Abstract
Pluripotent embryonic stem cells (ESCs) self-renew or differentiate into all tissues of the developing embryo and cell-specification factors are necessary to balance gene expression. Here we delineate the function of the PHD-finger protein 5a (Phf5a) in ESC self-renewal and ascribe its role in regulating pluripotency, cellular reprogramming and myoblast specification. We demonstrate that Phf5a is essential for maintaining pluripotency, since depleted ESCs exhibit hallmarks of differentiation. Mechanistically, we attribute Phf5a function to the stabilization of the Paf1 transcriptional complex and control of RNA polymerase II elongation on pluripotency loci. Apart from an ESC-specific factor, we demonstrate that Phf5a controls differentiation of adult myoblasts. Our findings suggest a potent mode of regulation by Phf5a in stem cells, which directs their transcriptional programme, ultimately regulating maintenance of pluripotency and cellular reprogramming.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981).
Martin, G. R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc. Natl Acad. Sci. USA 78, 7634–7638 (1981).
Orkin, S. H. & Hochedlinger, K. Chromatin connections to pluripotency and cellular reprogramming. Cell 145, 835–850 (2011).
Buckley, S. M. et al. Regulation of pluripotency and cellular reprogramming by the ubiquitin-proteasome system. Cell Stem Cell 11, 783–798 (2012).
Trappe, R. et al. Identification and characterization of a novel murine multigene family containing a PHD-finger-like motif. Biochem. Biophys. Res. Commun. 293, 816–826 (2002).
Trappe, R., Schulze, E., Rzymski, T., Frode, S. & Engel, W. The Caenorhabditis elegans ortholog of human PHF5a shows a muscle-specific expression domain and is essential for C. elegans morphogenetic development. Biochem. Biophys. Res. Commun. 297, 1049–1057 (2002).
Jaehning, J. A. The Paf1 complex: platform or player in RNA polymerase II transcription? Biochim. Biophys. Acta 1799, 379–388 (2010).
Tomson, B. N. & Arndt, K. M. The many roles of the conserved eukaryotic Paf1 complex in regulating transcription, histone modifications, and disease states. Biochim. Biophys. Acta 1829, 116–126 (2013).
Akanuma, T., Koshida, S., Kawamura, A., Kishimoto, Y. & Takada, S. Paf1 complex homologues are required for Notch-regulated transcription during somite segmentation. EMBO Rep. 8, 858–863 (2007).
Langenbacher, A. D. The PAF1 complex differentially regulates cardiomyocyte specification. Dev. Biol. 353, 19–28 (2011).
Nguyen, C. T., Langenbacher, A., Hsieh, M. & Chen, J. N. The PAF1 complex component Leo1 is essential for cardiac and neural crest development in zebrafish. Dev. Biol. 341, 167–175 (2010).
Schaniel, C. et al. Smarcc1/Baf155 couples self-renewal gene repression with changes in chromatin structure in mouse embryonic stem cells. Stem Cells 27, 2979–2991 (2009).
Walter, W., Sanchez-Cabo, F. & Ricote, M. GOplot: an R package for visually combining expression data with functional analysis. Bioinformatics 31, 2912–2914 (2015).
Premsrirut, P. K. et al. A rapid and scalable system for studying gene function in mice using conditional RNA interference. Cell 145, 145–158 (2011).
Lian, X. et al. Directed cardiomyocyte differentiation from human pluripotent stem cells by modulating Wnt/β-catenin signaling under fully defined conditions. Nat. Protoc. 8, 162–175 (2013).
Shelton, M. et al. Derivation and expansion of PAX7-positive muscle progenitors from human and mouse embryonic stem cells. Stem Cell Rep. 3, 516–529 (2014).
Madan, B. et al. The pluripotency-associated gene Dppa4 is dispensable for embryonic stem cell identity and germ cell development but essential for embryogenesis. Mol. Cell. Biol. 29, 3186–3203 (2009).
Stadtfeld, M., Maherali, N., Borkent, M. & Hochedlinger, K. A reprogrammable mouse strain from gene-targeted embryonic stem cells. Nat. Methods 7, 53–55 (2010).
Beard, C., Hochedlinger, K., Plath, K., Wutz, A. & Jaenisch, R. Efficient method to generate single-copy transgenic mice by site-specific integration in embryonic stem cells. Genesis 44, 23–28 (2006).
