Abstract
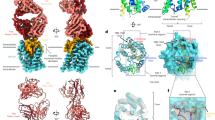
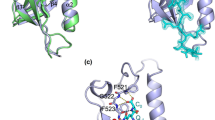
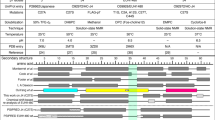
Hepatitis B virus (HBV) is an infectious, potentially lethal human pathogen. However, there are no effective therapies for chronic HBV infections. Antiviral development is hampered by the lack of high-resolution structures for essential HBV protein-protein interactions. The interaction between preS1, an HBV surface-protein domain, and its human binding partner, γ2-adaptin, subverts the membrane-trafficking apparatus to mediate virion export. This interaction is a putative drug target. We report here atomic-resolution descriptions of the binding thermodynamics and structural biology of the interaction between preS1 and the EAR domain of γ2-adaptin. NMR, protein engineering, X-ray crystallography and MS showed that preS1 contains multiple γ2-EAR–binding motifs that mimic the membrane-trafficking motifs (and binding modes) of host proteins. These motifs localize together to a relatively rigid, functionally important region of preS1, an intrinsically disordered protein. The preS1–γ2-EAR interaction was relatively weak and efficiently outcompeted by a synthetic peptide. Our data provide the structural road map for developing peptidomimetic antivirals targeting the γ2-EAR–preS1 interaction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Beck, J. & Nassal, M. Hepatitis B virus replication. World J. Gastroenterol. 13, 48–64 (2007).
Bruss, V. Envelopment of the hepatitis B virus nucleocapsid. Virus Res. 106, 199–209 (2004).
Nassal, M. Hepatitis B virus replication: novel roles for virus-host interactions. Intervirology 42, 100–116 (1999).
Steven, A.C. et al. Structure, assembly, and antigenicity of hepatitis B virus capsid proteins. Adv. Virus Res. 64, 125–164 (2005).
Glebe, D. & Urban, S. Viral and cellular determinants involved in hepadnaviral entry. World J. Gastroenterol. 13, 22–38 (2007).
Roseman, A.M., Berriman, J.A., Wynne, S.A., Butler, P.J.G. & Crowther, R.A. A structural model for maturation of the hepatitis B virus core. Proc. Natl. Acad. Sci. USA 102, 15821–15826 (2005).
Seitz, S., Urban, S., Antoni, C. & Böttcher, B. Cryo-electron microscopy of hepatitis B virions reveals variability in envelope capsid interactions. EMBO J. 26, 4160–4167 (2007).
Wynne, S.A., Crowther, R.A. & Leslie, A.G. The crystal structure of the human hepatitis B virus capsid. Mol. Cell 3, 771–780 (1999).
Awe, K., Lambert, C. & Prange, R. Mammalian BiP controls posttranslational ER translocation of the hepatitis B virus large envelope protein. FEBS Lett. 582, 3179–3184 (2008).
Yan, H. et al. Sodium taurocholate cotransporting polypeptide is a functional receptor for human hepatitis B and D virus. Elife 2012, e00049 (2012).
Prange, R. Host factors involved in hepatitis B virus maturation, assembly, and egress. Med. Microbiol. Immunol. (Berl.) 201, 449–461 (2012).
Hartmann-Stühler, C. & Prange, R. Hepatitis B virus large envelope protein interacts with γ2-adaptin, a clathrin adaptor-related protein. J. Virol. 75, 5343–5351 (2001).
Döring, T., Gotthardt, K., Stieler, J. & Prange, R. γ2-adaptin is functioning in the late endosomal sorting pathway and interacts with ESCRT-I and -III subunits. Biochim. Biophys. Acta 1803, 1252–1264 (2010).
Boehm, M. & Bonifacino, J.S. Adaptins: the final recount. Mol. Biol. Cell 12, 2907–2920 (2001).
Lewin, D.A. et al. Cloning, expression, and localization of a novel γ-adaptin–like molecule. FEBS Lett. 435, 263–268 (1998).
Rost, M., Döring, T. & Prange, R. γ2-Adaptin, a ubiquitin-interacting adaptor, is a substrate to coupled ubiquitination by the ubiquitin ligase Nedd4 and functions in the endosomal pathway. J. Biol. Chem. 283, 32119–32130 (2008).
Takatsu, H., Sakurai, M., Shin, H.W., Murakami, K. & Nakayama, K. Identification and characterization of novel clathrin adaptor-related proteins. J. Biol. Chem. 273, 24693–24700 (1998).
Zizioli, D. et al. Early embryonic death of mice deficient in γ-adaptin. J. Biol. Chem. 274, 5385–5390 (1999).
Lambert, C., Döring, T. & Prange, R. Hepatitis B virus maturation is sensitive to functional inhibition of ESCRT-III, Vps4, and γ2-adaptin. J. Virol. 81, 9050–9060 (2007).
