Abstract
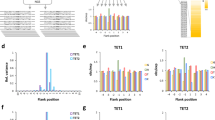
Ten-eleven translocation (TET) enzymes catalyze stepwise oxidation of 5-methylcytosine (mC) to yield 5-hydroxymethylcytosine (hmC) and the rarer bases 5-formylcytosine (fC) and 5-carboxylcytosine (caC). Stepwise oxidation obscures how each individual base forms and functions in epigenetic regulation, and prompts the question of whether TET enzymes primarily serve to generate hmC or are adapted to produce fC and caC as well. By mutating a single, conserved active site residue in human TET2, Thr1372, we uncovered enzyme variants that permit oxidation to hmC but largely eliminate fC and caC. Biochemical analyses, combined with molecular dynamics simulations, elucidated an active site scaffold that is required for wild-type (WT) stepwise oxidation and that, when perturbed, explains the mutants' hmC-stalling phenotype. Our results suggest that the TET2 active site is shaped to enable higher-order oxidation and provide the first TET variants that could be used to probe the biological functions of hmC separately from fC and caC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
Kriaucionis, S. & Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–930 (2009).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Pfaffeneder, T. et al. The discovery of 5-formylcytosine in embryonic stem cell DNA. Angew. Chem. Int. Edn Engl. 50, 7008–7012 (2011).
He, Y.F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
Kohli, R.M. & Zhang, Y. TET enzymes, TDG and the dynamics of DNA demethylation. Nature 502, 472–479 (2013).
Bachman, M. et al. 5-Hydroxymethylcytosine is a predominantly stable DNA modification. Nat. Chem. 6, 1049–1055 (2014).
Bachman, M. et al. 5-Formylcytosine can be a stable DNA modification in mammals. Nat. Chem. Biol. 11, 555–557 (2015).
Wu, H. & Zhang, Y. Charting oxidized methylcytosines at base resolution. Nat. Struct. Mol. Biol. 22, 656–661 (2015).
Liu, M.Y., DeNizio, J.E., Schutsky, E.K. & Kohli, R.M. The expanding scope and impact of epigenetic cytosine modifications. Curr. Opin. Chem. Biol. 33, 67–73 (2016).
Zheng, G., Fu, Y. & He, C. Nucleic acid oxidation in DNA damage repair and epigenetics. Chem. Rev. 114, 4602–4620 (2014).
Hu, L. et al. Structural insight into substrate preference for TET-mediated oxidation. Nature 527, 118–122 (2015).
Hu, L. et al. Crystal structure of TET2-DNA complex: insight into TET-mediated 5mC oxidation. Cell 155, 1545–1555 (2013).
Lu, J. et al. A computational investigation on the substrate preference of ten-eleven-translocation 2 (TET2). Phys. Chem. Chem. Phys. 18, 4728–4738 (2016).
Maiti, A. & Drohat, A.C. Thymine DNA glycosylase can rapidly excise 5-formylcytosine and 5-carboxylcytosine: potential implications for active demethylation of CpG sites. J. Biol. Chem. 286, 35334–35338 (2011).
Weber, A.R. et al. Biochemical reconstitution of TET1-TDG-BER-dependent active DNA demethylation reveals a highly coordinated mechanism. Nat. Commun. 7, 10806 (2016).
Iurlaro, M. et al. A screen for hydroxymethylcytosine and formylcytosine binding proteins suggests functions in transcription and chromatin regulation. Genome Biol. 14, R119 (2013).
Spruijt, C.G. et al. Dynamic readers for 5-(hydroxy)methylcytosine and its oxidized derivatives. Cell 152, 1146–1159 (2013).
Crawford, D.J. et al. Tet2 catalyzes stepwise 5-methylcytosine oxidation by an iterative and de novo mechanism. J. Am. Chem. Soc. 138, 730–733 (2016).
Fang, D., Lord, R.L. & Cisneros, G.A. Ab initio QM/MM calculations show an intersystem crossing in the hydrogen abstraction step in dealkylation catalyzed by AlkB. J. Phys. Chem. B 117, 6410–6420 (2013).
