Abstract
Integrins have key functions in cell adhesion and migration. How integrins are dynamically relocalized to the leading edge in highly polarized migratory cells has remained unexplored. Here, we demonstrate that β1 integrin (known as PAT-3 in Caenorhabditis elegans), but not β3, is transported from the plasma membrane to the trans-Golgi network, to be resecreted in a polarized manner. This retrograde trafficking is restricted to the non-ligand-bound conformation of β1 integrin. Retrograde trafficking inhibition abrogates several β1-integrin-specific functions such as cell adhesion in early embryonic development of mice, and persistent cell migration in the developing posterior gonad arm of C. elegans. Our results establish a paradigm according to which retrograde trafficking, and not endosomal recycling, is the key driver for β1 integrin function in highly polarized cells. These data more generally suggest that the retrograde route is used to relocalize plasma membrane machinery from previous sites of function to the leading edge of migratory cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bonifacino, J. S. & Rojas, R. Retrograde transport from endosomes to the trans-Golgi network. Nat. Rev. Mol. Cell. Biol. 7, 568–579 (2006).
Johannes, L. & Popoff, V. Tracing the retrograde route in protein trafficking. Cell 135, 1175–1187 (2008).
Arighi, C. N., Hartnell, L. M., Aguilar, R. C., Haft, C. R. & Bonifacino, J. S. Role of the mammalian retromer in sorting of the cation-independent mannose 6-phosphate receptor. J. Cell Biol. 165, 123–133 (2004).
Seaman, M. N. Cargo-selective endosomal sorting for retrieval to the Golgi requires retromer. J. Cell Biol. 165, 111–122 (2004).
Mallard, F. et al. Early/recycling endosomes-to-TGN transport involves two SNARE complexes and a Rab6 isoform. J. Cell Biol. 156, 653–664 (2002).
Perez-Victoria, F. J., Mardones, G. A. & Bonifacino, J. S. Requirement of the human GARP complex for mannose 6-phosphate-receptor-dependent sorting of cathepsin D to lysosomes. Mol. Biol. Cell 19, 2350–2362 (2008).
Franch-Marro, X. et al. Wingless secretion requires endosome-to-Golgi retrieval of Wntless/Evi/Sprinter by the retromer complex. Nat. Cell Biol. 10, 170–177 (2008).
Zhang, D. et al. RAB-6.2 and the retromer regulate glutamate receptor recycling through a retrograde pathway. J. Cell Biol. 196, 85–101 (2012).
Duncan, J. R. & Kornfeld, S. Intracellular movement of two mannose 6-phosphate receptors: return to the Golgi apparatus. J. Cell Biol. 106, 617–628 (1988).
Sandvig, K., Torgersen, M. L., Engedal, N., Skotland, T. & Iversen, T. G. Protein toxins from plants and bacteria: probes for intracellular transport and tools in medicine. FEBS Lett. 584, 2626–2634 (2010).
Johannes, L. & Römer, W. Shiga toxins—from cell biology to biomedical applications. Nat. Rev. Microbiol. 8, 105–116 (2010).
Hynes, R. O. Integrins: bidirectional, allosteric signaling machines. Cell 110, 673–687 (2002).
Wolfenson, H., Lavelin, I. & Geiger, B. Dynamic regulation of the structure and functions of integrin adhesions. Dev. Cell 24, 447–458 (2013).
Huttenlocher, A. & Horwitz, A. R. Integrins in cell migration. Cold Spring Harb. Perspect. Biol. 3, a005074 (2011).
Lobert, V. H. et al. Ubiquitination of α5β1 integrin controls fibroblast migration through lysosomal degradation of fibronectin-integrin complexes. Dev. Cell 19, 148–159 (2010).
Arjonen, A., Alanko, J., Veltel, S. & Ivaska, J. Distinct recycling of active and inactive β1 integrins. Traffic 13, 610–625 (2012).
Jones, M. C., Caswell, P. T. & Norman, J. C. Endocytic recycling pathways: emerging regulators of cell migration. Curr. Opin. Cell Biol. 18, 549–557 (2006).