Rost, M. et al. γ-adaptin, a novel ubiquitin-interacting adaptor, and Nedd4 ubiquitin ligase control hepatitis B virus maturation. J. Biol. Chem. 281, 29297–29308 (2006).
Chi, S.W., Kim, D., Lee, S., Chang, I. & Han, K. Pre-structured motifs in the natively unstructured preS1 surface antigen of hepatitis B virus. Protein Sci. 16, 2108–2117 (2007).
Tompa, P. & Fuxreiter, M. Fuzzy complexes: polymorphism and structural disorder in protein-protein interactions. Trends Biochem. Sci. 33, 2–8 (2008).
Dunker, A.K. et al. The unfoldomics decade: an update on intrinsically disordered proteins. BMC Genomics 9 (suppl. 2), S1 (2008).
Mattera, R., Ritter, B., Sidhu, S.S., McPherson, P.S. & Bonifacino, J.S. Definition of the consensus motif recognized by γ-adaptin ear domains. J. Biol. Chem. 279, 8018–8028 (2004).
Kent, H.M., McMahon, H.T., Evans, P.R., Benmerah, A. & Owen, D.J. γ-adaptin appendage domain: structure and binding site for Eps15 and γ-synergin. Structure 10, 1139–1148 (2002).
Miller, G.J., Mattera, R., Bonifacino, J.S. & Hurley, J.H. Recognition of accessory protein motifs by the γ-adaptin ear domain of GGA3. Nat. Struct. Biol. 10, 599–606 (2003).
Wishart, D.S. & Sykes, B.D. Chemical shifts as a tool for structure determination. Methods Enzymol. 239, 363–392 (1994).
Turnbull, W.B. & Daranas, A.H. On the value of c: can low affinity systems be studied by isothermal titration calorimetry? J. Am. Chem. Soc. 125, 14859–14866 (2003).
Inoue, M. et al. Molecular basis for autoregulatory interaction between GAE domain and hinge region of GGA1. Traffic 8, 904–913 (2007).
Mills, I.G. et al. EpsinR: an AP1/clathrin interacting protein involved in vesicle trafficking. J. Cell Biol. 160, 213–222 (2003).
Yamada, Y., Shiba, T., Kato, R. & Nakayama, K. Structure determination of GGA-GAE and γ1-ear in complex with peptides: crystallization of low-affinity complexes in membrane traffic. Acta Crystallogr. D Biol. Crystallogr. 61, 731–736 (2005).
Mészáros, B., Simon, I. & Dosztányi, Z. Prediction of protein binding regions in disordered proteins. PLoS Comput. Biol. 5, e1000376 (2009).
Mooney, C., Pollastri, G., Shields, D.C. & Haslam, N.J. Prediction of short linear protein binding regions. J. Mol. Biol. 415, 193–204 (2012).
Ferguson, N., Sharpe, T.D., Johnson, C.M., Schartau, P.J. & Fersht, A.R. Analysis of “downhill” protein folding. Nature 445, E14–E15 (2007).
Reddy, T. & Rainey, J.K. Interpretation of biomolecular NMR spin relaxation parameters. Biochem. Cell Biol. 88, 131–142 (2010).
Smith, D.P. et al. Deciphering drift time measurements from travelling wave ion mobility spectrometry–mass spectrometry studies. Eur. J. Mass Spectrom. (Chichester, Eng.) 15, 113–130 (2009).
Smith, D.P., Giles, K., Bateman, R.H., Radford, S.E. & Ashcroft, A.E. Monitoring copopulated conformational states during protein folding events using electrospray ionization–ion mobility spectrometry–mass spectrometry. J. Am. Soc. Mass Spectrom. 18, 2180–2190 (2007).
Clemmer, D.E., Hudgins, R.R. & Jarrold, M.F. Naked protein conformations: cytochrome c in the gas phase. J. Am. Chem. Soc. 117, 10141–10142 (1995).
Ruotolo, B.T. & Robinson, C.V. Aspects of native proteins are retained in vacuum. Curr. Opin. Chem. Biol. 10, 402–408 (2006).
Saikusa, K. et al. Characterisation of an intrinsically disordered protein complex of Swi5-Sfr1 by ion mobility mass spectrometry and small-angle X-ray scattering. Analyst 138, 1441–1449 (2013).
Pagel, K., Natan, E., Hall, Z., Fersht, A.R. & Robinson, C.V. Intrinsically disordered p53 and its complexes populate compact conformations in the gas phase. Angew. Chem. Int. Edn. Engl. 52, 361–365 (2013).
Davey, N.E., Travé, G. & Gibson, T.J. How viruses hijack cell regulation. Trends Biochem. Sci. 36, 159–169 (2011).
Maritzen, T. et al. A novel subtype of AP-1-binding motif within the palmitoylated trans-Golgi network/endosomal accessory protein Gadkin/γ-BAR. J. Biol. Chem. 285, 4074–4086 (2010).
Schulze, A., Schieck, A., Ni, Y., Mier, W. & Urban, S. Fine mapping of pre-S sequence requirements for hepatitis B virus large envelope protein–mediated receptor interaction. J. Virol. 84, 1989–2000 (2010).
