an extensive hydrogen-bonding network that stretches through Glu181EL2 to Glu1133.28, the PSB counter-ion (Fig. 5b, Supplementary Fig. 8 and Supplementary Note 7). Isomerization results in motion of the retinal PSB proton away from the stabilizing interaction with Glu1133.28. This ultimately leads to proton transfer from the PSB to Glu1133.28 in Meta-II. Nevertheless, the change in the position of the PSB proton must already be sensed by nearby residues in the Meta-I intermediate before proton transfer. The counter-ion shift mechanism envisions motion of the charged Glu181EL2 side chain towards the PSB end of the retinal in Meta-I (ref. 35). The large changes in the NMR chemical shifts of Ser186EL2 and Met2887.35 observed in Meta-I are consistent with Glu181EL2 motion. However, motion of Glu181EL2 in Meta-I precedes the change in the position of Tyr191EL2 and Tyr2686.51 (Fig. 5b) because the striking downfield chemical shift of Tyr191EL2 is not observed until Meta-II. Moreover, the proposed mechanism argues that the tilt of H6 occurs following PSB deprotonation and movement of Tyr191EL2, in agreement with EPR measurements50. These results are consistent with an earlier FTIR study suggesting a role for Tyr268 and H6 reorientation in rhodopsin activation51.
The C19 and C20 methyl groups contribute to the rearrangement of the extracellular hydrogen-bonding network. These methyl groups extend from the polyene chain and are located on opposite sides of the cis C11=C12 bond (Fig. 1a). The crystal structures of rhodopsin show that the C20 methyl group is primed to rotate in a clockwise direction on isomerization28,29,30, the only potential obstacle being Tyr2686.51. The C20-Tyr2686.51 crosspeak observed in Meta-II by NMR is consistent with this trajectory of the C20 methyl group and suggests that a steric contact with Tyr2686.51 contributes to the rearrangement of the hydrogen-bonding network stabilizing Meta-I on the extracellular surface of the receptor. In rhodopsin regenerated with an 11-cis retinal analogue lacking the C20 methyl group, the photoreaction is slowed52 and the quantum yield is reduced53.
The C19 methyl group of retinal is oriented towards EL2 in the dark state of rhodopsin and is tightly packed between Ile189EL2 and Tyr191EL2 (Supplementary Fig. 8). Retinal analogues lacking the C19 methyl group cannot activate rhodopsin54, while increasing the size of the C19 methyl to either an ethyl or propyl group increases receptor activity in the dark, even though the retinal still remains covalently bound and in its 11-cis configuration43,55,56. On isomerization, the C19 methyl group rotates in the opposite direction (that is, anticlockwise) to the C20 methyl group28,29,30, and consequently would have increased steric contact with Tyr191EL2. Removal of the C19 methyl group eliminates this interaction, while the ethyl or propyl substitutions at C19 result in a steric clash with Tyr191EL2, which would disrupt the Tyr191EL2-Tyr2686.51 hydrogen-bonding network in the dark, in a manner similar to light-driven isomerization in the wild-type receptor.
The net effect of PSB deprotonation and rearrangement of the hydrogen-bonding network involving EL2 is a shift of Tyr191EL2 away from H6. We propose that this motion allows the extracellular end of H6 to pivot inward. In the visual pigments, Tyr191EL2 has a high level of sequence identity (61%), and an overall level of conservation of 83% as either tyrosine or tryptophan. Interestingly, even though Tyr191EL2 and Tyr2686.51 contribute to Meta-I stability (Fig. 4), mutation of either of these residues results in a substantial drop in G protein transducin activation44,57. This dual influence of Tyr191EL2 and Tyr2686.51 on Meta-I stability and Meta-II activity is consistent with one set of hydrogen-bonding interactions stabilizing Meta-I and a second set stabilizing Meta-II (Fig. 5). The observation that Tyr191EL2 and Tyr2686.51 strongly stabilize the Meta-I state suggests that in crystal structures of opsin and/or Meta-II, the inactive-state hydrogen-bonding network reforms in this region under the conditions used for crystallization.