Sommer, C. A. et al. Induced pluripotent stem cell generation using a single lentiviral stem cell cassette. Stem Cells 27, 543–549 (2009).
Ding, L. et al. A genome-scale RNAi screen for Oct4 modulators defines a role of the Paf1 complex for embryonic stem cell identity. Cell Stem Cell 4, 403–415 (2009).
Ponnusamy, M. P. et al. RNA polymerase II associated factor 1/PD2 maintains self-renewal by its interaction with Oct3/4 in mouse embryonic stem cells. Stem Cells 27, 3001–3011 (2009).
Hubert, C. G. et al. Genome-wide RNAi screens in human brain tumor isolates reveal a novel viability requirement for PHF5A. Genes Dev. 27, 1032–1045 (2013).
Shen, S. et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc. Natl Acad. Sci. USA 111, E5593–E5601 (2014).
Rahl, P. B. et al. c-Myc regulates transcriptional pause release. Cell 141, 432–445 (2010).
Kim, J., Guermah, M. & Roeder, R. G. The human PAF1 complex acts in chromatin transcription elongation both independently and cooperatively with SII/TFIIS. Cell 140, 491–503 (2010).
Mueller, C. L., Porter, S. E., Hoffman, M. G. & Jaehning, J. A. The Paf1 complex has functions independent of actively transcribing RNA polymerase II. Mol. Cell 14, 447–456 (2004).
Nordick, K., Hoffman, M. G., Betz, J. L. & Jaehning, J. A. Direct interactions between the Paf1 complex and a cleavage and polyadenylation factor are revealed by dissociation of Paf1 from RNA polymerase II. Eukaryot. Cell 7, 1158–1167 (2008).
Smith, E. & Shilatifard, A. Transcriptional elongation checkpoint control in development and disease. Genes Dev. 27, 1079–1088 (2013).
Levine, M. Paused RNA polymerase II as a developmental checkpoint. Cell 145, 502–511 (2011).
Liu, L. et al. Transcriptional pause release is a rate-limiting step for somatic cell reprogramming. Cell Stem Cell 15, 574–588 (2014).
Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Jonkers, I., Kwak, H. & Lis, J. T. Genome-wide dynamics of Pol II elongation and its interplay with promoter proximal pausing, chromatin, and exons. eLife 3, e02407 (2014).
Bienz, M. The PHD finger, a nuclear protein-interaction domain. Trends Biochem. Sci. 31, 35–40 (2006).
Li, H. et al. Molecular basis for site-specific read-out of histone H3K4me3 by the BPTF PHD finger of NURF. Nature 442, 91–95 (2006).
Pena, P. V. et al. Molecular mechanism of histone H3K4me3 recognition by plant homeodomain of ING2. Nature 442, 100–103 (2006).
Taverna, S. D., Li, H., Ruthenburg, A. J., Allis, C. D. & Patel, D. J. How chromatin-binding modules interpret histone modifications: lessons from professional pocket pickers. Nat. Struct. Mol. Biol. 14, 1025–1040 (2007).
Rigbolt, K. T. et al. System-wide temporal characterization of the proteome and phosphoproteome of human embryonic stem cell differentiation. Sci. Signal. 4, rs3 (2011).
Chaudhary, K., Deb, S., Moniaux, N., Ponnusamy, M. P. & Batra, S. K. Human RNA polymerase II-associated factor complex: dysregulation in cancer. Oncogene 26, 7499–7507 (2007).
Yu, M. et al. RNA polymerase II-associated factor 1 regulates the release and phosphorylation of paused RNA polymerase II. Science 350, 1383–1386 (2015).
Chen, F. X. et al. PAF1, a molecular regulator of promoter-proximal pausing by RNA polymerase II. Cell 162, 1003–1015 (2015).
Falck, E. & Klinga-Levan, K. Expression patterns of Phf5a/PHF5A and Gja1/GJA1 in rat and human endometrial cancer. Cancer Cell Int. 13, 43 (2013).
Rzymski, T., Grzmil, P., Meinhardt, A., Wolf, S. & Burfeind, P. PHF5A represents a bridge protein between splicing proteins and ATP-dependent helicases and is differentially expressed during mouse spermatogenesis. Cytogenet. Genome Res. 121, 232–244 (2008).
Nijhawan, D. et al. Cancer vulnerabilities unveiled by genomic loss. Cell 150, 842–854 (2012).
