Abstract
Lytic polysaccharide monooxygenases (LPMOs) are copper-containing enzymes that oxidatively break down recalcitrant polysaccharides such as cellulose and chitin. Since their discovery, LPMOs have become integral factors in the industrial utilization of biomass, especially in the sustainable generation of cellulosic bioethanol. We report here a structural determination of an LPMO-oligosaccharide complex, yielding detailed insights into the mechanism of action of these enzymes. Using a combination of structure and electron paramagnetic resonance spectroscopy, we reveal the means by which LPMOs interact with saccharide substrates. We further uncover electronic and structural features of the enzyme active site, showing how LPMOs orchestrate the reaction of oxygen with polysaccharide chains.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Himmel, M.E. et al. Biomass recalcitrance: engineering plants and enzymes for biofuels production. Science 315, 804–807 (2007).
Sheldon, R.A. Green and sustainable manufacture of chemicals from biomass: state of the art. Green Chem. 16, 950–963 (2014).
Payne, C.M. et al. Fungal cellulases. Chem. Rev. 115, 1308–1448 (2015).
Beeson, W.T., Vu, V.V., Span, E.A., Phillips, C.M. & Marletta, M.A. Cellulose degradation by polysaccharide monooxygenases. Annu. Rev. Biochem. 84, 923–946 (2015).
Horn, S.J., Vaaje-Kolstad, G., Westereng, B. & Eijsink, V.G.H. Novel enzymes for the degradation of cellulose. Biotechnol. Biofuels 5, 45 (2012).
Müller, G., Várnai, A., Johansen, K.S., Eijsink, V.G.H. & Horn, S.J. Harnessing the potential of LPMO-containing cellulase cocktails poses new demands on processing conditions. Biotechnol. Biofuels 8, 187 (2015).
Hemsworth, G.R., Henrissat, B., Davies, G.J. & Walton, P.H. Discovery and characterization of a new family of lytic polysaccharide monooxygenases. Nat. Chem. Biol. 10, 122–126 (2014).
Lo Leggio, L. et al. Structure and boosting activity of a starch-degrading lytic polysaccharide monooxygenase. Nat. Commun. 6, 5961 (2015).
Cantarel, B.L. et al. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for glycogenomics. Nucleic Acids Res. 37, D233–D238 (2009).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P.M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–D495 (2014).
Merino, S.T. & Cherry, J. Progress and challenges in enzyme development for biomass utilization. Adv. Biochem. Eng. Biotechnol. 108, 95–120 (2007).
Harris, P.V. et al. Stimulation of lignocellulosic biomass hydrolysis by proteins of glycoside hydrolase family 61: structure and function of a large, enigmatic family. Biochemistry 49, 3305–3316 (2010).
Vaaje-Kolstad, G. et al. An oxidative enzyme boosting the enzymatic conversion of recalcitrant polysaccharides. Science 330, 219–222 (2010).
Forsberg, Z. et al. Cleavage of cellulose by a CBM33 protein. Protein Sci. 20, 1479–1483 (2011).
Quinlan, R.J. et al. Insights into the oxidative degradation of cellulose by a copper metalloenzyme that exploits biomass components. Proc. Natl. Acad. Sci. USA 108, 15079–15084 (2011).
Phillips, C.M., Beeson, W.T., Cate, J.H. & Marletta, M.A. Cellobiose dehydrogenase and a copper-dependent polysaccharide monooxygenase potentiate cellulose degradation by Neurospora crassa. ACS Chem. Biol. 6, 1399–1406 (2011).
Hemsworth, G.R., Davies, G.J. & Walton, P.H. Recent insights into copper-containing lytic polysaccharide mono-oxygenases. Curr. Opin. Struct. Biol. 23, 660–668 (2013).
Aachmann, F.L. et al. in Encyclopedia of Inorganic and Bioinorganic Chemistry (ed. Scott, R.A.) 111, 1–13 (Wiley, 2015).
Kjaergaard, C.H. et al. Spectroscopic and computational insight into the activation of O2 by the mononuclear Cu center in polysaccharide monooxygenases. Proc. Natl. Acad. Sci. USA 111, 8797–8802 (2014).
Kim, S., Ståhlberg, J., Sandgren, M., Paton, R.S. & Beckham, G.T. Quantum mechanical calculations suggest that lytic polysaccharide monooxygenases use a copper-oxyl, oxygen-rebound mechanism. Proc. Natl. Acad. Sci. USA 111, 149–154 (2014).
Gao, J., Thomas, D.A., Sohn, C.H. & Beauchamp, J.L. Biomimetic reagents for the selective free radical and acid-base chemistry of glycans: application to glycan structure determination by mass spectrometry. J. Am. Chem. Soc. 135, 10684–10692 (2013).