Roberts, M., Barry, S., Woods, A., van der Sluijs, P. & Norman, J. PDGF-regulated rab4-dependent recycling of αvβ3 integrin from early endosomes is necessary for cell adhesion and spreading. Curr. Biol. 11, 1392–1402 (2001).
Caswell, P. T. & Norman, J. C. Integrin trafficking and the control of cell migration. Traffic 7, 14–21 (2006).
Powelka, A. M. et al. Stimulation-dependent recycling of integrin β1 regulated by ARF6 and Rab11. Traffic 5, 20–36 (2004).
Pellinen, T. et al. Small GTPase Rab21 regulates cell adhesion and controls endosomal traffic of β1-integrins. J. Cell Biol. 173, 767–780 (2006).
Riggs, K. A. et al. Regulation of integrin endocytic recycling and chemotactic cell migration by syntaxin 6 and VAMP3 interaction. J. Cell Sci. 125, 3827–3839 (2012).
White, D. P., Caswell, P. T. & Norman, J. C. αvβ3 and α5β1 integrin recycling pathways dictate downstream Rho kinase signaling to regulate persistent cell migration. J. Cell Biol. 177, 515–525 (2007).
Johannes, L. & Shafaq-Zadah, M. SNAP-tagging the retrograde route. Methods Cell Biol. 118, 139–155 (2013).
Shi, G. et al. SNAP-tag based proteomics approach for studying retrograde transport. Traffic 13, 914–925 (2012).
Takada, Y., Huang, C. & Hemler, M. E. Fibronectin receptor structures in the VLA family of heterodimers. Nature 326, 607–609 (1987).
De Franceschi, N., Hamidi, H., Alanko, J., Sahgal, P. & Ivaska, J. Integrin traffic—the update. J. Cell Sci. 128, 839–852 (2015).
Schiller, H. B. et al. β1- and αv-class integrins cooperate to regulate myosin II during rigidity sensing of fibronectin-based microenvironments. Nat. Cell Biol. 15, 625–636 (2013).
Shattil, S. J., Kim, C. & Ginsberg, M. H. The final steps of integrin activation: the end game. Nat. Rev. Mol. Cell Biol. 11, 288–300 (2010).
Akiyama, S. K., Yamada, S. S., Chen, W. T. & Yamada, K. M. Analysis of fibronectin receptor function with monoclonal antibodies: roles in cell adhesion, migration, matrix assembly, and cytoskeletal organization. J. Cell Biol. 109, 863–875 (1989).
Mould, A. P., Garratt, A. N., Askari, J. A., Akiyama, S. K. & Humphries, M. J. Identification of a novel anti-integrin monoclonal antibody that recognises a ligand-induced binding site epitope on the β1 subunit. FEBS Lett. 363, 118–122 (1995).
Bazzoni, G., Shih, D. T., Buck, C. A. & Hemler, M. E. Monoclonal antibody 9EG7 defines a novel β1 integrin epitope induced by soluble ligand and manganese, but inhibited by calcium. J. Biol. Chem. 270, 25570–25577 (1995).
Takada, Y. & Puzon, W. Identification of a regulatory region of integrin β1 subunit using activating and inhibiting antibodies. J. Biol. Chem. 268, 17597–17601 (1993).
Ni, H., Li, A., Simonsen, N. & Wilkins, J. A. Integrin activation by dithiothreitol or Mn2+ induces a ligand-occupied conformation and exposure of a novel NH2-terminal regulatory site on the β1 integrin chain. J. Biol. Chem. 273, 7981–7987 (1998).
Wilcke, M. et al. Rab11 regulates the compartmentalization of early endosomes required for efficient transport from early endosomes to the trans-Golgi network. J. Cell Biol. 151, 1207–1220 (2000).
Schlaepfer, D. D., Hauck, C. R. & Sieg, D. J. Signaling through focal adhesion kinase. Prog. Biophys. Mol. Biol. 71, 435–478 (1999).