Reavie, L. et al. Regulation of hematopoietic stem cell differentiation by a single ubiquitin ligase-substrate complex. Nat. Immunol. 11, 207–215 (2010).
Blais, A., van Oevelen, C. J., Margueron, R., Acosta-Alvear, D. & Dynlacht, B. D. Retinoblastoma tumor suppressor protein-dependent methylation of histone H3 lysine 27 is associated with irreversible cell cycle exit. J. Cell Biol. 179, 1399–1412 (2007).
Springer, M. L., Rando, T. A. & Blau, H. M. in Current Protocols in Human Genetics (eds Haines, J. L. et al.) Ch. 13, Unit 13–14 (John Wiley, 2002).
Gloeckner, C. J., Boldt, K. & Ueffing, M. in Current Protocols in Protein Science (eds Coligan, J. E. et al.) Ch. 19, Unit 19–20 (John Wiley, 2009).
Hochedlinger, K., Yamada, Y., Beard, C. & Jaenisch, R. Ectopic expression of Oct-4 blocks progenitor-cell differentiation and causes dysplasia in epithelial tissues. Cell 121, 465–477 (2005).
Zhu, B. et al. The human PAF complex coordinates transcription with events downstream of RNA synthesis. Genes Dev. 19, 1668–1673 (2005).
Dignam, J. D., Lebovitz, R. M. & Roeder, R. G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11, 1475–1489 (1983).
Gao, Z. et al. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Mol. Cell 45, 344–356 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ntziachristos, P. et al. Genetic inactivation of the polycomb repressive complex 2 in T cell acute lymphoblastic leukemia. Nat. Med. 18, 298–301 (2012).
Chen, H. & Boutros, P. C. VennDiagram: a package for the generation of highly-customizable Venn and Euler diagrams in R. BMC Bioinformatics 12, 35 (2011).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Ntziachristos, P. et al. Contrasting roles of histone 3 lysine 27 demethylases in acute lymphoblastic leukaemia. Nature 514, 513–517 (2014).
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We thank all members of the Aifantis laboratory for useful comments and discussions throughout the duration of this project; Z. Gao and P. Voigt for experimental help with glycerol gradients and helpful discussions; K. J. Armache and P. De Ioannes Fernandez for help with protein purification, helpful discussions and manuscript preparation; H.-J. Fehling for the Dppa4-RFP/Brachyury-GFP reporter ESC line; A. Heguy and the NYU Genome Technology Center (supported in part by National Institutes of Health (NIH)/National Cancer Institute (NCI) grant P30CA016087-30) for expertize with sequencing experiments; the NYU Histology Core (5P30CA16087-31) for assistance; C. Loomis and L. Chiriboga for immunohistochemistry experiments; and H. Li and T. Liu at the Center for Advanced Proteomics Research, New Jersey School of Medicine for mass spectrometry. This work has used computing resources at the High Performance Computing Facility of the Center of Health Informatics and Bioinformatics at the NYU Medical Center. A.S. is supported by the NYSTEM institutional NYU Stem Cell Training Grant (C026880). I.A. is supported by the NIH (RO1CA133379, RO1CA105129, RO1CA149655, 5RO1CA173636, 1RO1CA194923) and the NYSTEM programme of the New York State Health Department (NYSTEM-N11G-255).
Author information
Authors and Affiliations
Contributions
I.A. and A.S. designed the experiments and wrote the manuscript. A.S. performed the experiments. A.T., C.L., T.T. and I.D. designed and performed the analysis of genome-wide data. P.N. provided expertize in sequencing experiments and contributed in manuscript preparation. A.L.G.N. performed histological examination of teratomas. S.B. contributed ideas. M.S., Y.Y., B.D.D., S.R. and B.D.S. provided materials and tips related to this study, helped with ideas and concepts and contributed to manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Phf5a depletion results in aberrant ESC differentiation.
