Abstract
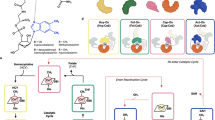
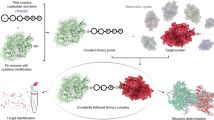
The biosynthesis of many vitamins and coenzymes has often proven difficult to elucidate owing to a combination of low abundance and kinetic lability of the pathway intermediates. Through a serial reconstruction of the cobalamin (vitamin B12) pathway in Escherichia coli and by His tagging the terminal enzyme in the reaction sequence, we have observed that many unstable intermediates can be isolated as tightly bound enzyme-product complexes. Together, these approaches have been used to extract intermediates between precorrin-4 and hydrogenobyrinic acid in their free acid form and permitted the delineation of the overall reaction catalyzed by CobL, including the formal elucidation of precorrin-7 as a metabolite. Furthermore, a substrate-carrier protein, CobE, that can also be used to stabilize some of the transient metabolic intermediates and enhance their onward transformation, has been identified. The tight association of pathway intermediates with enzymes provides evidence for a form of metabolite channeling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Webb, M.E., Marquet, A., Mendel, R.R., Rebeille, F. & Smith, A.G. Elucidating biosynthetic pathways for vitamins and cofactors. Nat. Prod. Rep. 24, 988–1008 (2007).
Battersby, A.R. How nature builds the pigments of life: the conquest of vitamin B12 . Science 264, 1551–1557 (1994).
Warren, M.J., Raux, E., Schubert, H.L. & Escalante-Semerena, J.C. The biosynthesis of adenosylcobalamin (vitamin B12). Nat. Prod. Rep. 19, 390–412 (2002).
Blanche, F. et al. Vitamin B12: how the problem of its biosynthesis was solved. Angew. Chem. Int. Edn. Engl. 34, 383–411 (1995).
Blanche, F. et al. Hydrogenobyrinic acid—isolation, biosynthesis, and function. Angew. Chem. Int. Edn. Engl. 29, 884–886 (1990).
McGoldrick, H.M. et al. Identification and characterization of a novel vitamin B12 (cobalamin) biosynthetic enzyme (CobZ) from Rhodobacter capsulatus, containing flavin, heme, and Fe-S cofactors. J. Biol. Chem. 280, 1086–1094 (2005).
Uzar, H.C., Battersby, A.R., Carpenter, T.A. & Leeper, F.J. Biosynthesis of porphyrins and related macrocycles. Part 28. Development of a pulse labeling method to determine the C-methylation sequence for vitamin B12 . J. Chem. Soc. Perkin Trans. 1, 1689–1696 (1987).
Blanche, F., Debussche, L., Thibaut, D., Crouzet, J. & Cameron, B. Purification and characterization of S-adenosyl-L-methionine: uroporphyrinogen III methyltransferase from Pseudomonas denitrificans. J. Bacteriol. 171, 4222–4231 (1989).
Warren, M.J. et al. Enzymatic synthesis and structure of precorrin-3, a trimethyldipyrrocorphin intermediate in vitamin B12 biosynthesis. Biochemistry 31, 603–609 (1992).
Debussche, L. et al. Biosynthesis of vitamin B12: structure of precorrin-3B, the trimethylated substrate of the enzyme catalysing ring contraction. J. Chem. Soc. Chem. Commun. 1100–1103 (1993).
Scott, A.I. et al. Biosynthesis of vitamin B12. Discovery of the enzymes for oxidative ring contraction and insertion of the fourth methyl group. FEBS Lett. 331, 105–108 (1993).
Thibaut, D. et al. Biosynthesis of vitamin-B12—the structure of factor-IV, the oxidized form of precorrin-4. J. Chem. Soc. Chem. Commun. 513–515 (1993).
Min, C. et al. Isolation, structure and genetically engineered synthesis of precorrin-5. J. Am. Chem. Soc. 115, 10380–10381 (1993).
Thibaut, D., Blanche, F., Debussche, L., Leeper, F.J. & Battersby, A.R. Biosynthesis of vitamin B12: structure of precorrin-6x octamethyl ester. Proc. Natl. Acad. Sci. USA 87, 8800–8804 (1990).
Blanche, F. et al. Precorrin-6x reductase from Pseudomonas denitrificans: purification and characterization of the enzyme and identification of the structural gene. J. Bacteriol. 174, 1036–1042 (1992).
Blanche, F. et al. Biosynthesis of vitamin B12 in Pseudomonas denitrificans: the biosynthetic sequence from precorrin-6y to precorrin-8x is catalyzed by the cobL gene product. J. Bacteriol. 174, 1050–1052 (1992).
Thibaut, D. et al. The final step in the biosynthesis of hydrogenobyrinic acid is catalyzed by the cobH gene product with precorrin-8x as the substrate. J. Bacteriol. 174, 1043–1049 (1992).
Santander, P.J., Kajiwara, Y., Williams, H.J. & Scott, A.I. Structural characterization of novel cobalt corrinoids synthesized by enzymes of the vitamin B12 anaerobic pathway. Bioorg. Med. Chem. 14, 724–731 (2006).
Scott, A.I., Warren, M.J., Roessner, C.A., Stolowich, N.J. & Santander, P.J. Development of an overmethylation strategy for corrin synthesis. Multi-enzyme preparation of pyrrocorphins. J. Chem. Soc. Chem. Commun. 593–597 (1990).
Shipman, L.W., Li, D., Roessner, C.A., Scott, A.I. & Sacchettini, J.C. Crystal structure of precorrin-8x methyl mutase. Structure 9, 587–596 (2001).