Giannone, G., Ronde, P., Gaire, M., Haiech, J. & Takeda, K. Calcium oscillations trigger focal adhesion disassembly in human U87 astrocytoma cells. J. Biol. Chem. 277, 26364–26371 (2002).
Bardin, S. et al. Phenotypic characterisation of RAB6A knockout mouse embryonic fibroblasts. Biol. Cell 10.1111/boc.201400083 (2015).
Moore, R., Tao, W., Smith, E. R. & Xu, X. X. The primitive endoderm segregates from the epiblast in β1 integrin-deficient early mouse embryos. Mol. Cell. Biol. 34, 560–572 (2014).
Fassler, R. & Meyer, M. Consequences of lack of β1 integrin gene expression in mice. Genes Dev. 9, 1896–1908 (1995).
Aumailley, M., Pesch, M., Tunggal, L., Gaill, F. & Fassler, R. Altered synthesis of laminin 1 and absence of basement membrane component deposition in β1 integrin-deficient embryoid bodies. J. Cell Sci. 113, 259–268 (2000).
Lohikangas, L., Gullberg, D. & Johansson, S. Assembly of laminin polymers is dependent on β1-integrins. Exp. Cell Res. 265, 135–144 (2001).
Yu, W. et al. Beta1-integrin orients epithelial polarity via Rac1 and laminin. Mol. Biol. Cell 16, 433–445 (2005).
Maiuri, P. et al. The first world cell race. Curr. Biol. 22, 673–675 (2012).
Pellinen, T. et al. Small GTPase Rab21 regulates cell adhesion and controls endosomal traffic of β1-integrins. J. Cell Biol. 173, 767–780 (2006).
Thery, M. et al. Anisotropy of cell adhesive microenvironment governs cell internal organization and orientation of polarity. Proc. Natl Acad. Sci. USA 103, 19771–19776 (2006).
Theisen, U., Straube, E. & Straube, A. Directional persistence of migrating cells requires Kif1C-mediated stabilization of trailing adhesions. Dev. Cell 23, 1153–1166 (2012).
Petrie, R. J., Doyle, A. D. & Yamada, K. M. Random versus directionally persistent cell migration. Nat. Rev. Mol. Cell. Biol. 10, 538–549 (2009).
White, D. P., Caswell, P. T. & Norman, J. C. αvβ3 and α5β1 integrin recycling pathways dictate downstream Rho kinase signaling to regulate persistent cell migration. J. Cell Biol. 177, 515–525 (2007).
Yadav, S., Puri, S. & Linstedt, A. D. A primary role for Golgi positioning in directed secretion, cell polarity, and wound healing. Mol. Biol. Cell 20, 1728–1736 (2009).
Darido, C. & Jane, S. M. Golgi feels its own wound. Adv. Wound Care 2, 87–92 (2013).
Lee, M., Cram, E. J., Shen, B. & Schwarzbauer, J. E. Roles for β(pat-3) integrins in development and function of Caenorhabditis elegans muscles and gonads. J. Biol. Chem. 276, 36404–36410 (2001).
Yang, P. T. et al. Wnt signaling requires retromer-dependent recycling of MIG-14/Wntless in Wnt-producing cells. Dev. Cell 14, 140–147 (2008).
Nishiwaki, K. Mutations affecting symmetrical migration of distal tip cells in Caenorhabditis elegans. Genetics 152, 985–997 (1999).
Ridley, A. J. et al. Cell migration: integrating signals from front to back. Science 302, 1704–1709 (2003).
Etienne-Manneville, S. Cdc42–the centre of polarity. J. Cell Sci. 117, 1291–1300 (2004).
Etienne-Manneville, S. & Hall, A. Integrin-mediated activation of Cdc42 controls cell polarity in migrating astrocytes through PKCzeta. Cell 106, 489–498 (2001).
Danen, E. H. et al. Integrins control motile strategy through a Rho-cofilin pathway. J. Cell Biol. 169, 515–526 (2005).
Bretscher, M. S. Circulating integrins: α5β1, α6β4 and Mac-1, but not α3β1, α4β1 or LFA-1. EMBO J. 11, 405–410 (1992).
