Abstract
Peroxisomes are highly metabolic, autonomously replicating organelles that generate reactive oxygen species (ROS) as a by-product of fatty acid β-oxidation. Consequently, cells must maintain peroxisome homeostasis, or risk pathologies associated with too few peroxisomes, such as peroxisome biogenesis disorders, or too many peroxisomes, inducing oxidative damage and promoting diseases such as cancer. We report that the PEX5 peroxisome import receptor binds ataxia-telangiectasia mutated (ATM) and localizes this kinase to the peroxisome. In response to ROS, ATM signalling activates ULK1 and inhibits mTORC1 to induce autophagy. Specificity for autophagy of peroxisomes (pexophagy) is provided by ATM phosphorylation of PEX5 at Ser 141, which promotes PEX5 monoubiquitylation at Lys 209, and recognition of ubiquitylated PEX5 by the autophagy adaptor protein p62, directing the autophagosome to peroxisomes to induce pexophagy. These data reveal an important new role for ATM in metabolism as a sensor of ROS that regulates pexophagy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Reddy, J. K. & Hashimoto, T. Peroxisomal beta-oxidation and peroxisome proliferator-activated receptor alpha: an adaptive metabolic system. Annu. Rev. Nutri. 21, 193–230 (2001).
Schrader, M. & Fahimi, H. D. Peroxisomes and oxidative stress. Biochim. Biophys. Acta 1763, 1755–1766 (2006).
Alexander, A. et al. ATM signals to TSC2 in the cytoplasm to regulate mTORC1 in response to ROS. Proc. Natl Acad. Sci. USA 107, 4153–4158 (2010).
Chen, Y., Azad, M. B. & Gibson, S. B. Superoxide is the major reactive oxygen species regulating autophagy. Cell Death Differ. 16, 1040–1052 (2009).
Holmstrom, K. M. & Finkel, T. Cellular mechanisms and physiological consequences of redox-dependent signalling. Nat. Rev. Mol. Cell Biol. 15, 411–421 (2014).
Scherz-Shouval, R. et al. Reactive oxygen species are essential for autophagy and specifically regulate the activity of Atg4. EMBO J. 26, 1749–1760 (2007).
Purdue, P. E. & Lazarow, P. B. Peroxisome biogenesis. Annu. Rev. Cell Dev. Biol. 17, 701–752 (2001).
Santos, M. J. et al. Peroxisomal proliferation protects from beta-amyloid neurodegeneration. J. Biol. Chem. 280, 41057–41068 (2005).
Singh, I., Singh, A. K. & Contreras, M. A. Peroxisomal dysfunction in inflammatory childhood white matter disorders: an unexpected contributor to neuropathology. J. Child Neurol. 24, 1147–1157 (2009).
Smith, J. J. & Aitchison, J. D. Peroxisomes take shape. Nat. Rev. Mol. Cell Biol. 14, 803–817 (2013).
Weller, S., Gould, S. J. & Valle, D. Peroxisome biogenesis disorders. Annu. Rev. Genomics Hum. Genet. 4, 165–211 (2003).
Reuber, B. E. et al. Mutations in PEX1 are the most common cause of peroxisome biogenesis disorders. Nat. Genet. 17, 445–448 (1997).
Fransen, M., Nordgren, M., Wang, B. & Apanasets, O. Role of peroxisomes in ROS/RNS-metabolism: implications for human disease. Biochim. Biophys. Acta 1822, 1363–1373 (2012).
Dunn, W. A. Jr et al. Pexophagy: the selective autophagy of peroxisomes. Autophagy 1, 75–83 (2005).
Farre, J. C. & Subramani, S. Peroxisome turnover by micropexophagy: an autophagy-related process. Trends Cell Biol. 14, 515–523 (2004).
Iwata, J. et al. Excess peroxisomes are degraded by autophagic machinery in mammals. J. Biol. Chem. 281, 4035–4041 (2006).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 8, 445–544 (2012).
Till, A., Lakhani, R., Burnett, S. F. & Subramani, S. Pexophagy: the selective degradation of peroxisomes. Int. J. Cell Biol. 2012, 512721 (2012).
Kirkin, V., McEwan, D. G., Novak, I. & Dikic, I. A role for ubiquitin in selective autophagy. Mol. Cell 34, 259–269 (2009).
Kim, P. K., Hailey, D. W., Mullen, R. T. & Lippincott-Schwartz, J. Ubiquitin signals autophagic degradation of cytosolic proteins and peroxisomes. Proc. Natl Acad. Sci. USA 105, 20567–20574 (2008).
Guo, Z., Kozlov, S., Lavin, M. F., Person, M. D. & Paull, T. T. ATM activation by oxidative stress. Science 330, 517–521 (2010).