The proposed two-stage trigger mechanism builds on the global toggle switch mechanism developed on the basis of ligand-activated GPCRs, which envisions a vertical see-saw movement of helix H6 with conserved Pro2676.50 serving as the pivot point58. In the case of the visual pigments, Meta-I corresponds to the inactive state of a ligand-activated GPCR, where ligand binding simply allows H6 to pivot to open up the intracellular G protein-binding pocket.
Methods
Expression and purification of 13C-labelled bovine rhodopsin
Isotope-enriched bovine opsin59 was expressed using inducible HEK293S cell lines60, which are widely used for the production of recombinant proteins and viruses. The primers used for construction of the rhodopsin mutants are listed in Supplementary Table 6. The original cell lines were obtained from Jeremy Nathans (Johns Hopkins University), but not authenticated or tested for mycoplasma contamination as they were used only for protein expression. The expressed opsin was generated into rhodopsin through incubation with ∼30 μM 11-cis retinal and extracted from membranes using 1% (w/v) n-β-D-dodecyl maltopyranoside (DDM) in PBS pH 7.4. The detergent-solubilized lysates were separated from the remaining cell debris via centrifugation at 25,000g for 30 min at 4 °C. The detergent-solubilized lysate was applied to a rho-1D4-Sepharose column via gravity flow to capture rhodopsin. The flow-through volume was saved and the amount of unbound receptor assayed by western blot using rho-1D4. The column was then washed with 10 volumes of 0.02% DDM in PBS (pH 7.2). A subsequent wash step with 0.02% DDM in sodium phosphate buffer (pH 6.0) was used to equilibrate the column for elution. The receptor was then eluted from the rho-1D4-column with 0.02% DDM in sodium phosphate buffer (pH 6.0) supplemented with a nonapeptide (100 μM) whose sequence corresponds to the carboxyl terminus of rhodopsin (TETSQVAPA).
For capture of Meta-I, the DDM of the solubilized rhodopsin was reduced to 0.02% w/v and subsequently exchanged for 0.02–0.05% w/v digitonin on the 1D4-Sepharose column38. Rhodopsin was eluted in 2 mM phosphate buffer (pH 7.0) containing digitonin (0.02–0.05% w/v) and 100 μM C-terminal nonapeptide. The pooled, eluted rhodopsin fractions were concentrated to a final volume of ∼400 μl using Centricon devices with a 10 kDa molecular weight cutoff (Amicon, Bedford, MA), followed by further concentration under argon gas to a volume of ∼100 μl. All buffers were prepared fresh before purification38.
Synthesis of 13C-labelled retinals
The 13C-labelled retinals were synthesized using unlabelled and 13C-labelled small molecule precursors. 13C5, 13C18 retinal was prepared by condensation of 4-bromo-2-methyl-2-butane with 1,2-13C-ethyl acetoacetate, and conversion of the labelled 6-methyl-5-hepta-2-one to 13C5, 13C18 retinal using conventional methods61. 13C10, 13C11 retinal was synthesized by condensation of β-ionone with 13CH313CN, which yields labelled β-ionylideneacetonitrile after dehydration. The latter was converted to 13C10, 13C11 retinal61. 13C12, 13C20 retinal was synthesized using aldol condensation of 2,3-dimethyl-5-(2',6',6'-trimethyl-1'-cyclohexene-1'-yl)-2,4-pentadienal with 1,3-13C acetone and conversion to labelled retinal. 13C14, 13C15 retinal was prepared by condensation of 5,6-dimethyl-8-(2',6',6'-trimethyl-1'-cyclohexene-1'-yl)-3,5,7-octatriene-2-one with di-13C-labelled acetonitrile and transformation to labelled retinal by conventional methods61. 13C18, 13C20 retinal was prepared by condensation of citral with 1,3-di-13C acetone and conversion of ketone product to labelled retinal61.
