Abstract
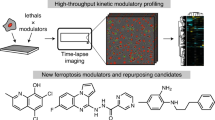
Apoptosis is one type of programmed cell death. Increasingly, non-apoptotic cell death is recognized as being genetically controlled, or 'regulated'. However, the full extent and diversity of alternative cell death mechanisms remain uncharted. Here we surveyed the landscape of pharmacologically accessible cell death mechanisms. In an examination of 56 caspase-independent lethal compounds, modulatory profiling showed that 10 compounds induced three different types of regulated non-apoptotic cell death. Optimization of one of those ten resulted in the discovery of FIN56, a specific inducer of ferroptosis. Ferroptosis has been found to occur when the lipid-repair enzyme GPX4 is inhibited. FIN56 promoted degradation of GPX4. FIN56 also bound to and activated squalene synthase, an enzyme involved in isoprenoid biosynthesis, independent of GPX4 degradation. These discoveries show that dysregulation of lipid metabolism is associated with ferroptosis. This systematic approach is a means to discover and characterize novel cell death phenotypes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Fuchs, Y. & Steller, H. Programmed cell death in animal development and disease. Cell 147, 742–758 (2011).
Aravind, L., Dixit, V.M. & Koonin, E.V. The domains of death: evolution of the apoptosis machinery. Trends Biochem. Sci. 24, 47–53 (1999).
Degterev, A. et al. Identification of RIP1 kinase as a specific cellular target of necrostatins. Nat. Chem. Biol. 4, 313–321 (2008).
Vanden Berghe, T., Linkermann, A., Jouan-Lanhouet, S., Walczak, H. & Vandenabeele, P. Regulated necrosis: the expanding network of non-apoptotic cell death pathways. Nat. Rev. Mol. Cell Biol. 15, 135–147 (2014).
Kaczmarek, A., Vandenabeele, P. & Krysko, D.V. Necroptosis: the release of damage-associated molecular patterns and its physiological relevance. Immunity 38, 209–223 (2013).
Kono, H. & Rock, K.L. How dying cells alert the immune system to danger. Nat. Rev. Immunol. 8, 279–289 (2008).
Skouta, R. et al. Ferrostatins inhibit oxidative lipid damage and cell death in diverse disease models. J. Am. Chem. Soc. 136, 4551–4556 (2014).
Linkermann, A. et al. Synchronized renal tubular cell death involves ferroptosis. Proc. Natl. Acad. Sci. USA 111, 16836–16841 (2014).
Dixon, S.J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Dixon, S.J. et al. Pharmacological inhibition of cystine-glutamate exchange induces endoplasmic reticulum stress and ferroptosis. eLife 3, e02523 (2014).
Hayano, M., Yang, W.S., Corn, C.K., Pagano, N.C. & Stockwell, B.R. Loss of cysteinyl-tRNA synthetase (CARS) induces the transsulfuration pathway and inhibits ferroptosis induced by cystine deprivation. Cell Death Differ. 23, 270–278 (2016).
Yang, W.S. et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 156, 317–331 (2014).
Imai, H. & Nakagawa, Y. Biological significance of phospholipid hydroperoxide glutathione peroxidase (PHGPx, GPx4) in mammalian cells. Free Radic. Biol. Med. 34, 145–169 (2003).
Linkermann, A., Stockwell, B.R., Krautwald, S. & Anders, H.-J. Regulated cell death and inflammation: an auto-amplification loop causes organ failure. Nat. Rev. Immunol. 14, 759–767 (2014).
Hitomi, J. et al. Identification of a molecular signaling network that regulates a cellular necrotic cell death pathway. Cell 135, 1311–1323 (2008).
Hahn, W.C. et al. Creation of human tumour cells with defined genetic elements. Nature 400, 464–468 (1999).
Wolpaw, A.J. et al. Modulatory profiling identifies mechanisms of small molecule-induced cell death. Proc. Natl. Acad. Sci. USA 108, E771–E780 (2011).
Degterev, A. et al. Chemical inhibitor of nonapoptotic cell death with therapeutic potential for ischemic brain injury. Nat. Chem. Biol. 1, 112–119 (2005).
Takahashi, N. et al. Necrostatin-1 analogues: critical issues on the specificity, activity and in vivo use in experimental disease models. Cell Death Dis. 3, e437 (2012).
Yang, W.S. & Stockwell, B.R. Synthetic lethal screening identifies compounds activating iron-dependent, nonapoptotic cell death in oncogenic-RAS-harboring cancer cells. Chem. Biol. 15, 234–245 (2008).
Shoemaker, R.H. The NCI60 human tumour cell line anticancer drug screen. Nat. Rev. Cancer 6, 813–823 (2006).
Shimada, K., Hayano, M., Pagano, N.C. & Stockwell, B.R. Cell-line selectivity improves the predictive power of pharmacogenomic analyses and helps identify NADPH as biomarker for ferroptosis sensitivity. Cell Chem. Biol. 23, 225–235 (2016).
Romanowska, M. et al. Effects of selenium supplementation on expression of glutathione peroxidase isoforms in cultured human lung adenocarcinoma cell lines. Lung Cancer 55, 35–42 (2007).
Fradejas, N. et al. Mammalian Trit1 is a tRNA([Ser]Sec)-isopentenyl transferase required for full selenoprotein expression. Biochem. J. 450, 427–432 (2013).