Fang, D. & Cisneros, G.A. Alternative pathway for the reaction catalyzed by DNA dealkylase AlkB from ab initio QM/MM calculations. J. Chem. Theory Comput. 10, 5136–5148 (2014).
Hashimoto, H. et al. Structure of Naegleria Tet-like dioxygenase (NgTet1) in complexes with a reaction intermediate 5-hydroxymethylcytosine DNA. Nucleic Acids Res. 43, 10713–10721 (2015).
Abdel-Wahab, O. et al. Genetic characterization of TET1, TET2, and TET3 alterations in myeloid malignancies. Blood 114, 144–147 (2009).
Scourzic, L., Mouly, E. & Bernard, O.A. TET proteins and the control of cytosine demethylation in cancer. Genome Med. 7, 9 (2015).
Cliffe, L.J. et al. JBP1 and JBP2 are two distinct thymidine hydroxylases involved in J biosynthesis in genomic DNA of African trypanosomes. Nucleic Acids Res. 37, 1452–1462 (2009).
Bullard, W., Lopes da Rosa-Spiegler, J., Liu, S., Wang, Y. & Sabatini, R. Identification of the glucosyltransferase that converts hydroxymethyluracil to base J in the trypanosomatid genome. J. Biol. Chem. 289, 20273–20282 (2014).
Pei, J., Kim, B.H. & Grishin, N.V. PROMALS3D: a tool for multiple protein sequence and structure alignments. Nucleic Acids Res. 36, 2295–2300 (2008).
Hu, X. et al. Tet and TDG mediate DNA demethylation essential for mesenchymal-to-epithelial transition in somatic cell reprogramming. Cell Stem Cell 14, 512–522 (2014).
Lian, C.G. et al. Loss of 5-hydroxymethylcytosine is an epigenetic hallmark of melanoma. Cell 150, 1135–1146 (2012).
Neri, F. et al. TET1 is a tumour suppressor that inhibits colon cancer growth by derepressing inhibitors of the WNT pathway. Oncogene 34, 4168–4176 (2015).
Gu, T.P. et al. The role of Tet3 DNA dioxygenase in epigenetic reprogramming by oocytes. Nature 477, 606–610 (2011).
Liu, M.Y., DeNizio, J.E. & Kohli, R.M. Quantification of oxidized 5-methylcytosine bases and TET enzyme activity. Methods Enzymol. 573, 365–385 (2016).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Schafmeister, C.E.A.F., Ross, W.S. & Romanovski, V. The Leap Module of AMBER (School of Pharmacy, University of California, San Francisco, 1995).
Case, D.A. et al. The Amber biomolecular simulation programs. J. Comput. Chem. 26, 1668–1688 (2005).
Jorgensen, W.L., Chandrasekhar, J., Madura, J.D., Impey, R.W. & Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926 (1983).
Dolinsky, T.J., Nielsen, J.E., McCammon, J.A. & Baker, N.A. PDB2PQR: an automated pipeline for the setup of Poisson-Boltzmann electrostatics calculations. Nucleic Acids Res. 32, W665–W667 (2004).
Dolinsky, T.J. et al. PDB2PQR: expanding and upgrading automated preparation of biomolecular structures for molecular simulations. Nucleic Acids Res. 35, W522–W525 (2007).
Olsson, M.H., Søndergaard, C.R., Rostkowski, M. & Jensen, J.H. PROPKA3: Consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 7, 525–537 (2011).
Bradbrook, G.M. et al. X-Ray and molecular dynamics studies of concanavalin-A glucoside and mannoside complexes relating structure to thermodynamics of binding. J. Chem. Soc., Faraday Trans. 94, 1603–1611 (1998).
Oda, A., Yamaotsu, N. & Hirono, S. New AMBER force field parameters of heme iron for cytochrome P450s determined by quantum chemical calculations of simplified models. J. Comput. Chem. 26, 818–826 (2005).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Roe, D.R. & Cheatham, T.E. III. PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38, 27–28 (1996).
Johnson, E.R. et al. Revealing noncovalent interactions. J. Am. Chem. Soc. 132, 6498–6506 (2010).
Contreras-García, J. et al. NCIPLOT: a program for plotting non-covalent interaction regions. J. Chem. Theory Comput. 7, 625–632 (2011).
