Abstract
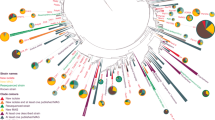
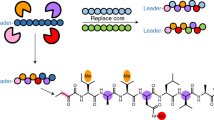
A large family of cytotoxic cyclic peptides exemplified by the patellamides has been isolated from ascidians harboring the obligate cyanobacterial symbionts Prochloron spp.1,2,3,4,5. Genome sequence analysis of these symbionts has revealed that Prochloron spp. synthesize patellamides by a ribosomal pathway6. To understand how this pathway evolved to produce a suite of related metabolites, we analyzed 46 prochloron-containing ascidians from the tropical Pacific Ocean for the presence of patellamide biosynthetic genes and taxonomic markers. Here, we show that Prochloron spp. generate a diverse library of patellamides using small, hypervariable cassettes within a conserved genetic background. Each symbiont strain contains a single pathway, and mixtures of symbionts within ascidians lead to the accumulation of chemical libraries. We used this information to engineer the production of a new cyclic peptide in Escherichia coli, thereby demonstrating the power of comparative analysis of closely related symbiotic pathways to direct the genetic synthesis of new molecules.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Sings, H.L. & Rinehart, K.L. Compounds produced from potential tunicate-blue-green algal symbiosis: a review. J. Ind. Microbiol. Biotechnol. 17, 385–396 (1996).
Schmidt, E.W., Sudek, S. & Haygood, M.G. Genetic evidence supports secondary metabolic diversity in Prochloron spp., the cyanobacterial symbiont of a tropical ascidian. J. Nat. Prod. 67, 1341–1345 (2004).
Degnan, B.M. et al. New cyclic peptides with cytotoxic activity from the ascidian Lissoclinum patella. J. Med. Chem. 32, 1349–1354 (1989).
Fu, X., Do, T., Schmitz, F.J., Andrusevich, V. & Engel, M.H. New cyclic peptides from the ascidian Lissoclinum patella. J. Nat. Prod. 61, 1547–1551 (1998).
Sesin, D.F., Simon, J.G. & Ireland, C.M. The chemistry of Lissoclinum patella. Bull. Soc. Chim. Belg. 95, 853–867 (1986).
Schmidt, E.W. et al. Patellamide A and C biosynthesis by a microcin-like pathway in Prochloron didemni, the cyanobacterial symbiont of Lissoclinum patella. Proc. Natl. Acad. Sci. USA 102, 7315–7320 (2005).
Newman, D.J., Cragg, G.M. & Snader, K.M. Natural products as sources of new drugs over the period 1981–2002. J. Nat. Prod. 66, 1022–1037 (2003).
Floss, H.G. Combinatorial biosynthesis–potential and problems. J. Biotechnol. 124, 242–257 (2006).
Walsh, C.T. Combinatorial biosynthesis of antibiotics: challenges and opportunities. ChemBioChem 3, 125–134 (2002).
Xie, L. et al. Lacticin 481: in vitro reconstitution of lantibiotic synthetase activity. Science 303, 679–682 (2004).
Austin, M.B., Bowman, M.E., Ferrer, J.-L., Schröder, J. & Noel, J.P. An aldol switch discovered in stilbene synthases mediates cyclization specificity of type III polyketide synthases. Chem. Biol. 11, 1179–1194 (2004).
Schmidt, E.W., Obraztsova, A.Y., Davidson, S.K., Faulkner, D.J. & Haygood, M.G. Identification of the antifungal peptide-containing symbiont of the marine sponge Theonella swinhoei as a novel delta-proteobacterium, Candidatus entotheonella palauensis. Mar. Biol. 136, 969–977 (2000).
Piel, J. A polyketide synthase-peptide synthetase gene cluster from an uncultured bacterial symbiont of Paederus beetles. Proc. Natl. Acad. Sci. USA 99, 14002–14007 (2002).
Baumann, P. Biology of bacteriocyte-associated endosymbionts of plant sap-sucking insects. Annu. Rev. Microbiol. 59, 155–189 (2005).
Partida-Martinez, L.P. & Hertweck, C. Pathogenic fungus harbours endosymbiotic bacteria for toxin production. Nature 437, 884–888 (2005).
Withers, N., Vidaver, W. & Lewin, R.A. Pigment composition, photosynthesis and fine-structure of a non-blue-green prokaryotic algal symbiont (Prochloron sp.) in a didemnid ascidian from Hawaiian waters. Phycologia 17, 167–171 (1978).
Lewin, R.A. & Cheng, L. (eds.) Prochloron: a Microbial Enigma (Chapman and Hall, New York, 1989).
Burns, B.P., Goh, F., Allen, M. & Neilan, B.A. Microbial diversity of extant stromatolites in the hypersaline marine environment of Shark Bay, Australia. Environ. Microbiol. 6, 1096–1101 (2004).
Long, P.F., Dunlap, W.C., Battershill, C.N. & Jaspars, M. Shotgun cloning and heterologous expression of the patellamide gene cluster as a strategy to achieve sustained metabolite production. ChemBioChem 6, 1760–1765 (2005).
Yokobori, S., Kurabayashi, A., Neilan, B.A., Maruyama, T. & Hirose, E. Multiple origins of the ascidian-Prochloron symbiosis: molecular phylogeny of photosymbiotic and non-symbiotic colonial ascidians inferred from 18S rDNA sequences. Mol. Phylogenet. Evol. 40, 8–19 (2006).