Song, J.H. et al. Deletion of Pim kinases elevates the cellular levels of reactive oxygen species and sensitizes to K-Ras-induced cell killing. Oncogene 34, 3728–3736 (2015).
Tansey, T.R. & Shechter, I. Structure and regulation of mammalian squalene synthase. Biochim. Biophys. Acta 1529, 49–62 (2000).
Liu, C.-I. et al. Acta Crystallogr. D Biol. Crystallogr. 70, 231–241 (2014).
Chugh, A., Ray, A. & Gupta, J.B. Squalene epoxidase as hypocholesterolemic drug target revisited. Prog. Lipid Res. 42, 37–50 (2003).
Gueven, N., Woolley, K. & Smith, J. Border between natural product and drug: comparison of the related benzoquinones idebenone and coenzyme Q10 . Redox Biol. 4, 289–295 (2015).
Dixon, S.J. et al. Human haploid cell genetics reveals roles for lipid metabolism genes in nonapoptotic cell death. ACS Chem. Biol. 10, 1604–1609 (2015).
Hirsch, H.A. et al. A transcriptional signature and common gene networks link cancer with lipid metabolism and diverse human diseases. Cancer Cell 17, 348–361 (2010).
Santos, C.R. & Schulze, A. Lipid metabolism in cancer. FEBS J. 279, 2610–2623 (2012).
Kamphorst, J.J., Fan, J., Lu, W., White, E. & Rabinowitz, J.D. Liquid chromatography-high resolution mass spectrometry analysis of fatty acid metabolism. Anal. Chem. 83, 9114–9122 (2011).
Yagoda, N. et al. RAS-RAF-MEK-dependent oxidative cell death involving voltage-dependent anion channels. Nature 447, 864–868 (2007).
Cholody, W.M. et al. Derivatives of fluorene, anthracene, xanthene, dibenzosuberone and acridine and uses thereof. US patent application PCT/US2008/006015 (2008).
Backman, T.W.H., Cao, Y. & Girke, T. ChemMine tools: an online service for analyzing and clustering small molecules. Nucleic Acids Res. 39, W486–W491 (2011).
Alegre-Aguarón, E. et al. Growth factor priming differentially modulates components of the extracellular matrix proteome in chondrocytes and synovium-derived stem cells. PLoS One 9, e88053 (2014).
Acknowledgements
We thank H. Li (Columbia University, New York, New York, USA) and C. Henderson (Biogen Idec, Cambridge, Massachusetts, USA) for providing mevalonic acid and farnesyl pyrophosphate, C.-I. Liu and A.H.J. Wang (Academia Sinica, Taipei, Taiwan) for the truncated human squalene synthase construct, R. Weinberg (Whitehead Institute, Cambridge, Massachusetts, USA) for engineered BJ cell lines (BJeLR, DRD, BJeHLT, and BJeH), E. Schon (Columbia University, New York, New York, USA) for 143B cells, E. Lee for assistance with ACC experiments, and V. Viswanathan for helpful discussions. This research was funded by the Howard Hughes Medical Institute, the US National Institutes of Health (grants 5R01CA097061, 5R01GM085081, and R01CA161061 to B.R.S.), New York Stem Cell Science (grant C026715 to B.R.S.) and the US National Cancer Institute (K99 Pathway to Independence Award 1K99CA166517-01 to S.J.D.).
Author information
Authors and Affiliations
Contributions
K.S. and B.R.S. conceived of the project, designed the experiments, analyzed the data, and wrote the manuscript. K.S. performed all experiments and analyses except the following: R.S. and C.A.V. synthesized CIL56 analogs; A.K. expressed and purified SQS protein and performed confirmatory SQS pulldown experiments and competition assays; W.S.Y. performed the GPX4 enzymatic assay; and M.H. performed confirmatory siRNA experiments. L.M.B. performed proteomic analysis for target identification. S.J.D. participated in characterization of TOFA's effect on FIN56. A.J.W. assisted in modulatory profiling experiments. S.J.D. assisted in writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–14 and Supplementary Tables 1–3. (PDF 5029 kb)
Supplementary Data Set 1
Lists of lethal compounds, chemical death modulators, cell lines, and modulatory profiles of characterized and uncharacterized lethal compounds. (XLSX 104 kb)
Supplementary Note 1
Supplementary Note 1 (PDF 2402 kb)
Supplementary Note 2
Synthetic Procedures (PDF 1066 kb)
Rights and permissions
About this article
Cite this article
Shimada, K., Skouta, R., Kaplan, A. et al. Global survey of cell death mechanisms reveals metabolic regulation of ferroptosis. Nat Chem Biol 12, 497–503 (2016). https://doi.org/10.1038/nchembio.2079
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2079
This article is cited by
-
Lipids as mediators of cancer progression and metastasis
Nature Cancer (2024)
-
Apomorphine is a potent inhibitor of ferroptosis independent of dopaminergic receptors
Scientific Reports (2024)
-
Current and Potential Roles of Ferroptosis in Bladder Cancer
Current Medical Science (2024)
-
Effect and Mechanism of LIN28 on Ferroptosis in Mg2+-free Rat Hippocampal Neuron Model of Epilepsy
Neurochemical Research (2024)
-
Ferroptosis: potential targets and emerging roles in pancreatic diseases
Archives of Toxicology (2024)