Amessou, M. et al. Syntaxin 16 and syntaxin 5 control retrograde transport of several exogenous and endogenous cargo proteins. J. Cell. Sci. 120, 1457–1468 (2007).
Schauer, K., Duong, T., Gomes-Santos, C. S. & Goud, B. Studying intracellular trafficking pathways with probabilistic density maps. Methods Cell Biol. 118, 325–343 (2013).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nature Methods 9, 671–675 (2012).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Bolte, S. & Cordelieres, F. P. A guided tour into subcellular colocalization analysis in light microscopy. J. Microsc. 224, 213–232 (2006).
Dessau, R. B. & Pipper, C. B. “R”–project for statistical computing. Ugeskr. Laeger 170, 328–330 (2008).
Vielemeyer, O. et al. Characterization of single chain antibody targets through yeast two hybrid. BMC Biotechnol. 10, 59 (2010).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Kamath, R. S. & Ahringer, J. Genome-wide RNAi screening in Caenorhabditis elegans. Methods 30, 313–321 (2003).
Acknowledgements
We thank M. Piel, A. M. Lennon-Duménil and D. Bourc’his for helpful discussions. We acknowledge the following people for providing materials: J. Bonifacino (National Institutes of Health, Bethesda, USA) for anti-vps35 antibody (no. 764), K. Schauer (Institut Curie, Paris, France) for the RPE-1 GFP–Rab1 stable cell line, G. Michaux (Institut de Génétique et Développement de Rennes, France) and R. Legouis (Institut de Biologie Intégrative de la Cellule, Gif-sur-Yvette, France) for C. elegans reagents and materials, the CGC (University of Minnesota, USA) for providing strains. We thank the staff of the animal facility at Institut Curie for mouse breeding and crossing. C.S.G.-S. was financially supported by a Marie Curie Fellowship PIEF-GA-2011-299756. This work was supported by grants from the Agence Nationale pour la Recherche (ANR-11 BSV2 014 03 and ANR-14-CE14-0002-02 to L.J.), Human Frontier Science Program grant RGP0029-2014 to L.J., and by European Research Council advanced grants (project 340485 to L.J. and project 339847 ‘MYODYN’ to B.G.). We acknowledge the Recombinant Protein and Antibody Platform of the Institut Curie (http://umr144.curie.fr/en/plateform/protein-and-antibody-laboratory-001279) for the production of human recombinant antibodies against Rab6:GTP. The Johannes and Goud teams are members of Labex CelTisPhyBio (11-LBX-0038) and Idex Paris Sciences et Lettres (ANR-10-IDEX-0001-02 PSL). The facilities as well as scientific and technical assistance from staff in the PICT-IBiSA/Nikon Imaging Centre at Institut Curie-CNRS, Proteomics and Mass Spectrometry Laboratory, Institut Curie (Damarys Loew and Florent Dingli), Cell and Tissue Imaging Platform—PICT-IBiSA (member of France–Bioimaging), the Genetics and Developmental Biology Department (UMR3215/U934), and the France–BioImaging infrastructure (ANR-10-INSB-04) are acknowledged.
Author information
Authors and Affiliations
Contributions
M.S.-Z., C.S.G.-S., P.C., B.G. and L.J. designed the experiments and wrote the manuscript. M.S.-Z. and C.S.G.-S. performed experiments and analysed data. S.B. and J.I. contributed to animal experiments. P.M., M.M., A.G. and C.L. analysed data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 β1 integrin is transported to the Golgi.