Lewin, R.A. & Withers, N.W. Extraordinary pigment composition of a prokaryotic alga. Nature 256, 735–737 (1975).
Lewin, R.A. Prochlorophyta as a proposed new division of algae. Nature 261, 697–698 (1976).
Tomitani, A. et al. Chlorophyll b and phycobilins in the common ancestor of cyanobacteria and chloroplasts. Nature 400, 159–162 (1999).
Criss, A.K., Kline, K.A. & Seifert, H.S. The frequency and rate of pilin antigenic variation in Neisseria gonorrhoeae. Mol. Microbiol. 58, 510–519 (2005).
Bendich, A.J. & Drlica, K. Prokaryotic and eukaryotic chromosomes: what's the difference? Bioessays 22, 481–486 (2000).
Binder, B.J. & Chisholm, S.W. Relationship between DNA cycle and growth rate in Synechococcus sp. strain PCC 6301. J. Bacteriol. 172, 2313–2319 (1990).
Scarborough, R.M. Development of eptifibatide. Am. Heart J. 138, 1093–1104 (1999).
Litman, G.W., Cannon, J.P. & Dishaw, L.J. Reconstructing immune phylogeny: new perspectives. Nat. Rev. Immunol. 5, 866–879 (2005).
Espiritu, D.J. et al. Venomous cone snails: molecular phylogeny and the generation of toxin diversity. Toxicon 39, 1899–1916 (2001).
Duda, T.F. & Palumbi, S.R. Molecular genetics of ecological diversification: duplication and rapid evolution of toxin genes of the venomous gastropod Conus. Proc. Natl. Acad. Sci. USA 96, 6820–6823 (1999).
Schroeder, F.C. et al. Polyazamacrolides from ladybird beetles: ring-size selective oligomerization. Proc. Natl. Acad. Sci. USA 95, 13387–13391 (1998).
Sudek, S., Haygood, M.G., Youssef, D.T. & Schmidt, E.W. Trichamide, a cyclic peptide from the bloom-forming cyanobacterium Trichodesmium erythraeum predicted from the genome sequence. Appl. Environ. Microbiol. 72, 4382–4387 (2006).
Williams, A.B. & Jacobs, R.S. A marine natural product, patellamide D, reverses multidrug resistance in a human leukemic cell line. Cancer Lett. 71, 97–102 (1993).
Morita, M., Shibuya, M., Kushiro, T., Masuda, K. & Ebizuka, Y. Molecular cloning and functional expression of triterpene synthases from pea (Pisum sativum). Eur. J. Biochem. 267, 3453–3460 (2000).
Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F. & Higgins, D.G. The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882 (1997).
Salomon, C.E. & Faulkner, D.J. Localization studies of bioactive cyclic peptides in the ascidian Lissoclinum patella. J. Nat. Prod. 65, 689–692 (2002).
Ireland, C.M., Durso, A.R., Newman, R.A. & Hacker, M.P. Antineoplastic cyclic peptides from the marine tunicate Lissoclinum patella. J. Org. Chem. 47, 1807–1811 (1982).
Acknowledgements
This work was supported by grants from the US National Science Foundation (EF-0412226) and the US National Institutes of Health (R01 GM071425-01A1), and by a Willard L. Eccles Fellowship to B.J.H. We thank D.J. Faulkner (University of California San Diego), C. Ireland (University of Utah), L. Matainaho (University of Papua New Guinea) and the governments of the Republic of Palau and Papua New Guinea for the opportunities to collect the samples used in these studies. We also thank K. Rai and D. Jones (University of Utah) and L. Jiang (TIGR) for their help in the quantitative PCR experiments. T. Bugni (University of Utah) helped with mass measurements, and J. Sims (University of Utah) aided with sample processing.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Complete list of patE variants (patE1–patE29) identified by sequencing clone libraries. (PDF 167 kb)
Supplementary Fig. 2
Amino acid sequences of the compounds coded by the variants in Supplementary Figure 1. (PDF 151 kb)
Supplementary Fig. 3
An example of the SNPs observed in the raw Sequencher file. (PDF 185 kb)
Supplementary Fig. 4
Cladogram of prochloron strains derived from maximum likelihood analysis. (PDF 107 kb)
Supplementary Table 1
Data from the best-characterized 11 samples from this study. (PDF 88 kb)
Supplementary Table 2
Data from other samples examined in this study. (PDF 83 kb)
Supplementary Methods
Detailed methods, NMR and mass spectra, and tables of primers used in the study. (PDF 1370 kb)
Rights and permissions
About this article
Cite this article
Donia, M., Hathaway, B., Sudek, S. et al. Natural combinatorial peptide libraries in cyanobacterial symbionts of marine ascidians. Nat Chem Biol 2, 729–735 (2006). https://doi.org/10.1038/nchembio829
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio829
This article is cited by
-
Core-dependent post-translational modifications guide the biosynthesis of a new class of hypermodified peptides
Nature Communications (2023)
-
Isolation and identification of marine microbial products
Journal of Genetic Engineering and Biotechnology (2021)
-
Roussoelins A and B: two phenols with antioxidant capacity from ascidian-derived fungus Roussoella siamensis SYSU-MS4723
Marine Life Science & Technology (2021)
-
Ascidian-associated photosymbionts from Manado, Indonesia: secondary metabolites, bioactivity simulation, and biosynthetic insight
Symbiosis (2021)
-
Recent advances in the biosynthesis of RiPPs from multicore-containing precursor peptides
Journal of Industrial Microbiology and Biotechnology (2020)