(a) Alignment of Phf5a protein sequence from different organisms. Green: Conserved cysteine residues, which coordinate Zn2+. Orange: Nuclear localization signal (NLS). (b) Relative expression of Phf5a transcript by qRT-PCR in ESC differentiation. n = 6 biologically independent replicates (see Supplementary Table 5). ∗∗p = 0.0001, respectively, two-sided Student’s t-test, values represent the mean ± s.d. (c) Nanog-GFP ESC colonies following knockdown with shControl or shPhf5a respectively. Scale bars, 100 μm. (d) FACS plots showing levels of apoptosis and cell death following shPhf5a knockdown in ESCs. Proteasome inhibition by MG132 is used as positive control. (e) Western blot analysis of total and cleaved Caspase-3 following shPhf5a knockdown in ESCs and differentiation. Proteasome inhibition by MG132 is used as positive control. (f) Gene Ontology (GO) analysis of significantly downregulated or upregulated genes, respectively, after RNA-sequencing in ESCs using shControl or shPhf5a silencing (g) Schematic of Phf5a mRNA indicating distinct target regions for shRNAs, gRNAs and siRNA used in the study. (h) Western blot analysis of Oct4 levels following siPhf5a knockdown in ESCs. (i) Alkaline phosphatase (AP) staining of ESCs following knockdown using siRNA transfections. Scale bars, 100 μm. (j) Western blot analysis of pluripotency factors following shPhf5a knockdown in ESCs. (k) Bar graphs showing expression levels of Phf5a in ESCs of different genetic backgrounds. n = 6 biologically independent replicates (see Supplementary Table 5). MK6 and CCE: D2 and D4 ∗∗p = 0.0001, respectively. KH2: D2 and D4 p = ∗∗0.0155 and 0.0052, respectively, two-sided Student’s t-test, values represent the mean ± s.d. (l) Western blot analysis of pluripotency factors following shPhf5a knockdown in MK6 ESCs (C57BL/6).
Supplementary Figure 2 Silencing of Phf5a affects ESC pluripotency and iPS generation.
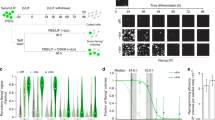
(a) Schematic of Tet-inducible mir-30 shRNA expression cassette at the targeted Col1a1 locus in ESCs. (b) Brightfield and fluorescent images of ESCs targeted with shControl and shPhf5a mir-30 shRNA expression cassettes at the Col1a1 locus following the addition of doxyxycline. Scale bars, 100 μm. (c) Comparison of Phf5a expression levels by qRT-PCR in ESCs targeted with shControl and shPhf5a mir-30 shRNA expression cassettes at the Col1a1 locus following the addition of doxycycline. n = 3 biologically independent replicates (see Supplementary Table 5). shPhf5a-2 hairpin: ∗∗p = 0.0072 and shPhf5a-3 hairpin ∗∗p = 0.008, respectively, two-sided Student’s t-test, values represent the mean ± s.d. (d) Western blot analysis of Phf5a protein levels in ESCs targeted with shControl and shPhf5a mir-30 shRNA expression cassettes at the Col1a1 locus following the addition of doxycycline. (e–g) Histology and comparison of shControl and shPhf5a teratomas using H&E or PAS stain (e and f) and desmin, nestin or cytokeratin immunohistochemistry (g), respectively. Teratomas tissue formations include respiratory cilia, keratinized skin, cartilage and goblet cells, representative of all germ layers. Arrows indicate ciliated respiratory epithelium; stars indicate goblet cells stained positive for PAS stain. Scale bars, 100 μm. (h) FACS plots showing levels of Dppa4-RFP and Brachyury-GFP following induction of mesoderm differentiation in the presence or absence of Phf5a. (i) Western blot analysis of Brachyury protein levels following induction of mesoderm differentiation in the presence or absence of Phf5a. (j) Levels of several mesoderm markers following shPhf5a depletion. n = 3 biologically independent replicates (see Supplementary Table 5). Brachyury: ∗∗p = 0.0078, Msgn: n.s: non-significant p = 0.0694, Nkx2-5: ∗∗p = 0.0010, n.s: non-significant Isl1: p = 0.2223, two-sided Student’s t-test, values represent the mean ± s.d. (k and l) Proliferation assays of ESCs (k) or MEFs (l), respectively, following shControl or shPhf5a depletion. n = 3 biologically independent replicates (see Supplementary Table 5). ESCs Day2 and Day4: ∗∗p = 0.0001, respectively. MEFs Day2 and Day4: n.s: non-signifficant, p = 0.4169 and p = 0.8769, respectively, two-sided Student’s t-test, values represent the mean ± s.d. (m and n) Expression levels of pluripotency factors (m) and western blot for Nanog protein levels (n), respectively, during ESC differentiation, in the presence or absence of ectopic expression of Phf5a. n = 3 biologically independent replicates (see Supplementary Table 5). Phf5a: ∗∗p = 0.0016, Nanog: ∗∗p = 0.0451, Pou5f1: ∗∗p = 0.0335, Sox2: p = 0.0485, Zfp42: n.s: non-signifficant p = 0.0908, Nr0b1: p = 0.0209, two-sided Student’s t-test, values represent the mean ± s.d. (o) Representative iPSC colony morphology and AP-staining following shPhf5a knockdown. Scale bars, 100 μm. (p) Western blot analysis of reprogramming markers in OKSM MEFs following shPhf5a knockdown, on day 14 post-initial doxycycline induction.