Solid-state NMR spectroscopy
Solid-state NMR experiments were conducted at static field strengths of either 500 or 600 MHz using a 4 mm magic angle spinning (MAS) probe with a spinning rate of 10–12 kHz. NMR measurements were first made on rhodopsin in the dark at −83 °C. For Meta- I and -II, the NMR MAS rotor containing rhodopsin was ejected from the NMR probe, the NMR cap on the rotor was removed and the sample was illuminated for 1–2 min using a 400 W lamp with a 495 nm long-pass filter at room temperature (Meta-II)62 or 4 °C (Meta-I)38. The cap was then replaced and the rotor inserted into a pre-cooled NMR probe where the sample temperature was able to reach −83 °C within ∼5 min. We estimate that the conversion from rhodopsin to Meta-I in digitonin is >90% on the basis of ultraviolet/visible absorption and NMR spectra38. After conversion, we estimate that there is a loss of <10% of the Meta-I intermediate to Meta-II and opsin before the sample is cooled to −83 °C for NMR measurements38. We estimate that the conversion from rhodopsin to Meta-II in DDM is >90% and the loss of Meta-II to opsin is <5% before the sample is cooled to −83 °C (ref. 62).
The build-up curves were obtained by collecting DARR NMR spectra as a function of the mixing time during which magnetization is exchanged. The known curves were derived from measurements within rhodopsin at fixed distances (for example, retinal C5-C18, 1.4 Å; C8-C19, C12-C20, 2.4 Å; Cys110 Cβ-Cys187 Cβ, 3.6 Å; Cys187 Cβ-Gly188 Cα, 4.6 Å; Cys187 Cβ-Gly188 C=O, 5.3 Å). Spin diffusion is limited by the sparse labelling schemes that are typically used. The NMR assignments were based on mutation and/or mapping out specific correlations with unique resonances that have previously been assigned18,19,34,38,63,64.
All 13C solid-state MAS NMR spectra were externally referenced to the 13C resonance of neat TMS at 0 p.p.m. at room temperature. Using TMS as the external reference, we calibrated the carbonyl resonance of solid glycine at 176.46 p.p.m. The chemical shift difference between 13C of DSS in D2O relative to neat TMS is 2.01 p.p.m. The spectra shown were generally repeated at least twice on samples purified from different cell growths without deviations of more than ±0.2 p.p.m. in the reported chemical shifts.
FTIR spectroscopy
FTIR difference spectroscopy was performed with a Bruker Vertex 70 spectrometer with a mercury–cadmium–telluride detector22,23,24. Spectra were recorded by using time-resolved rapid-scan FTIR methods with a spectral resolution of 4 cm−1. The wild-type and mutant rhodopsin samples were purified from HEK293S cell membranes in DDM and reconstituted at a 1:200 molar ratio into egg phosphocholine using biobeads for detergent removal. The results shown in Fig. 4 are for samples in phosphocholine membranes obtained at 0 °C. Samples were prepared by drying solutions of rhodopsin between two CaF2 windows and then pre-equilibrating the sample with buffer (200 mM Bis-Tris propane or MES at pH 5.0 and 5.5) at the appropriate pH. Photolysis was carried out using an LED array centred at 530 nm for 1 s, and experiments were performed with an acquisition time of 12 s.