Graham, S.E., Syeda, F. & Cisneros, G.A. Computational prediction of residues involved in fidelity checking for DNA synthesis in DNA polymerase I. Biochemistry 51, 2569–2578 (2012).
Elias, A.A. & Cisneros, G.A. Computational study of putative residues involved in DNA synthesis fidelity checking in Thermus aquaticus DNA polymerase I. Adv. Protein Chem. Struct. Biol. 96, 39–75 (2014).
Dewage, S.W. & Cisneros, G.A. Computational analysis of ammonia transfer along two intramolecular tunnels in Staphylococcus aureus glutamine-dependent amidotransferase (GatCAB). J. Phys. Chem. B 119, 3669–3677 (2015).
Cui, Q. & Karplus, M. Catalysis and specificity in enzymes: a study of triosephosphate isomerase and comparison with methyl glyoxal synthase. Adv. Protein Chem. 66, 315–372 (2003).
Martí, S. et al. Preorganization and reorganization as related factors in enzyme catalysis: the chorismate mutase case. Chemistry 9, 984–991 (2003).
Senn, H.M., O'Hagan, D. & Thiel, W. Insight into enzymatic C-F bond formation from QM and QM/MM calculations. J. Am. Chem. Soc. 127, 13643–13655 (2005).
Cisneros, G.A. et al. Reaction mechanism of the epsilon subunit of E. coli DNA polymerase III: insights into active site metal coordination and catalytically significant residues. J. Am. Chem. Soc. 131, 1550–1556 (2009).
Maiti, A., Michelson, A.Z., Armwood, C.J., Lee, J.K. & Drohat, A.C. Divergent mechanisms for enzymatic excision of 5-formylcytosine and 5-carboxylcytosine from DNA. J. Am. Chem. Soc. 135, 15813–15822 (2013).
Morgan, M.T., Bennett, M.T. & Drohat, A.C. Excision of 5-halogenated uracils by human thymine DNA glycosylase. Robust activity for DNA contexts other than CpG. J. Biol. Chem. 282, 27578–27586 (2007).
Acknowledgements
We thank B. Niedziolka and the Wistar Institute Protein Expression Facility for help with protein expression in Sf9 cells and all members of our labs for insightful discussions. Computing time from Wayne State C&IT and additional mass spectrometry resources from I. Blair's lab are gratefully acknowledged. This work was supported by the Rita Allen Foundation Scholar Award to R.M.K. and by NIH grants (R01 GM110174 to B.A.G., R01 GM108583 to G.A.C., and F30 CA196097 to M.Y.L.).
Author information
Authors and Affiliations
Contributions
R.M.K., G.A.C., M.Y.L., and H.T. conceived the experiments. R.M.K., M.Y.L., D.J.C., J.E.D., X.-J.C., and B.A.G. were involved in the design and optimization of biochemical and cellular experiments, which were performed and analyzed by M.Y.L., J.E.D., and R.M.K. For MD simulations, G.A.C., H.T., M.Y.L., and R.M.K. were involved in the design of experiments, which were performed and analyzed by H.T. and G.A.C. The manuscript was written by M.Y.L., R.M.K., H.T., and G.A.C. and edited by all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–10 and Supplementary Tables 1–9. (PDF 2502 kb)
Supplementary Dataset 1
Force field parameters. (TAR 81 kb)
Rights and permissions
About this article
Cite this article
Liu, M., Torabifard, H., Crawford, D. et al. Mutations along a TET2 active site scaffold stall oxidation at 5-hydroxymethylcytosine. Nat Chem Biol 13, 181–187 (2017). https://doi.org/10.1038/nchembio.2250
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2250
This article is cited by
-
The level of active DNA demethylation compounds in leukocytes and urine samples as potential epigenetic biomarkers in breast cancer patients
Scientific Reports (2024)
-
Vitamin C boosts DNA demethylation in TET2 germline mutation carriers
Clinical Epigenetics (2023)
-
Simultaneous sequencing of genetic and epigenetic bases in DNA
Nature Biotechnology (2023)
-
TETology: Epigenetic Mastermind in Action
Applied Biochemistry and Biotechnology (2021)