(a) Schematic representation of the SNAP-tag strategy for vectorial proteomics. BG-modified plasma membrane proteins (BG-Prot-X) that reach the Golgi compartment are covalently captured by TGN-localized GalT-GFP-SNAP fusion protein. After GFP-trap pulldown, retrograde cargoes are identified by mass spectrometry (4 independent proteomics experiments). The table summarizes integrins that were found on the hit list (see Supplementary Table 1). (b) BG-modified anti-α5 integrin antibodies localize to Golgi in GalT-GFP-SNAP expressing HeLa cells, as opposed to BG-modified anti-β3 integrin antibodies. Representative images of 3 independent experiments for both β3 integrin and α5 integrin. (c) HeLa cells were colabeled with 12G10 antibody and BG-FN-50K-Cy3 to validate the activity of this fibronectin fragment after BG and Cy3 labeling. (d) HeLa cells were stimulated with TS2/16 antibody or with fibronectin/manganese (FN-Mn2 +), and labeled with non-ligand-bound β1 integrin conformation-specific antibody mAb13, or with ligand-bound conformation-specific antibodies 12G10 and 9EG7 (representative images from 5 independent experiments for TS2/16, and 2 independent experiments for FN-Mn2 +). The percentage of mAb13, 12G10 or 9EG7 signal in the Golgi compartment was then quantified (1 representative of 2 independent experiments. mAb13-No Stim. n = 13 cells; mAb13-Stim. n = 15 cells; mAb13-Mn2+ Stim. n = 16 cells; 12G10-No Stim. n = 10 cells; 12G10-Stim. n = 16 cells; 9EG7-No Stim. n = 22 cells; 9EG7-Stim. n = 30 cells). Insets show the Golgi compartment. Unpaired t-test. (e) HeLa cells were incubated with mAb13-BG antibody on ice, washed, incubated for 30 min at 37 °C in the absence, and then another 60 min in the presence of TS2/16. The formation of conjugates with GalT-GFP-SNAP was quantified as in Fig. 1b (n = 3 independent experiments). Paired t-test. (f) Colocalization analysis for α5/β1 integrins with the retromer subunits Snx6 or Vps26 (mean ± s.d. of n = 10 cells per condition, 1 representative of 2 independent experiments is shown). (g, h) Western blotting on HeLa cell lysates was used to determine total cellular levels of β1 integrin using TS2/16 antibody (g; means ± s.d., n = 5 independent experiments for ctrl siRNA and synt16 siRNA, n = 4 independent experiments for Rab6 siRNA), and FACS for cell surface levels (h; means ± s.d., n = 3 independent experiments, MFI = Mean Fluorescence Intensity) under conditions of inhibition of retrograde transport by depletion of syntaxin 16 or Rab6. Paired t-test. (i) Colocalization analysis of β1 integrin with LAMP1 on HeLa cells. Quantification by Manders’ coefficient (1 representative of 2 independent experiments. ctrl siRNA n = 14 cells, synt16 siRNA n = 18 cells, Rab6 siRNA = 17 cells)), mean ± s.d. For all images, scale bars = 10 μm. Unpaired t-test unless otherwise stated. ∗P < 0.05, ∗∗P < 0.01, ∗∗∗P < 0.001. Statistics source data can be found in Supplementary Table 4. Unprocessed blots can be found in Supplementary Fig. 7.
Supplementary Figure 2 Retrograde machinery is required for cell spreading and adhesion.
(a) Box-and-Whisker plots show RPE-1 cell spreading areas at different times after adhesion on fibronectin-coated coverslips (1 representative of 3 independent experiments. 20 min.: ctrl-siRNA n = 31 cells, Rab6-siRNA n = 29 cells; 2 h: ctrl-siRNA n = 12 cells, Rab6-siRNA n = 26 cells; 24 h: ctrl-siRNA n = 18 cells, Rab6-siRNA n = 22 cells). The ends of the Whiskers are set at 1.5× Interquartile Range above the third quartile, and 1.5× Interquartile Range below the first quartile). (b) Cell size in suspension measured by flow cytometry. Numbers of independent experiments: HeLa n = 4, RPE-1 n = 6. (c) Quantification by Western blotting of phosphorylation levels of focal adhesion kinase (pFAK) (n = 3 and 2 independent experiments for synt16 siRNA and Rab6 siRNA, respectively) in HeLa cells. (d) Percentage of RPE-1 cells labeled with the focal adhesion marker vinculin at different times after adhesion (n = 5 independent experiments). (e) Average number of RPE-1 cells adherent to fibronectin-coated coverslips per field of a ×10 objective, after 1 h of adhesion and 3 washes with PBS++ (1 representative of 2 independent experiments. ctrl-siRNA n = 13 fields, synt16-siRNA n = 19 fields, Rab6-siRNA n = 17 fields). (f) Number of RPE-1 cells after a 5 min incubation with trypsin (1 representative of 2 independent experiments. ctrl-siRNA n = 4 fields, synt16-siRNA n = 3 fields, Rab6-siRNA n = 5 fields). Means ± s.d., ∗P < 0.05, ∗∗P < 0.01, ∗∗∗P < 0.001, unpaired t-test.
