Abstract
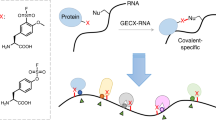
Excluding the ribosome and riboswitches, developing small molecules that selectively target RNA is a longstanding problem in chemical biology. A typical cellular RNA is difficult to target because it has little tertiary, but abundant secondary structure. We designed allele-selective compounds that target such an RNA, the toxic noncoding repeat expansion (r(CUG)exp) that causes myotonic dystrophy type 1 (DM1). We developed several strategies to generate allele-selective small molecules, including non-covalent binding, covalent binding, cleavage and on-site probe synthesis. Covalent binding and cleavage enabled target profiling in cells derived from individuals with DM1, showing precise recognition of r(CUG)exp. In the on-site probe synthesis approach, small molecules bound adjacent sites in r(CUG)exp and reacted to afford picomolar inhibitors via a proximity-based click reaction only in DM1-affected cells. We expanded this approach to image r(CUG)exp in its natural context.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Schlünzen, F. et al. Structural basis for the interaction of antibiotics with the peptidyl transferase center in eubacteria. Nature 413, 814–821 (2001).
Carter, A.P. et al. Functional insights from the structure of the 30S ribosomal subunit and its interactions with antibiotics. Nature 407, 340–348 (2000).
Blount, K.F. et al. Novel riboswitch-binding flavin analog that protects mice against Clostridium difficile infection without inhibiting cecal flora. Antimicrob. Agents Chemother. 59, 5736–5746 (2015).
Howe, J.A. et al. Selective small-molecule inhibition of an RNA structural element. Nature 526, 672–677 (2015).
Brook, J.D. et al. Molecular basis of myotonic dystrophy: expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a protein kinase family member. Cell 68, 799–808 (1992).
Jiang, H., Mankodi, A., Swanson, M.S., Moxley, R.T. & Thornton, C.A. Myotonic dystrophy type 1 is associated with nuclear foci of mutant RNA, sequestration of muscleblind proteins and deregulated alternative splicing in neurons. Hum. Mol. Genet. 13, 3079–3088 (2004).
Taneja, K.L., McCurrach, M., Schalling, M., Housman, D. & Singer, R.H. Foci of trinucleotide repeat transcripts in nuclei of myotonic dystrophy cells and tissues. J. Cell Biol. 128, 995–1002 (1995).
Furling, D., Lemieux, D., Taneja, K. & Puymirat, J. Decreased levels of myotonic dystrophy protein kinase (DMPK) and delayed differentiation in human myotonic dystrophy myoblasts. Neuromuscul. Disord. 11, 728–735 (2001).
Hamshere, M.G., Newman, E.E., Alwazzan, M., Athwal, B.S. & Brook, J.D. Transcriptional abnormality in myotonic dystrophy affects DMPK but not neighboring genes. Proc. Natl. Acad. Sci. USA 94, 7394–7399 (1997).
Johnson, L.F., Abelson, H.T., Penman, S. & Green, H. The relative amounts of the cytoplasmic RNA species in normal, transformed and senescent cultured cell lines. J. Cell. Physiol. 90, 465–470 (1977).
Velagapudi, S.P., Gallo, S.M. & Disney, M.D. Sequence-based design of bioactive small molecules that target precursor microRNAs. Nat. Chem. Biol. 10, 291–297 (2014).
Disney, M.D. et al. Inforna 2.0: a platform for the sequence-based design of small molecules targeting structured RNAs. ACS Chem. Biol. 11, 1720–1728 (2016).
Pushechnikov, A. et al. Rational design of ligands targeting triplet repeating transcripts that cause RNA dominant disease: application to myotonic muscular dystrophy type 1 and spinocerebellar ataxia type 3. J. Am. Chem. Soc. 131, 9767–9779 (2009).
Lee, M.M., Pushechnikov, A. & Disney, M.D. Rational and modular design of potent ligands targeting the RNA that causes myotonic dystrophy 2. ACS Chem. Biol. 4, 345–355 (2009).
Childs-Disney, J.L., Hoskins, J., Rzuczek, S.G., Thornton, C.A. & Disney, M.D. Rationally designed small molecules targeting the RNA that causes myotonic dystrophy type 1 are potently bioactive. ACS Chem. Biol. 7, 856–862 (2012).
Velagapudi, S.P. et al. Design of a small molecule against an oncogenic noncoding RNA. Proc. Natl. Acad. Sci. USA 113, 5898–5903 (2016).
Rzuczek, S.G. et al. Features of modularly assembled compounds that impart bioactivity against an RNA target. ACS Chem. Biol. 8, 2312–2321 (2013).
Childs-Disney, J.L. et al. Induction and reversal of myotonic dystrophy type 1 pre-mRNA splicing defects by small molecules. Nat. Commun. 4, 2044 (2013).
Wojtkowiak-Szlachcic, A. et al. Short antisense-locked nucleic acids (all-LNAs) correct alternative splicing abnormalities in myotonic dystrophy. Nucleic Acids Res. 43, 3318–3331 (2015).
Mastroyiannopoulos, N.P., Feldman, M.L., Uney, J.B., Mahadevan, M.S. & Phylactou, L.A. Woodchuck post-transcriptional element induces nuclear export of myotonic dystrophy 3′ untranslated region transcripts. EMBO Rep. 6, 458–463 (2005).