The conditions for NMR and FTIR are different. For NMR, the analysis relies on complete conversion (>90%) to Meta-I or Meta-II, which is facilitated by solubilization in digitonin or DDM, respectively. Rhodopsin is monomeric in DDM and is able to activate the G protein transducin65. Digitonin is unusual in that its hydrophobic end is composed of a rigid spirostan steroid moiety rather than flexible fatty acyl chains. The rigid framework effectively blocks the transition from Meta-I to Meta-II 66. For FTIR, the analysis uses difference methods in which the FTIR spectrum of Meta-I or Meta-II is subtracted from the spectrum of rhodopsin. Only the vibrations that change in frequency or intensity contribute to the difference spectrum. Changes in pH or temperature can be used to shift the equilibrium between Meta-I and Meta-II. At low temperatures (below ∼10 °C), the Meta-I ⇔ Meta-II equilibrium reflects a two-state transition in both DDM and phosphocholine bilayers, which breaks down into a series of Meta-II substates at higher temperature22,50. In both DDM and phosphocholine bilayers the conversion to Meta-IIbH+ happens rapidly (millisecond–second timescale) at 20 °C (refs 22, 50), which requires the use of time-resolved methods to follow the transition. For NMR, we convert fully to Meta-II at room temperature, but require several minutes to low-temperature trap the Meta-II intermediate, which we assume is predominately Meta-IIbH+, before it decays.
Data availability
NMR data that support the findings of this study have been deposited in the BMRB databank with the accession codes 26812 (rhodopsin) and 26813 (Meta-II) (http://www.bmrb.wisc.edu). The coordinates of Meta-II from the guided molecular dynamics simulations and remaining data are available on reasonable request from the corresponding author.
Additional information
How to cite this article: Kimata, N. et al. Retinal orientation and interactions in rhodopsin reveal a two-stage trigger mechanism for activation. Nat. Commun. 7:12683 doi: 10.1038/ncomms12683 (2016).
References
Smith, S. O. Structure and activation of the visual pigment rhodopsin. Ann. Rev. Biophys. 39, 309–328 (2010).
Park, P. S. H., Lodowski, D. T. & Palczewski, K. Activation of G protein-coupled receptors: beyond two-state models and tertiary conformational changes. Annu. Rev. Pharmacol. Toxicol. 48, 107–141 (2008).
Kobilka, B. K. G protein coupled receptor structure and activation. Biophys. Biochim. Acta Biomem. 1768, 794–807 (2007).
Katritch, V., Cherezov, V. & Stevens, R. C. Diversity and modularity of G protein-coupled receptor structures. Trends Pharmacol. Sci. 33, 17–27 (2012).
Venkatakrishnan, A. J. et al. Molecular signatures of G-protein-coupled receptors. Nature 494, 185–194 (2013).
Park, J. H., Scheerer, P., Hofmann, K. P., Choe, H. W. & Ernst, O. P. Crystal structure of the ligand-free G-protein-coupled receptor opsin. Nature 454, 183–187 (2008).
Scheerer, P. et al. Crystal structure of opsin in its G-protein-interacting conformation. Nature 455, 497–502 (2008).
Rasmussen, S. G. F. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011).
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three dimensional models and computational probing of structure-function relations in G-protein coupled receptors. Methods Neurosci. 25, 366–428 (1995).
Matsumoto, H. & Yoshizawa, T. Recognition of opsin to longitudinal length of retinal isomers in formation of rhodopsin. Vision Res. 18, 607–609 (1978).
Deupi, X. & Kobilka, B. K. Energy landscapes as a tool to integrate GPCR structure, dynamics, and function. Physiology 25, 293–303 (2010).
Standfuss, J. et al. The structural basis of agonist-induced activation in constitutively active rhodopsin. Nature 471, 656–660 (2011).
Deupi, X. et al. Stabilized G protein binding site in the structure of constitutively active metarhodopsin-II. Proc. Natl Acad. Sci. USA 109, 119–124 (2012).
Farrens, D. L., Altenbach, C., Yang, K., Hubbell, W. L. & Khorana, H. G. Requirement of rigid-body motion of transmembrane helices for light activation of rhodopsin. Science 274, 768–770 (1996).
Deupi, X. Relevance of rhodopsin studies for GPCR activation. Biochim. Biophys. Acta 1837, 674–682 (2014).