Supplementary Figure 3 Rab6 KO mice show impaired early embryonic development.
(a) Organization of embryonic layers at 4.5 dpc or 6 dpc of gestation of Rab6+/− or Rab6−/− mouse embryos (representative images of 2 Rab6+/− and 3 Rab6−/− embryos at 4.5 dpc, and 6 Rab6+/− and 6 Rab6−/− embryos at 6 dpc). Scale bars = 20 μm for 4.5 dpc, 40 μm for 6 dpc. Nanog (epiblast marker), Gata4 (visceral endoderm marker). (b) 5.5 dpc wild-type embryos labeled with Rab6-GTP and β1 integrin-specific antibodies. Scale bar = 20 μm.
Supplementary Figure 4 Distribution of β1 integrin after retrograde machinery depletion.
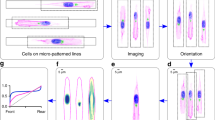
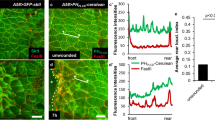
(a) Normalized mean cells showing distribution of both non-ligand-bound (mAb13) and ligand-bound (12G10) β1 integrin on HeLa or RPE-1 cells that were plated on crossbow-shaped micropatterns. Images represent n = 15 cells per condition (1 representative of 2 independent experiments). Scale bars = 10 μm. (b) Intensity distribution profiles and normalized images of mean cells showing the surface distribution of ligand-bound β1 integrin (12G10) on RPE-1 cells, migrating on fibronectin-coated micropatterned lines. n = 10 cells per condition in one representative experiment. mean ± s.d. Numbers of independent experiments: ctrl siRNA = 4, synt16 siRNA = 3, Rab6 siRNA = 3, and Rab11 siRNA = 2. Scale bar = 10μm.
Supplementary Figure 5 Retrograde trafficking is required for persistent cell migration.
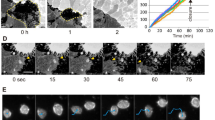
(a) Schematic representation of path persistence: ratio of effective maximum displacement, d, to actual trajectory length, D. (b) Speed distribution of RPE-1 cells migrating at least 50 μm in 2D random migration on fibronectin-coated coverslips (pooled cells from independent experiments are represented in the graph. Number of independent experiments: ctrl siRNA = 6 (n = 456 cells), synt16 siRNA = 3 (n = 58 cells), Rab6 siRNA = 5 (n = 361 cells), and Rab11 siRNA = 4 (n = 447 cells), or in 1D migration on fibronectin-coated micropatterned lines (pooled cells from independent experiments are represented in the graph. Number of independent experiments: ctrl siRNA = 5 (n = 836 cells), synt16 siRNA = 3 (n = 552 cells), Rab6 siRNA = 3 (n = 218 cells), and Rab11 siRNA = 2 (n = 507 cells). 10 representative trajectories for the 1D migration condition are shown (17 h of migration). Note the loss of track linearity for the syntaxin 16 and Rab6 depletion conditions. (c) RPE-1 cells were plated on fibronectin-coated non-micropatterned surface (culture dishes), or on fibronectin-coated line micropatterns before be fixed, permeabilized and immunostained with mAb13. Insets show the Golgi compartment. Scale bar = 10 μm. (d) Transmission microscopy images representing the edge progression in wound healing assays with RPE-1 cells Scale bar = 40 μm. Wound edge velocity is quantified. 1 representative of 3 independent experiments for ctrl siRNA and Rab6 siRNA and 2 independent experiments for synt16 siRNA are shown. ×10 objective fields of 2 wounds were quantified: ctrl siRNA n = 18 fields, synt16 siRNA n = 12 fields, Rab6-siRNA n = 13 fields, mean ± s.d.). (e) Box-and-Whiskers plots representing the migratory persistence of edge cells in wound healing assays. 