Eriksson, K.-E., Pettersson, B. & Westermark, U. Oxidation: an important enzyme reaction in fungal degradation of cellulose. FEBS Lett. 49, 282–285 (1974).
Hemsworth, G.R. et al. The copper active site of CBM33 polysaccharide oxygenases. J. Am. Chem. Soc. 135, 6069–6077 (2013).
Li, X., Beeson, W.T. IV, Phillips, C.M., Marletta, M.A. & Cate, J.H.D. Structural basis for substrate targeting and catalysis by fungal polysaccharide monooxygenases. Structure 20, 1051–1061 (2012).
Agger, J.W. et al. Discovery of LPMO activity on hemicelluloses shows the importance of oxidative processes in plant cell wall degradation. Proc. Natl. Acad. Sci. USA 111, 6287–6292 (2014).
Borisova, A.S. et al. Structural and functional characterization of a lytic polysaccharide monooxygenase with broad substrate specificity. J. Biol. Chem. 290, 22955–22969 (2015).
Bennati-Granier, C. et al. Substrate specificity and regioselectivity of fungal AA9 lytic polysaccharide monooxygenases secreted by Podospora anserina. Biotechnol. Biofuels 8, 90 (2015).
Loose, J.S.M., Forsberg, Z., Fraaije, M.W., Eijsink, V.G.H. & Vaaje-Kolstad, G. A rapid quantitative activity assay shows that the Vibrio cholerae colonization factor GbpA is an active lytic polysaccharide monooxygenase. FEBS Lett. 588, 3435–3440 (2014).
Gudmundsson, M. et al. Structural and electronic snapshots during the transition from a Cu(II) to Cu(I) metal center of a lytic polysaccharide monooxygenase by X-ray photoreduction. J. Biol. Chem. 289, 18782–18792 (2014).
Asensio, J.L., Ardá, A., Cañada, F.J. & Jiménez-Barbero, J. Carbohydrate-aromatic interactions. Acc. Chem. Res. 46, 946–954 (2013).
Nishio, M. The CH/π hydrogen bond in chemistry. Conformation, supramolecules, optical resolution and interactions involving carbohydrates. Phys. Chem. Chem. Phys. 13, 13873–13900 (2011).
Davies, G.J., Planas, A. & Rovira, C. Conformational analyses of the reaction coordinate of glycosidases. Acc. Chem. Res. 45, 308–316 (2012).
Lee, J.Y. & Karlin, K.D. Elaboration of copper-oxygen mediated C-H activation chemistry in consideration of future fuel and feedstock generation. Curr. Opin. Chem. Biol. 25, 184–193 (2015).
Iwaizumi, M., Kudo, T. & Kita, S. Correlation between the hyperfine coupling constants of donor nitrogens and the structures of the first coordination sphere in copper complexes as studied by nitrogen-14 ENDOR spectroscopy. Inorg. Chem. 25, 1546–1550 (1986).
Kim, D., Kim, N.H. & Kim, S.H. 34 GHz pulsed ENDOR characterization of the copper coordination of an amyloid-β peptide relevant to Alzheimer's disease. Angew. Chem. Int. Ed. Engl. 52, 1139–1142 (2013).
Aachmann, F.L., Sørlie, M., Skjåk-Bræk, G., Eijsink, V.G.H. & Vaaje-Kolstad, G. NMR structure of a lytic polysaccharide monooxygenase provides insight into copper binding, protein dynamics, and substrate interactions. Proc. Natl. Acad. Sci. USA 109, 18779–18784 (2012).
Karlin, K.D. & Itoh, S. Copper-Oxygen Chemistry (Wiley, 2011).
Donoghue, P.J. et al. Rapid C-H bond activation by a monocopper(III)-hydroxide complex. J. Am. Chem. Soc. 133, 17602–17605 (2011).
Wu, M. et al. Crystal structure and computational characterization of the lytic polysaccharide monooxygenase GH61D from the Basidiomycota fungus Phanerochaete chrysosporium. J. Biol. Chem. 288, 12828–12839 (2013).
Dhar, D. & Tolman, W.B. Hydrogen atom abstraction from hydrocarbons by a copper(III)-hydroxide complex. J. Am. Chem. Soc. 137, 1322–1329 (2015).
Singh, S.K. & Das, A. The n → π* interaction: a rapidly emerging non-covalent interaction. Phys. Chem. Chem. Phys. 17, 9596–9612 (2015).
Egli, M. & Gessner, R.V. Stereoelectronic effects of deoxyribose O4′ on DNA conformation. Proc. Natl. Acad. Sci. USA 92, 180–184 (1995).