Cooper, A. Energy uptake in the first step of visual excitation. Nature 282, 531–533 (1979).
Wald, G., Durell, J. & St George, R. C. C. The light reaction in the bleaching of rhodopsin. Science 111, 179–181 (1950).
Ahuja, S. et al. Helix movement is coupled to displacement of the second extracellular loop in rhodopsin activation. Nat. Struct. Mol. Biol. 16, 168–175 (2009).
Ahuja, S. et al. Location of the retinal chromophore in the activated state of rhodopsin. J. Biol. Chem. 284, 10190–10201 (2009).
Choe, H. W. et al. Crystal structure of metarhodopsin II. Nature 471, 651–655 (2011).
Okada, T. et al. X-ray diffraction analysis of three-dimensional crystals of bovine rhodopsin obtained from mixed micelles. J. Struct. Biol. 130, 73–80 (2000).
Mahalingam, M., Martinez-Mayorga, K., Brown, M. F. & Vogel, R. Two protonation switches control rhodopsin activation in membranes. Proc. Natl Acad. Sci. USA 105, 17795–17800 (2008).
Standfuss, J., Zaitseva, E., Mahalingam, M. & Vogel, R. Structural impact of the E113Q counterion mutation on the activation and deactivation pathways of the G protein-coupled receptor rhodopsin. J. Mol. Biol. 380, 145–157 (2008).
Vogel, R., Siebert, F., Lüdeke, S., Hirshfeld, A. & Sheves, M. Agonists and partial agonists of rhodopsin: retinals with ring modifications. Biochemistry 44, 11684–11699 (2005).
Han, M., Smith, S. O. & Sakmar, T. P. Constitutive activation of opsin by mutation of methionine 257 on transmembrane helix 6. Biochemistry 37, 8253–8261 (1998).
Okada, T. et al. The retinal conformation and its environment in rhodopsin in light of a new 2.2 Å crystal structure. J. Mol. Biol. 342, 571–583 (2004).
Patel, A. B. et al. Coupling of retinal isomerization to the activation of rhodopsin. Proc. Natl Acad. Sci. USA 101, 10048–10053 (2004).
Hornak, V. et al. Light activation of rhodopsin: Insights from molecular dynamics simulations guided by solid-state NMR distance restraints. J. Mol. Biol. 396, 510–527 (2010).
Nakamichi, H. & Okada, T. Crystallographic analysis of primary visual photochemistry. Angew. Chem. Int. Ed. 45, 4270–4273 (2006).
Schreiber, M., Sugihara, M., Okada, T. & Buss, V. Quantum mechanical studies on the crystallographic model of bathorhodopsin. Angew. Chem. Int. Ed. 45, 4274–4277 (2006).
Takegoshi, K., Nakamura, S. & Terao, T. 13C-1H dipolar-assisted rotational resonance in magic-angle spinning NMR. Chem. Phys. Lett. 344, 631–637 (2001).
Takegoshi, K., Nakamura, S. & Terao, T. 13C-1H dipolar-driven 13C-13C recoupling without 13C rf irradiation in nuclear magnetic resonance of rotating solids. J. Chem. Phys. 118, 2325–2341 (2003).
Li, J., Edwards, P. C., Burghammer, M., Villa, C. & Schertler, G. F. X. Structure of bovine rhodopsin in a trigonal crystal form. J. Mol. Biol. 343, 1409–1438 (2004).
Ahuja, S. et al. 6-s-cis conformation and polar binding pocket of the retinal chromophore in the photoactivated state of rhodopsin. J. Am. Chem. Soc. 131, 15160–15169 (2009).
Yan, E. C. Y. et al. Retinal counterion switch in the photoactivation of the G protein-coupled receptor rhodopsin. Proc. Natl Acad. Sci. USA 100, 9262–9267 (2003).
