Abstract
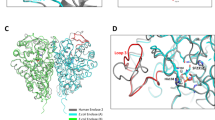
Despite being crucial for energy generation in most forms of life, few if any microbial antibiotics specifically inhibit glycolysis. To develop a specific inhibitor of the glycolytic enzyme enolase 2 (ENO2) for the treatment of cancers with deletion of ENO1 (encoding enolase 1), we modeled the synthetic tool compound inhibitor phosphonoacetohydroxamate (PhAH) into the active site of human ENO2. A ring-stabilized analog of PhAH, in which the hydroxamic nitrogen is linked to Cα by an ethylene bridge, was predicted to increase binding affinity by stabilizing the inhibitor in a bound conformation. Unexpectedly, a structure-based search revealed that our hypothesized backbone-stabilized PhAH bears strong similarity to SF2312, a phosphonate antibiotic of unknown mode of action produced by the actinomycete Micromonospora, which is active under anaerobic conditions. Here, we present multiple lines of evidence, including a novel X-ray structure, that SF2312 is a highly potent, low-nanomolar inhibitor of enolase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Fothergill-Gilmore, L.A. & Michels, P.A. Evolution of glycolysis. Prog. Biophys. Mol. Biol. 59, 105–235 (1993).
Deutscher, D., Meilijson, I., Kupiec, M. & Ruppin, E. Multiple knockout analysis of genetic robustness in the yeast metabolic network. Nat. Genet. 38, 993–998 (2006).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Vander Heiden, M.G., Cantley, L.C. & Thompson, C.B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Muller, F.L. et al. Passenger deletions generate therapeutic vulnerabilities in cancer. Nature 488, 337–342 (2012).
Poyner, R.R. & Reed, G.H. Structure of the bis divalent cation complex with phosphonoacetohydroxamate at the active site of enolase. Biochemistry 31, 7166–7173 (1992).
Anderson, V.E., Weiss, P.M. & Cleland, W.W. Reaction intermediate analogues for enolase. Biochemistry 23, 2779–2786 (1984).
Qin, J., Chai, G., Brewer, J.M., Lovelace, L.L. & Lebioda, L. Structures of asymmetric complexes of human neuron specific enolase with resolved substrate and product and an analogous complex with two inhibitors indicate subunit interaction and inhibitor cooperativity. J. Inorg. Biochem. 111, 187–194 (2012).
de AS Navarro, M.V. et al. Structural flexibility in Trypanosoma brucei enolase revealed by X-ray crystallography and molecular dynamics. FEBS J. 274, 5077–5089 (2007).
Wedekind, J.E., Poyner, R.R., Reed, G.H. & Rayment, I. Chelation of serine 39 to Mg2+ latches a gate at the active site of enolase: structure of the bis(Mg2+) complex of yeast enolase and the intermediate analog phosphonoacetohydroxamate at 2.1-Å resolution. Biochemistry 33, 9333–9342 (1994).
Zhang, E., Hatada, M., Brewer, J.M. & Lebioda, L. Catalytic metal ion binding in enolase: the crystal structure of an enolase-Mn2+-phosphonoacetohydroxamate complex at 2.4-Å resolution. Biochemistry 33, 6295–6300 (1994).
Watanabe, H., Yoshida, J., Tanaka, E., Ito, M., Miyadoh, S. & Shomura, T. Studies on a new phosphonic acid antibiotic, SF-2312. Scientific Reports of Meiji Seika Kaisha 25, 12–17 (1986).
Hanaya, T. & Itoh, C. An efficient synthesis of antibiotic SF-2312 (3-dihydroxyphosphoryl-1,5-dihydroxy-2-pyrrolidone). Heterocycles 82, 1675–1683 (2011).
Liu, Y., Jacobs, H.K. & Gopalan, A.S. A new approach to cyclic hydroxamic acids: intramolecular cyclization of N-benzyloxy carbamates with carbon nucleophiles. Tetrahedron 67, 2206–2214 (2011).
Martinez Molina, D. et al. Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science 341, 84–87 (2013).
Brewer, J.M. & Wampler, J.E. A differential scanning calorimetric study of the effects of metal ions, substrate/product, substrate analogues and chaotropic anions on the thermal denaturation of yeast enolase 1. Int. J. Biol. Macromol. 28, 213–218 (2001).
Zhang, E., Brewer, J.M., Minor, W., Carreira, L.A. & Lebioda, L. Mechanism of enolase: the crystal structure of asymmetric dimer enolase-2-phospho-D-glycerate/enolase-phosphoenolpyruvate at 2.0 Å resolution. Biochemistry 36, 12526–12534 (1997).
Marangos, P.J., Parma, A.M. & Goodwin, F.K. Functional properties of neuronal and glial isoenzymes of brain enolase. J. Neurochem. 31, 727–732 (1978).
Muller, F.L., Fletcher-Sananikone, E., Colla, S., Aquilanti, E. & DePinho, R. Collateral gene inactivation biomarkers and targets for cancer therapy. US patent WO2013090732A2 (2013).
Dette, G.A., Knothe, H., Schönenbach, B. & Plage, G. Comparative study of fosfomycin activity in Mueller–Hinton media and in tissues. J. Antimicrob. Chemother. 11, 517–524 (1983).
Horii, S. et al. Structure of alahopcin (nourseimycin), a new dipeptide antibiotic. J. Antibiot. (Tokyo) 38, 302–311 (1985).
Higashide, E., Kanamaru, T., Fukase, H. & Horii, S. Isolation of dealanylalahopcin, a new amino acid, and its biological activity. J. Antibiot. (Tokyo) 38, 296–301 (1985).
Gerlt, J.A., Babbitt, P.C. & Rayment, I. Divergent evolution in the enolase superfamily: the interplay of mechanism and specificity. Arch. Biochem. Biophys. 433, 59–70 (2005).
