Abstract
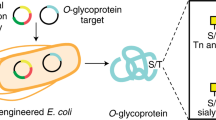
We describe a new method for producing homogeneous eukaryotic N-glycoproteins. The method involves the engineering and functional transfer of the Campylobacter jejuni glycosylation machinery in Escherichia coli to express glycosylated proteins with the key GlcNAc-Asn linkage. The bacterial glycans were then trimmed and remodeled in vitro by enzymatic transglycosylation to fulfill a eukaryotic N-glycosylation. It provides a potentially general platform for producing eukaryotic N-glycoproteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Change history
18 March 2010
In the version of this article initially published online, an extra blue square corresponding to a GlcNAc monomer was shown in the transglycosylation step in Figure 1 and the Graphical Abstract. Also, panels g and h of Figure 2 were mistakenly swapped. The errors have been corrected for the print, PDF and HTML versions of this article.
References
Dwek, R.A. Chem. Rev. 96, 683–720 (1996).
Helenius, A. & Aebi, M. Science 291, 2364–2369 (2001).
Gamblin, D.P., Scanlan, E.M. & Davis, B.G. Chem. Rev. 109, 131–163 (2009).
Rich, J.R. & Withers, S.G. Nat. Chem. Biol. 5, 206–215 (2009).
Bennett, C.S. & Wong, C.H. Chem. Soc. Rev. 36, 1227–1238 (2007).
Szymanski, C.M., Yao, R., Ewing, C.P., Trust, T.J. & Guerry, P. Mol. Microbiol. 32, 1022–1030 (1999).
Young, N.M. et al. J. Biol. Chem. 277, 42530–42539 (2002).
Wacker, M. et al. Science 298, 1790–1793 (2002).
Kowarik, M. et al. Science 314, 1148–1150 (2006).
Li, B., Zeng, Y., Hauser, S., Song, H. & Wang, L.X. J. Am. Chem. Soc. 127, 9692–9693 (2005).
Li, B., Song, H., Hauser, S. & Wang, L.X. Org. Lett. 8, 3081–3084 (2006).
Ochiai, H., Huang, W. & Wang, L.X. J. Am. Chem. Soc. 130, 13790–13803 (2008).
Wei, Y. et al. Biochemistry 47, 10294–10304 (2008).
Umekawa, M. et al. J. Biol. Chem. 283, 4469–4479 (2008).
Huang, W. et al. J. Am. Chem. Soc. 131, 2214–2223 (2009).
Feldman, M.F. et al. Proc. Natl. Acad. Sci. USA 102, 3016–3021 (2005).
Wacker, M. et al. Proc. Natl. Acad. Sci. USA 103, 7088–7093 (2006).
Alaimo, C. et al. EMBO J. 25, 967–976 (2006).
Lehrer, J., Vigeant, K.A., Tatar, L.D. & Valvano, M.A. J. Bacteriol. 189, 2618–2628 (2007).
Kowarik, M. et al. EMBO J. 25, 1957–1966 (2006).
Troutman, J.M. & Imperiali, B. Biochemistry 48, 2807–2816 (2009).
Jefferis, R. Biotechnol. Prog. 21, 11–16 (2005).
Anthony, R.M. et al. Science 320, 373–376 (2008).
Villa, A. et al. Int. J. Cancer 122, 2405–2413 (2008).
Holliger, P. & Hudson, P.J. Nat. Biotechnol. 23, 1126–1136 (2005).
Acknowledgements
We thank members of the Aebi and Wang labs for fruitful discussions. This work was supported in part by the Swiss National Science Foundation (grant 3100AQ-105541 to M.A.), the Eidgenössische Technische Hochschule Zurich and the US National Institutes of Health (grant GM080374 to L.-X.W.). We thank K. Takegawa (Kyushu University) for providing the pGEX-2T/Endo-A plasmid and K. Yamamoto (Kyoto University) for providing the pET23b-Endo-M plasmid, which were used to express the endo enzymes. F.S. and C. Lizak are members of the Zurich PhD Program in Molecular Life Sciences.
Author information
Authors and Affiliations
Contributions
F.S. engineered the glycosylation pathway, characterized proteins and wrote the manuscript. W.H. trimmed and remodeled glycans and characterized products. C. Li remodeled glycans and characterized products. B.L.S. performed preliminary MS analyses of glycosylated AcrA. C. Lizak engineered the CH2 domain. A.P. characterized the F8 protein. S.N. conceived the original idea and performed preliminary experiments. D.N. developed F8 and supervised the research. M.A. supervised the research and wrote the manuscript. L.-X.W. conceived the original idea, supervised the research and wrote the manuscript. All authors contributed to editing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.N. is a co-founder and shareholder of Philogen, the company that owns the F8 antibody.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Results, and Supplementary Figures 1–9 (PDF 645 kb)
Rights and permissions
About this article
Cite this article
Schwarz, F., Huang, W., Li, C. et al. A combined method for producing homogeneous glycoproteins with eukaryotic N-glycosylation. Nat Chem Biol 6, 264–266 (2010). https://doi.org/10.1038/nchembio.314
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.314
This article is cited by
-
Genetic and process engineering strategies for enhanced recombinant N-glycoprotein production in bacteria
Microbial Cell Factories (2021)
-
Engineering the flagellar type III secretion system: improving capacity for secretion of recombinant protein
Microbial Cell Factories (2019)
-
Legionella effector SetA as a general O-glucosyltransferase for eukaryotic proteins
Nature Chemical Biology (2019)
-
Single-pot glycoprotein biosynthesis using a cell-free transcription-translation system enriched with glycosylation machinery
Nature Communications (2018)