Supplementary Figure 3 Phf5a interacts with the Paf1C.
(a) Western blot analysis of Phf5a and control tagged-proteins following doxycycline induction and cytoplasmic or nuclear fractionation in ESCs. (b) Flag immunofluorescence showing nuclear localization of tagged-Phf5a following doxycycline induction in ESCs. Scale bars, 100 μm. (c) Western blot analysis of Paf1C subunits in ESC differentiation. (d) Western blot analysis of interacting proteins following doxycycline induction and streptactin purification of Paf1C subunits or GFP control in ESCs. (e) Western blot analysis of Phf5a- or control- interacting proteins following doxycycline induction and tandem affinity purification using strep-tagII and flag tags in ESCs. (f) Representative time point analysis for presence of nucleic acids following lysis of ESCs in the presence of benzonase nuclease prior to immunoprecipitation in ESCs.
Supplementary Figure 4 Phf5a regulates Paf1C subunit composition.
(a) Venn diagrams showing number of all significantly differentially expressed genes, as well as significantly downregulated and upregulated genes using RNA-sequencing after shPhf5a and shPaf1 depletion in ESCs. (b) Western blot of fractions following glycerol gradient sedimentation analysis of Paf1C subunits using one-step purification from ESCs in the presence or absence of shPhf5a knockdown. (c–e) Graphs showing quantification of percent distribution for the Paf1C subunits Paf1, Wdr61 and Cdc73 shown in (c), in the presence or absence of Phf5a, respectively, in ESCs. (f) Western blot of fractions following glycerol gradient sedimentation analysis of Swi/Snf complex subunit Smarca4 and NELF complex subunit NELF-A using one-step purification from ESCs in the presence or absence of shPhf5a knockdown. (g) Binding profiles for genomic distribution of Leo1 peaks (upstream, promoter, coding region, 5’UTR, 3’UTR, downstream and intergenic) in ESCs, showing preferential binding (42%) within gene bodies of downregulated genes, but preferential binding (31%) within promoters of upregulated genes. (h) A Tet-inducible “knock-in” tagged Phf5a ESC line was used to perform ChIP-sequencing of Phf5a using a HA-epitope in the presence of absence of doxycycline. Snapshots of Phf5a binding on representative pluripotency gene targets are shown (Pou5f1, Sall4, Prdm14, Esrrb) in the presence (blue) of absence (gray) of doxycycline.
Supplementary Figure 5 Phf5a regulates Paf1C functions on transcriptional elongation in ESCs.
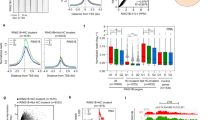
(a) Western blot analysis of Ser2-phospho- and Total RNA PolII in ESCs following CRISPR-Cas9 mediated Phf5a depletion. (b) Comparison of GRO-seq read density profiles of downregulated genes 72h following shControl or differentiated ESCs in the absence of LIF. RPKM: Reads Per Kilobase per Million total reads. (c) Comparison of GRO-seq read density profiles of downregulated genes 72h in all conditions tested (shControl, shPhf5a or differentiated ESCs in the absence of LIF). RPKM: Reads Per Kilobase per Million total reads. (d) Box plot showing comparison of log2 pausing index for downregulated genes, 72h following shControl, shPhf5a, ESC differentiated in the absence of LIF, or flavopiridol-treated ESCs, respectively, using GRO-seq analysis. Flavopiridol treatment is used as a positive control of pause-release block. n = 3 biologically independent replicates, Wilcoxon signed rank test non-parametric. (e) (Upper) Venn diagrams between Leo1 ChIP-sequencing targets in ESCs and either downregulated or upregulated genes using RNA-sequencing after shPhf5a knockdown. (Lower) Box plot showing pausing index ratios after GRO-seq analysis for the direct Leo1 targets shown above. Only downregulated targets exhibit significant promoter-proximal pausing. n = 3 biologically independent replicates, Wilcoxon signed rank test non-parametric. (f and g) Comparison of read density profiles for Ser5- (on TSSs) or Ser2- (on gene bodies) phosphorylated RNA PolII, respectively, for the specific Gene Ontology Terms chromatin organization (f) and positive regulation of transcription (g), respectively. n = 3 biologically independent replicates, lines represent the mean ± s.d. In box plots (d and e) the central mark is the median, and the edges of the box are the first and third quartiles. Whiskers extend to the most extreme non-outlier data points.