1 representative of 3 independent experiments for ctrl siRNA, and 2 independent experiments for synt16 siRNA and Rab6 siRNA is shown. (ctrl siRNA n = 47 cells, synt16 siRNA n = 49 cells, Rab6 siRNA n = 50 cells). 10 representative trajectories are shown to the right. (f) Golgi polarization towards the wound, derived as the angle between the wound migration direction (defined as a line perpendicular to the wound, passing via the center of the nucleus) and the position of the Golgi, visualized by GFP-Rab1. 1 experiment. (g) Non-ligand-bound (mAb13) and ligand-bound (12G10) β1 integrin labeling of wound edge cells. For (d), data represent means ± s.d., for (b, e), two-tailed Mann-Whitney and for (d) unpaired t-test. ∗P < 0.05, ∗∗P < 0.01, ∗∗∗P < 0.001. The ends of the Whiskers are set at 1.5× Interquartile Range above the third quartile, and 1.5× Interquartile Range below the first quartile.
Supplementary Figure 6 Inhibition of the retrograde trafficking machinery impairs DTC migration in C. elegans.
(a) C. elegans gonad migration defects in rab-6.2(ok2254) and vps-35(hu68) mutants obtained from the Caenorhabditis Genetic Center (CGC). Representative images of 30 worms per condition from 2 independent experiments. Scale bar = 25 μm. (b) Schematic representation of type 2 and type 3 Gonad migration defects.
Supplementary Figure 7 Full scans of original immunoblots presented in this work.
a–d correspond to Fig. 1b–e, g, respectively. Blots were probed with anti-SNAP (a, b and d), anti-β1 integrin, anti-vps35, anti-vps26, anti-snx6 and anti-rab6 (c) or anti-α tubulin antibodies (d). e, fcorrespond to Supplementary Fig. 1e, g, respectively. Blots were probed with anti-SNAP (e), anti-β1 integrin and anti-α tubulin antibodies (f).
Supplementary information
Supplementary Information
Supplementary Information (PDF 3002 kb)
Supplementary Table 1
Supplementary Information (XLSX 53 kb)
Supplementary Table 2
Supplementary Information (XLSX 97 kb)
Supplementary Table 3
Supplementary Information (XLSX 12 kb)
Supplementary Table 4
Supplementary Information (XLSX 73 kb)
1D migration.
Video microscopy of 1D migration of RPE-1 cells plated on fibronectin-coated 9 μm width micropatterned lines. Scale bar = 50 μm, 10 frames per sec. (MOV 455 kb)
Wound healing assay.
Wound healing migration of RPE-1 cells. Scale bar = 50 μm, 10 frames per sec. (MOV 2960 kb)
Rights and permissions
About this article
Cite this article
Shafaq-Zadah, M., Gomes-Santos, C., Bardin, S. et al. Persistent cell migration and adhesion rely on retrograde transport of β1 integrin. Nat Cell Biol 18, 54–64 (2016). https://doi.org/10.1038/ncb3287
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3287
This article is cited by
-
Caveolin-1 dolines form a distinct and rapid caveolae-independent mechanoadaptation system
Nature Cell Biology (2023)
-
Drebrin controls scar formation and astrocyte reactivity upon traumatic brain injury by regulating membrane trafficking
Nature Communications (2021)
-
The EMT activator ZEB1 accelerates endosomal trafficking to establish a polarity axis in lung adenocarcinoma cells
Nature Communications (2021)
-
Cargo-specific recruitment in clathrin- and dynamin-independent endocytosis
Nature Cell Biology (2021)
-
Integrin trafficking in cells and tissues
Nature Cell Biology (2019)