Yan, E. C. Y. et al. Photointermediates of the rhodopsin S186A mutant as a probe of the hydrogen-bond network in the chromophore pocket and the mechanism of counterion switch. J. Phys. Chem. C 111, 8843–8848 (2007).
Wald, G. Molecular basis of visual excitation. Science 162, 230–239 (1968).
Eilers, M. et al. Structural transitions of transmembrane helix 6 in the formation of metarhodopsin I. J. Phys. Chem. B 116, 10477–10489 (2012).
Ye, S. X. et al. Tracking G-protein-coupled receptor activation using genetically encoded infrared probes. Nature 464, 1386–1389 (2010).
Ballesteros, J. A. et al. Activation of the β2-adrenergic receptor involves disruption of an ionic lock between the cytoplasmic ends of transmembrane segments 3 and 6. J. Biol. Chem. 276, 29171–29177 (2001).
Ruprecht, J. J., Mielke, T., Vogel, R., Villa, C. & Schertler, G. F. X. Electron crystallography reveals the structure of metarhodopsin I. EMBO J. 23, 3609–3620 (2004).
Elgeti, M. et al. Conserved Tyr223(5.58) plays different roles in the activation and G-protein interaction of rhodopsin. J. Am. Chem. Soc. 133, 7159–7165 (2011).
Han, M., Lin, S. W., Minkova, M., Smith, S. O. & Sakmar, T. P. Functional interaction of transmembrane helices 3 and 6 in rhodopsin—replacement of phenylalanine 261 by alanine causes reversion of phenotype of a glycine 121 replacement mutant. J. Biol. Chem. 271, 32337–32342 (1996).
Nakayama, T. A. & Khorana, H. G. Mapping of the amino acids in membrane-embedded helices that interact with the retinal chromophore in bovine rhodopsin. J. Biol. Chem. 266, 4269–4275 (1991).
Deupi, X. & Standfuss, J. Structural insights into agonist-induced activation of G-protein-coupled receptors. Curr. Opin. Struct. Biol. 21, 541–551 (2011).
Kimata, N. et al. Free backbone carbonyls mediate rhodopsin activation. Nat. Struct. Mol. Biol. 23, 738–743 (2016).
Imai, H. et al. Single amino acid residue as a functional determinant of rod and cone visual pigments. Proc. Natl Acad. Sci. USA 94, 2322–2326 (1997).
Jäger, F. et al. Interactions of the β-ionone ring with the protein in the visual pigment rhodopsin control the activation mechanism. An FTIR and fluorescence study on artificial vertebrate rhodopsins. Biochemistry 33, 7389–7397 (1994).
Heck, M. et al. Signaling states of rhodopsin: Formation of the storage form, metarhodopsin III, from active metarhodopsin II. J. Biol. Chem. 278, 3162–3169 (2003).
Knierim, B., Hofmann, K. P., Ernst, O. P. & Hubbell, W. L. Sequence of late molecular events in the activation of rhodopsin. Proc. Natl Acad. Sci. USA 104, 20290–20295 (2007).
DeLange, F. et al. Tyrosine structural changes detected during the photoactivation of rhodopsin. J. Biol. Chem. 273, 23735–23739 (1998).
Wang, Q. et al. Femtosecond spectroscopy of a 13-demethylrhodopsin visual pigment analogue: the role of nonbonded interactions in the isomerization process. J. Phys. Chem. 100, 17388–17394 (1996).
Kochendoerfer, G. G., Verdegem, P. J. E., van der Hoef, I., Lugtenburg, J. & Mathies, R. A. Retinal analog study of the role of steric interactions in the excited state isomerization dynamics of rhodopsin. Biochemistry 35, 16230–16240 (1996).
Corson, D. W. et al. Relief of opsin desensitization and prolonged excitation of rod photoreceptors by 9-desmethylretinal. Proc. Natl Acad. Sci. USA 91, 6958–6962 (1994).