Guan, L. & Disney, M.D. Covalent small-molecule-RNA complex formation enables cellular profiling of small-molecule-RNA interactions. Angew. Chem. Int. Edn Engl. 52, 10010–10013 (2013).
Yang, W.Y., Wilson, H.D., Velagapudi, S.P. & Disney, M.D. Inhibition of non-ATG translational events in cells via covalent small molecules targeting RNA. J. Am. Chem. Soc. 137, 5336–5345 (2015).
Carter, B.J. et al. Site-specific cleavage of RNA by Fe(II)-bleomycin. Proc. Natl. Acad. Sci. USA 87, 9373–9377 (1990).
Lewis, W.G. et al. Click chemistry in situ: acetylcholinesterase as a reaction vessel for the selective assembly of a femtomolar inhibitor from an array of building blocks. Angew. Chem. Int. Edn Engl. 41, 1053–1057 (2002).
Rzuczek, S.G., Park, H. & Disney, M.D. A toxic RNA catalyzes the in cellulo synthesis of its own inhibitor. Angew. Chem. Int. Edn Engl. 53, 10956–10959 (2014).
Wang, E.T. et al. Transcriptome-wide regulation of pre-mRNA splicing and mRNA localization by muscleblind proteins. Cell 150, 710–724 (2012).
Wheeler, T.M. et al. Reversal of RNA dominance by displacement of protein sequestered on triplet repeat RNA. Science 325, 336–339 (2009).
Lima, W.F., Monia, B.P., Ecker, D.J. & Freier, S.M. Implication of RNA structure on antisense oligonucleotide hybridization kinetics. Biochemistry 31, 12055–12061 (1992).
Lakowicz, J.R. Principles of Fluorescence Spectroscopy (Springer, 2006).
Paige, J.S., Wu, K.Y. & Jaffrey, S.R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Strack, R.L., Disney, M.D. & Jaffrey, S.R. A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat-containing RNA. Nat. Methods 10, 1219–1224 (2013).
Bertrand, E. et al. Localization of ASH1 mRNA particles in living yeast. Mol. Cell 2, 437–445 (1998).
Baker, M. RNA imaging in situ. Nat. Methods 9, 787–790 (2012).
Chen, C.Z. et al. Two high-throughput screening assays for aberrant RNA-protein interactions in myotonic dystrophy type 1. Anal. Bioanal. Chem. 402, 1889–1898 (2012).
François, V. et al. Selective silencing of mutated mRNAs in DM1 by using modified hU7-snRNAs. Nat. Struct. Mol. Biol. 18, 85–87 (2011).
Shapiro, I.M. et al. An EMT-driven alternative splicing program occurs in human breast cancer and modulates cellular phenotype. PLoS Genet. 7, e1002218 (2011).
Warf, M.B., Nakamori, M., Matthys, C.M., Thornton, C.A. & Berglund, J.A. Pentamidine reverses the splicing defects associated with myotonic dystrophy. Proc. Natl. Acad. Sci. USA 106, 18551–18556 (2009).
Holt, I. et al. Defective mRNA in myotonic dystrophy accumulates at the periphery of nuclear splicing speckles. Genes Cells 12, 1035–1048 (2007).
Hampf, M. & Gossen, M. A protocol for combined Photinus and Renilla luciferase quantification compatible with protein assays. Anal. Biochem. 356, 94–99 (2006).
Acknowledgements
We thank T. Kodadek, G. Joyce, W. Ja, J. Childs-Disney, K. Sobczak and J. Cleveland for advice and critical review of the manuscript, and M.D.D. acknowledges J. and H. (nee McDougall) Disney. We also thank the platform for immortalization of human cells from the Institut de Myologie. This work was funded by the US National Institutes of Health (grants DP1NS096898 to M.D.D. and DP1NS096787 to R.Y.) and the Muscular Dystrophy Association (grant 380467 to M.D.D.). S.G.R. was partially supported by a postdoctoral fellowship from the Myotonic Dystrophy Foundation.
Author information
Authors and Affiliations
Contributions
M.D.D. directed the study, conceived of the ideas and designed experiments. S.G.R. designed experiments, synthesized all of the compounds and conducted all of the biochemical and cellular studies. Y.N. contributed to the synthesis of compounds. L.A.C. performed the FRET imaging. R.Y. contributed to the FRET studies. M.D.C. contributed to compound stability studies. D.F. provided critical reagents.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–14 and Supplementary Tables 1 and 2. (PDF 3349 kb)
Supplementary Note
Synthetic Procedures. (PDF 3737 kb)
Rights and permissions
About this article
Cite this article
Rzuczek, S., Colgan, L., Nakai, Y. et al. Precise small-molecule recognition of a toxic CUG RNA repeat expansion. Nat Chem Biol 13, 188–193 (2017). https://doi.org/10.1038/nchembio.2251
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2251
This article is cited by
-
Amplifying gene expression with RNA-targeted therapeutics
Nature Reviews Drug Discovery (2023)
-
Drugging RNA
Nature Biotechnology (2023)
-
Establishment of quantitative and consistent in vitro skeletal muscle pathological models of myotonic dystrophy type 1 using patient-derived iPSCs
Scientific Reports (2023)
-
Pervasive transcriptome interactions of protein-targeted drugs
Nature Chemistry (2023)
-
Targeting RNA structures with small molecules
Nature Reviews Drug Discovery (2022)