Supplementary Figure 6 Phf5a silencing inhibits myotube differentiation through Paf1C destabilization.
(a) Brightfield images of self-renewing myoblasts or 72h-differentiated myotubes. (b) Western blot analysis of self-renewal and differentiation markers in C2C12 cells following knockdown of Paf1C subunits and 72h differentiation. (c) Myocin heavy chain (MHC) immunofluorescence on differentiated myotubes for 72h following shControl or shPhf5a knockdown. Scale bars, 100 μm. (d) Quantification of myocin heavy chain (MHC) immunofluorescence intensity of Rosa26rtTACol1a1TREshPhf5a primary myotubes following addition of doxycycline. n = 3 biologically independent replicates (see Supplementary Table 5). ∗∗p = 0.0001, two-sided Student’s t-test, values represent the mean ± s.d. (e) Relative expression of Phf5a and multiple Paf1C subunits using qRT-PCR showing higher levels of most of Paf1C subunits in primary mouse myotubes. n = 3 biologically independent replicates (see Supplementary Table 5). Phf5a: p = 0.0193, Paf1: p = 0.0001, Cdc73: p = 0.0031, Leo1: p = 0.0378, Wdr61: p = 0.0238, Rtf1: n.s, non-significant, p = 0.1103, two-sided Student’s t-test, values represent the mean ± s.d. (f) Desmin immunofluorescence on 72h differentiated myotubes following CRISPR-Cas9 depletion of Phf5a. Scale bars, 100 μm. (g) Western blot analysis of the differentiation marker myocin heavy chain following CRISPR-Cas9 depletion of Phf5a. (h–j) Gene Ontology (GO) analysis of Leo1-bound genes after ChIP-sequencing in myoblasts (h) and myotubes (i) under shControl conditions, or myotubes (j) after shPhf5a depletion, respectively.
Supplementary Figure 7 Scans of unprocessed key blots.
(a) Scans from Figure 1a (b) Scans from Figure 1i (c) Scans from Figure 2d (d) Scans from Figure 3b (e) Scans from Figure 3c (f) Scans from Figure 3d (g) Scans from Figure 3e (h) Scans from Figure 4a (i) Scans from Figure 5a (j) Scans from Figure 7a (k) Scans from Figure 7e (l) Scans from Figure 7j.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1522 kb)
Supplementary Table 1
Supplementary Information (XLSX 593 kb)
Supplementary Table 2
Supplementary Information (XLSX 14 kb)
Supplementary Table 3
Supplementary Information (XLSX 4003 kb)
Supplementary Table 4
Supplementary Information (XLSX 39 kb)
Supplementary Table 5
Supplementary Information (XLSX 58 kb)
Rights and permissions
About this article
Cite this article
Strikoudis, A., Lazaris, C., Trimarchi, T. et al. Regulation of transcriptional elongation in pluripotency and cell differentiation by the PHD-finger protein Phf5a. Nat Cell Biol 18, 1127–1138 (2016). https://doi.org/10.1038/ncb3424
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3424
This article is cited by
-
Integrative modeling of lncRNA-chromatin interaction maps reveals diverse mechanisms of nuclear retention
BMC Genomics (2023)
-
PHF5A facilitates the development and progression of gastric cancer through SKP2-mediated stabilization of FOS
Journal of Translational Medicine (2023)
-
PHF5A regulates the expression of the DOCK5 variant to promote HNSCC progression through p38 MAPK activation
Biology Direct (2023)
-
PHF5A is a potential diagnostic, prognostic, and immunological biomarker in pan-cancer
Scientific Reports (2023)
-
Overlapping roles of spliceosomal components SF3B1 and PHF5A in rice splicing regulation
Communications Biology (2021)