Le Seyec, J., Chouteau, P., Cannie, I., Gripon, P. & Guguen-Guillouzo, C. Infection process of the hepatitis B virus depends on the presence of a defined sequence in the pre-S1 domain. J. Virol. 73, 2052–2057 (1999).
Bruss, V. & Thomssen, R. Mapping a region of the large envelope protein required for hepatitis B virion maturation. J. Virol. 68, 1643–1650 (1994).
Böttcher, B. et al. Peptides that block hepatitis B virus assembly: analysis by cryomicroscopy, mutagenesis and transfection. EMBO J. 17, 6839–6845 (1998).
Freund, S.M.V., Johnson, C.M., Jaulent, A.M. & Ferguson, N. Moving towards high-resolution descriptions of the molecular interactions and structural rearrangements of the human hepatitis B core protein. J. Mol. Biol. 384, 1301–1313 (2008).
Nogi, T. et al. Structural basis for the accessory protein recruitment by the γ-adaptin ear domain. Nat. Struct. Biol. 9, 527–531 (2002).
Sattler, M., Schleucher, J. & Griesinger, C. Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog. Nucl. Magn. Reson. Spectrosc. 34, 93–158 (1999).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D Biol. Crystallogr. 62, 72–82 (2006).
Long, F., Vagin, A.A., Young, P. & Murshudov, G.N. BALBES: a molecular-replacement pipeline. Acta Crystallogr. D Biol. Crystallogr. 64, 125–132 (2008).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Langer, G., Cohen, S.X., Lamzin, V.S. & Perrakis, A. Automated macromolecular model building for X-ray crystallography using ARP/wARP version 7. Nat. Protoc. 3, 1171–1179 (2008).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Ruotolo, B.T., Benesch, J.L.P., Sandercock, A.M., Hyung, S.-J. & Robinson, C.V. Ion mobility–mass spectrometry analysis of large protein complexes. Nat. Protoc. 3, 1139–1152 (2008).
Acknowledgements
We would like to thank K. Nakayama (University of Kyoto) for the γ2-adaptin clone; J. Lyons (Trinity College Dublin) and Deutsches Elektronen-Synchrotron–European Molecular Biology Laboratory beamline scientists for assistance with X-ray data collection; N. Moran (Royal College of Surgeons in Ireland) for access to biophysical instrumentation; M. Caffrey (Trinity College Dublin) for access to X-ray apparatus; and D. Shields and C. Mooney for bioinformatics help. This work was supported by Science Foundation Ireland (SFI)–President of Ireland Young Researcher Award (09/YI/B1682 to N.F.), Stokes Lecturer Award (07/SK/B1224a to N.F.), Royal Irish Academy–Royal Society Exchange award (IE111031 to N.F. and A.E.A.), SFI grant (07/IN.1/B1836) and US National Institutes of Health grants (GM75915, P50GM073210 U54GM094599) to M. Caffrey (funding V.E.P. and X-ray generator). D.A.S. was funded by an Engineering and Physical Sciences Research Council PhD Studentship. The Synapt mass spectrometer was purchased with Biotechnology and Biological Sciences Research Council UK funds (BB/E012558/1).
Author information
Authors and Affiliations
Contributions
M.C.J. purified proteins, performed ITC and NMR experiments, determined crystal structures, analyzed data and wrote the paper; J.V. purified proteins, performed ITC and NMR experiments, analyzed data and wrote the paper; G.J.P.R. performed and analyzed NMR experiments and wrote the paper; J.M. performed NMR experiments; D.A.S. performed and analyzed MS experiments and wrote the paper; V.E.P. determined crystal structures and wrote the paper; C.M.J. performed and analyzed calorimetry experiments; A.E.A. designed and interpreted MS experiments and wrote the paper; S.M.V.F. performed and analyzed NMR experiments and wrote the paper; N.F. directed research, designed and interpreted experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–6 and Supplementary Tables 1–2. (PDF 4090 kb)
Rights and permissions
About this article
Cite this article
Jürgens, M., Vörös, J., Rautureau, G. et al. The hepatitis B virus preS1 domain hijacks host trafficking proteins by motif mimicry. Nat Chem Biol 9, 540–547 (2013). https://doi.org/10.1038/nchembio.1294
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1294
This article is cited by
-
Determining buffer conditions for downstream processing of VLP-based recombinant hepatitis B surface antigen using multimodal resins in bind-elute and flow-through purification modes
Scientific Reports (2023)
-
Insights into the client protein release mechanism of the ATP-independent chaperone Spy
Nature Communications (2022)
-
Recognition by host nuclear transport proteins drives disorder-to-order transition in Hendra virus V
Scientific Reports (2018)
-
Structure-based rational design of peptide inhibitors to disrupt the recognition and interaction between hepatitis B virus large envelope protein and human hepatocyte receptor γ2-adaptin
Medicinal Chemistry Research (2017)