Pettersson, G. No convincing evidence is available for metabolite channeling between enzymes forming dynamic complexes. J. Theor. Biol. 152, 65–69 (1991).
Wu, X.M., Gutfreund, H., Lakatos, S. & Chock, P.B. Substrate channeling in glycolysis: a phantom phenomenon. Proc. Natl. Acad. Sci. USA 88, 497–501 (1991).
Huang, X., Holden, H.M. & Raushel, F.M. Channeling of substrates and intermediates in enzyme-catalyzed reactions. Annu. Rev. Biochem. 70, 149–180 (2001).
Jørgensen, K. et al. Metabolon formation and metabolic channeling in the biosynthesis of plant natural products. Curr. Opin. Plant Biol. 8, 280–291 (2005).
Miles, E.W., Rhee, S. & Davies, D.R. The molecular basis of substrate channeling. J. Biol. Chem. 274, 12193–12196 (1999).
McGuffee, S.R. & Elcock, A.H. Diffusion, crowding & protein stability in a dynamic molecular model of the bacterial cytoplasm. PLoS Comput. Biol. 6, e1000694 (2010).
Mika, J.T., van den Bogaart, G., Veenhoff, L., Krasnikov, V. & Poolman, B. Molecular sieving properties of the cytoplasm of Escherichia coli and consequences of osmotic stress. Mol. Microbiol. 77, 200–207 (2010).
Robinson, J.B. Jr., Inman, L., Sumegi, B. & Srere, P.A. Further characterization of the Krebs tricarboxylic acid cycle metabolon. J. Biol. Chem. 262, 1786–1790 (1987).
Dogutan, D.K. et al. Hangman corroles: efficient synthesis and oxygen reaction chemistry. J. Am. Chem. Soc. 133, 131–140 (2011).
Lee, C.H., Dogutan, D.K. & Nocera, D.G. Hydrogen generation by hangman metalloporphyrins. J. Am. Chem. Soc. 133, 8775–8777 (2011).
Waibel, R. et al. New derivatives of vitamin B12 show preferential targeting of tumors. Cancer Res. 68, 2904–2911 (2008).
Sambrook, J. & Russell, D. Site-specific mutagenesis by overlap extension in Molecular Cloning: A Laboratory Manual 3rd edn., Ch. 13, 13.36–13.39 (Cold Spring Harbor Laboratory Press, 2001).
Wishart, D.S. & Sykes, B.D. Chemical-shifts as a tool for structure determination. Methods Enzymol. 239, 363–392 (1994).
Delaglio, F. et al. NMRpipe—a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Vranken, W.F. et al. The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59, 687–696 (2005).
Vévodová, J., Graham, R.M., Raux, E., Warren, M.J. & Wilson, K.S. Crystallization and preliminary structure analysis of CobE, an essential protein of cobalamin (vitamin B12) biosynthesis. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 61, 442–444 (2005).
Seyedarabi, A. et al. Cloning, purification and preliminary crystallographic analysis of cobalamin methyltransferases from Rhodobacter capsulatus. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 66, 1652–1656 (2010).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D Biol. Crystallogr. 66, 22–25 (2010).
Brunger, A.T. Version 1.2 of the Crystallography and NMR system. Nat. Protoc. 2, 2728–2733 (2007).
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D Biol. Crystallogr. 62, 1002–1011 (2006).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Battye, T.G.G., Kontogiannis, L., Johnson, O., Powell, H.R. & Leslie, A.G.W. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D Biol. Crystallogr. 67, 271–281 (2011).
Blanc, E. et al. Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr. D Biol. Crystallogr. 60, 2210–2221 (2004).
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010).
Acknowledgements
This work was supported by grants from the Biotechnology and Biological Sciences Research Council (BB/E024203 and BB/I013334) and the Wellcome Trust (091163/Z/10/Z). We thank M. Rowe for additional NMR technical support. Diffraction data were collected at the European Synchrotron Radiation Facility, Grenoble, France (for CobE) and the Diamond Light Source, Oxfordshire, UK (for CobL and CobH). We thank C. Roessner (Texas A&M University) for a clone of the P. denitrificans cobG.
Author information
Authors and Affiliations
Contributions
E.D. designed and performed most of the experiments and analysis with support from A.D.L. and S.S.; A.D.L. performed MS analysis. S.L.T. performed all NMR data acquisition, which was analyzed with M.J.H. S.S., A.S. and R.W.P. contributed the CobLC and CobH–HBA structures and CobL and CobE–HBA models. J.W. and K.S.W. contributed the CobE structure. D.B. and S.S. determined the CobH–5-desmethyl-HBA structure. M.A.G. provided insight into substrate channeling. M.J.W. directed all aspects of the project. E.D. and M.J.W. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 9560 kb)
Rights and permissions
About this article
Cite this article
Deery, E., Schroeder, S., Lawrence, A. et al. An enzyme-trap approach allows isolation of intermediates in cobalamin biosynthesis. Nat Chem Biol 8, 933–940 (2012). https://doi.org/10.1038/nchembio.1086
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1086
This article is cited by
-
A synthetic cell-free 36-enzyme reaction system for vitamin B12 production
Nature Communications (2023)
-
Structure of full-length cobalamin-dependent methionine synthase and cofactor loading captured in crystallo
Nature Communications (2023)
-
Calculating metalation in cells reveals CobW acquires CoII for vitamin B12 biosynthesis while related proteins prefer ZnII
Nature Communications (2021)
-
Optimization of hydrogenobyrinic acid biosynthesis in Escherichia coli using multi-level metabolic engineering strategies
Microbial Cell Factories (2020)
-
Metabolic engineering of Escherichia coli for de novo biosynthesis of vitamin B12
Nature Communications (2018)