Han, M., Groesbeek, M., Sakmar, T. P. & Smith, S. O. The C9 methyl group of retinal interacts with glycine-121 in rhodopsin. Proc. Natl Acad. Sci. USA 94, 13442–13447 (1997).
Han, M., Lin, S. W., Smith, S. O. & Sakmar, T. P. The effects of amino acid replacements of glycine 121 on transmembrane helix 3 of rhodopsin. J. Biol. Chem. 271, 32330–32336 (1996).
Doi, T., Molday, R. S. & Khorana, H. G. Role of the intradiscal domain in rhodopsin assembly and function. Proc. Natl Acad. Sci. USA 87, 4991–4995 (1990).
Schwartz, T. W., Frimurer, T. M., Holst, B., Rosenkilde, M. M. & Elling, C. E. Molecular mechanism of 7TM receptor activation—a global toggle switch model. Annu. Rev. Pharmacol. Toxicol. 46, 481–519 (2006).
Oprian, D. D., Molday, R. S., Kaufman, R. J. & Khorana, H. G. Expression of a synthetic bovine rhodopsin gene in monkey kidney cells. Proc. Natl Acad. Sci. USA 84, 8874–8878 (1987).
Reeves, P. J., Thurmond, R. L. & Khorana, H. G. Structure and function in rhodopsin: high level expression of a synthetic bovine opsin gene and its mutants in stable mammalian cell lines. Proc. Natl Acad. Sci. USA 93, 11487–11492 (1996).
Lugtenburg, J. The synthesis of 13C-labeled retinals. Pure Appl. Chem. 57, 753–762 (1985).
Eilers, M., Ying, W. W., Reeves, P. J., Khorana, H. G. & Smith, S. O. Magic angle spinning nuclear magnetic resonance of isotopically labeled rhodopsin. Methods Enzymol. 343, 212–222 (2002).
Crocker, E. et al. Location of Trp265 in metarhodopsin II: implications for the activation mechanism of the visual receptor rhodopsin. J. Mol. Biol. 357, 163–172 (2006).
Goncalves, J. A. et al. Highly conserved tyrosine stabilizes the active state of rhodopsin. Proc. Natl Acad. Sci. USA 107, 19861–19866 (2010).
Ernst, O. P., Gramse, V., Kolbe, M., Hofmann, K. P. & Heck, M. Monomeric G protein-coupled receptor rhodopsin in solution activates its G protein transducin at the diffusion limit. Proc. Natl Acad. Sci. USA 104, 10859–10864 (2007).
Matthews, R. G., Hubbard, R., Brown, P. K. & Wald, G. Tautomeric forms of metarhodopsin. J. Gen. Physiol. 47, 215–240 (1963).
Acknowledgements
This work was supported by a grant from the NIH (GM 41412) to S.O.S.
Author information
Authors and Affiliations
Contributions
S.O.S. and P.J.R. conceived and designed the work; N.K., A.P. and M.E. prepared isotopically labelled rhodopsin; C.A.O. and P.J.R. prepared rhodopsin mutants; A.H. and M.S. synthesized isotopically labelled retinals; N.K., M.E. and M.Z. collected and analysed NMR data; E.Z. and R.V. collected and analysed FTIR data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-8, Supplementary Tables 1-6, Supplementary Notes 1-7, Supplementary References (PDF 8457 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kimata, N., Pope, A., Eilers, M. et al. Retinal orientation and interactions in rhodopsin reveal a two-stage trigger mechanism for activation. Nat Commun 7, 12683 (2016). https://doi.org/10.1038/ncomms12683
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms12683
This article is cited by
-
Strategies for acquisition of resonance assignment spectra of highly dynamic membrane proteins: a GPCR case study
Journal of Biomolecular NMR (2023)
-
Ultrafast structural changes direct the first molecular events of vision
Nature (2023)
-
Conformational perturbation, allosteric modulation of cellular signaling pathways, and disease in P23H rhodopsin
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.