Brewer, J.M. Yeast enolase: mechanism of activation by metal ions. CRC Crit. Rev. Biochem. 11, 209–254 (1981).
Lee, J. & Kim, S.H. High-throughput T7 LIC vector for introducing C-terminal poly-histidine tags with variable lengths without extra sequences. Protein Expr. Purif. 63, 58–61 (2009).
Studier, F.W. Protein production by auto-induction in high-density shaking cultures. Protein Exp. Purif. 41, 207–234 (2005).
Battye, T.G., Kontogiannis, L., Johnson, O., Powell, H.R. & Leslie, A.G. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D Biol. Crystallogr. 67, 271–281 (2011).
Evans, P.R. & Murshudov, G.N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Chai, G. et al. Expression, purification and the 1.8 angstroms resolution crystal structure of human neuron specific enolase. J. Mol. Biol. 341, 1015–1021 (2004).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Duncan, C.G. et al. Integrated genomic analyses identify ERRFI1 and TACC3 as glioblastoma-targeted genes. Oncotarget 1, 265–277 (2010).
Stommel, J.M. et al. Coactivation of receptor tyrosine kinases affects the response of tumor cells to targeted therapies. Science 318, 287–290 (2007).
Bady, P. et al. DNA fingerprinting of glioma cell lines and considerations on similarity measurements. Neuro-oncol. 14, 701–711 (2012).
Torsvik, A. et al. U-251 revisited: genetic drift and phenotypic consequences of long-term cultures of glioblastoma cells. Cancer Med. 3, 812–824 (2014).
Yuan, M., Breitkopf, S.B., Yang, X. & Asara, J.M. A positive/negative ion-switching, targeted mass spectrometry-based metabolomics platform for bodily fluids, cells, and fresh and fixed tissue. Nat. Protoc. 7, 872–881 (2012).
Ying, H. et al. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 149, 656–670 (2012).
Acknowledgements
We thank J. Holton and G. Meigs for their assistance with X-ray diffraction data collection at the Advanced Light Source, beamline 8.3.1. We thank K. Kalhurachi and J. McMurray for assistance with 13C-NMR measurements. We thank S. Millward, J. Marszalek and Y. Sun for critical comments and suggestions. We thank M. Yuan and S. Breitkopf for help with MS experiments. We thank M. Protopopova for assistance with Operetta cell imaging. We thank L. Lebioda, G. Reed and R. Poyner for useful discussions about the catalytic mechanism of enolase. We thank R. Zielinski for assistance with hypoxia experiments. The cell line D423-MG (D423) was kindly shared by D. Bigner (Duke University Medical Center) while the Gli56 cell line was shared by D. Louis (Massachusetts General Hospital) and TS neurospheres were shared by C. Brennan (Memorial Sloan Kettering Cancer Center). Inhibitors were provided by the Pharmaceutical Chemistry Facility at MD Anderson, supported by the US National Institutes of Health (NIH) National Cancer Institute under award number P30CA016672. Financial support was provided by NIH-NCI grant 7P01CA095616-10 (R.A.D.); Cancer Prevention Research Institute of Texas (CPRIT) grant RP140612 (R.A.D.); NCI grants 5P01CA120964 (J.M.A.) and 5P30CA006516 (J.M.A.) and NIH Career Development Project through the Specialized Programs of Research Excellence (SPORE) P50CA127001-07 (F.L.M.). F.L.M. was supported by Research Scholar Grant RSG-15-145-01-CDD from the American Cancer Society.
Author information
Authors and Affiliations
Contributions
D.M. performed structural modeling and docking; F.L.M. and D.M. conceived the cyclized inhibitors, which were synthesized by Z.P., D.S., B.A.B.P. and W.B.; P.G.L., G.R.L. and T.M.L. performed isolation of recombinant protein and X-ray crystallography; M.E.D.F., F.P., F.L.M. and B.C. repeated chemical syntheses and wrote synthetic procedures with characterizations. F.P. and F.L.M. performed chiral chromatography. F.L.M. and N.S. performed in vitro enzymatic activity experiments. N.S., N.H. and Y.-H.L. performed cell culture, western blots, 13C-NMR tracing and biochemical profiling experiments. N.S. performed thermal shift assays. J.M.A. performed MS small-molecule metabolite analysis. R.A.D., Y.A.W., F.L.M., N.S. and P.G.L. oversaw overall experimental design and data analysis and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–2 and Supplementary Figures 1–26. (PDF 5726 kb)
Supplementary Note 1
Synthetic procedures. (PDF 115 kb)
Source Data
Source data for Figures 1, 2, 4, and 5. (ZIP 142 kb)
Rights and permissions
About this article
Cite this article
Leonard, P., Satani, N., Maxwell, D. et al. SF2312 is a natural phosphonate inhibitor of enolase. Nat Chem Biol 12, 1053–1058 (2016). https://doi.org/10.1038/nchembio.2195
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2195
This article is cited by
-
ENO2-derived phosphoenolpyruvate functions as an endogenous inhibitor of HDAC1 and confers resistance to antiangiogenic therapy
Nature Metabolism (2023)
-
Metabolic collateral lethal target identification reveals MTHFD2 paralogue dependency in ovarian cancer
Nature Metabolism (2022)
-
Molecular docking of alpha-enolase to elucidate the promising candidates against Streptococcus pneumoniae infection
DARU Journal of Pharmaceutical Sciences (2021)
-
Robust detection of oncometabolic aberrations by 1H–13C heteronuclear single quantum correlation in intact biological specimens
Communications Biology (2020)
-
In silico prediction of a new lead compound targeting enolase of trypanosomatids through structure-based virtual screening and molecular dynamic studies
Journal of Molecular Modeling (2020)