Pavlakos, I. et al. Noncovalent lone pair···(No-π!)-heteroarene interactions: the Janus-faced hydroxy group. Angew. Chem. Int. Ed. Engl. 54, 8169–8174 (2015).
Roeser, D. et al. A general binding mechanism for all human sulfatases by the formylglycine-generating enzyme. Proc. Natl. Acad. Sci. USA 103, 81–86 (2006).
Spodsberg, N., Shaghasi, T., Sweeney, M., Xu, F. & Schnorr, K. Polypeptides having cellulolytic enhancing activity and polynucleotides encoding same. WO patent 2014066141 A3 (2014).
Liu, Y. et al. Use of a fluorescence plate reader for measuring kinetic parameters with inner filter effect correction. Anal. Biochem. 267, 331–335 (1999).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D Biol. Crystallogr. 66, 22–25 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Vagin, A.A. et al. REFMAC5 dictionary: organization of prior chemical knowledge and guidelines for its use. Acta Crystallogr. D Biol. Crystallogr. 60, 2184–2195 (2004).
Gemperle, C., Aebli, G., Schweiger, A. & Ernst, R.R. Phase cycling in pulse EPR. J. Magn. Reson. 88, 241–256 (1990).
Acknowledgements
We thank K. Rasmussen and R.M. Borup for experimental assistance, and MAXLAB, Sweden and the European Synchrotron Radiation Facility (ESRF), France, for synchrotron beam time and assistance. This work was supported by the UK Biotechnology and Biological Sciences Research Council (grant numbers BB/L000423 to P.D., G.J.D. and P.H.W., and BB/L021633/1 to G.J.D. and P.H.W.), Agence Française de l'Environnement et de la Maîtrise de l'Energie (grant number 1201C102 to B.H.), the Danish Council for Strategic Research (grant numbers 12-134923 to L.L.L. and 12-134922 to K.S.J.). Travel to synchrotrons was supported by the Danish Ministry of Higher Education and Science through the Instrument Center DANSCATT and the European Community's Seventh Framework Programme (FP7/2007-2013) under BioStruct-X (grant agreement 283570). L.M., S.F., S.C. and H.D. were supported by Institut de Chimie Moléculaire de Grenoble FR 2607, LabEx ARCANE (ANR-11-LABX-0003-01), the PolyNat Carnot Institute and the French Agence Nationale de la Recherche (PNRB2005-11).
Author information
Authors and Affiliations
Contributions
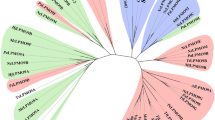
K.E.H.F. crystallized protein, collected and analyzed crystallographic data, solved crystal structures and made structural figures and tables; P.D. and T.J.S. conceived the activity, oxidation state and MS experiments, and T.J.S. performed them; J.-C.N.P. crystallized protein and collected crystallographic data; G.R.H. designed and performed the FRET kinetics experiments; L.C. performed EPR experiments and simulations; E.M.J. performed EPR experiments; M.T. and K.S.J. oversaw and directed the work of P.v.F., who purified the recombinant enzymes; L.M., S.C., S.F. and H.D. conceived and performed the FRET substrate synthesis; B.H. and N.L. performed bioinformatics analyses and alignments; F.T. collected pulsed EPR data; A.B. collected and simulated pulsed EPR spectra; G.J.D. conceived the FRET kinetics study; L.L.L. conceived the crystallographic study, collected and analyzed crystallographic data and solved crystal structures. P.H.W. conceived the EPR study. P.H.W. and L.L.L. wrote the paper with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
M.T. and P.v.F. are employees of Novozymes, a producer of enzymes for industrial use.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–6, Supplementary Notes 1–3 and Supplementary Figures 1–9. (PDF 1465 kb)
Rights and permissions
About this article
Cite this article
Frandsen, K., Simmons, T., Dupree, P. et al. The molecular basis of polysaccharide cleavage by lytic polysaccharide monooxygenases. Nat Chem Biol 12, 298–303 (2016). https://doi.org/10.1038/nchembio.2029
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2029
This article is cited by
-
Expanding the catalytic landscape of metalloenzymes with lytic polysaccharide monooxygenases
Nature Reviews Chemistry (2024)
-
Functional characterization of a lytic polysaccharide monooxygenase from Schizophyllum commune that degrades non-crystalline substrates
Scientific Reports (2023)
-
Synergistic action of lytic polysaccharide monooxygenase with glycoside hydrolase for lignocellulosic waste valorization: a review
Biomass Conversion and Biorefinery (2023)
-
Chitinolytic enzymes contribute to the pathogenicity of Aliivibrio salmonicida LFI1238 in the invasive phase of cold-water vibriosis
BMC Microbiology (2022)
-
Integrative structure determination reveals functional global flexibility for an ultra-multimodular arabinanase
Communications Biology (2022)