Lim, D. S. et al. ATM binds to beta-adaptin in cytoplasmic vesicles. Proc. Natl Acad. Sci. USA 95, 10146–10151 (1998).
Watters, D. et al. Localization of a portion of extranuclear ATM to peroxisomes. J. Biol. Chem. 274, 34277–34282 (1999).
Benjamin, D. & Hall, M. N. TSC on the peroxisome controls mTORC1. Nat. Cell Biol. 15, 1135–1136 (2013).
Zhang, J. et al. A tuberous sclerosis complex signalling node at the peroxisome regulates mTORC1 and autophagy in response to ROS. Nat. Cell Biol. 15, 1186–1196 (2013).
Kastan, M. B. & Bartek, J. Cell-cycle checkpoints and cancer. Nature 432, 316–323 (2004).
Kastan, M. B. & Lim, D. S. The many substrates and functions of ATM. Nat. Rev. Mol. Cell Biol. 1, 179–186 (2000).
Ma, C., Agrawal, G. & Subramani, S. Peroxisome assembly: matrix and membrane protein biogenesis. J. Cell Biol. 193, 7–16 (2011).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, 132–141 (2011).
Kim, S. T., Lim, D. S., Canman, C. E. & Kastan, M. B. Substrate specificities and identification of putative substrates of ATM kinase family members. J. Biol. Chem. 274, 37538–37543 (1999).
Brown, A. I., Kim, P. K. & Rutenberg, A. D. PEX5 and ubiquitin dynamics on mammalian peroxisome membranes. PLoS Comput. Biol. 10, e1003426 (2014).
Carvalho, A. F. et al. Ubiquitination of mammalian Pex5p, the peroxisomal import receptor. J. Biol. Chem. 282, 31267–31272 (2007).
Grou, C. P. et al. Members of the E2D (UbcH5) family mediate the ubiquitination of the conserved cysteine of Pex5p, the peroxisomal import receptor. J. Biol. Chem. 283, 14190–14197 (2008).
Platta, H. W. et al. Pex2 and pex12 function as protein-ubiquitin ligases in peroxisomal protein import. Mol. Cell. Biol. 29, 5505–5516 (2009).
Thoms, S. & Erdmann, R. Peroxisomal matrix protein receptor ubiquitination and recycling. Biochim. Biophys. Acta 1763, 1620–1628 (2006).
Wagner, S. A. et al. A proteome-wide, quantitative survey of in vivo ubiquitylation sites reveals widespread regulatory roles. Mol. Cell. Proteomic. 10, M111.013284 (2011).
Hunter, T. The age of crosstalk: phosphorylation, ubiquitination, and beyond. Mol. Cell 28, 730–738 (2007).
Miyata, N. & Fujiki, Y. Shuttling mechanism of peroxisome targeting signal type 1 receptor Pex5: ATP-independent import and ATP-dependent export. Mol. Cell. Biol. 25, 10822–10832 (2005).
Fujita, N. et al. Recruitment of the autophagic machinery to endosomes during infection is mediated by ubiquitin. J. Cell Biol. 203, 115–128 (2013).
Gammoh, N., Florey, O., Overholtzer, M. & Jiang, X. Interaction between FIP200 and ATG16L1 distinguishes ULK1 complex-dependent and -independent autophagy. Nat. Struct. Mol. Biol. 20, 144–149 (2013).
Nazarko, T. Y. et al. Peroxisomal Atg37 binds Atg30 or palmitoyl-CoA to regulate phagophore formation during pexophagy. J. Cell Biol. 204, 541–557 (2014).
Stolz, A., Ernst, A. & Dikic, I. Cargo recognition and trafficking in selective autophagy. Nat. Cell Biol. 16, 495–501 (2014).
Chen, Y. & Dorn, G. W. 2nd PINK1-phosphorylated mitofusin 2 is a Parkin receptor for culling damaged mitochondria. Science 340, 471–475 (2013).
Geisler, S. et al. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat. Cell Biol. 12, 119–131 (2010).
Shiba-Fukushima, K. et al. PINK1-mediated phosphorylation of the Parkin ubiquitin-like domain primes mitochondrial translocation of Parkin and regulates mitophagy. Sci. Rep. 2, 1002 (2012).
Green, D. R. & Levine, B. To be or not to be? How selective autophagy and cell death govern cell fate. Cell 157, 65–75 (2014).
Apanasets, O. et al. PEX5, the shuttling import receptor for peroxisomal matrix proteins, is a redox-sensitive protein. Traffic 15, 94–103 (2014).
Ma, C., Hagstrom, D., Polley, S. G. & Subramani, S. Redox-regulated cargo binding and release by the peroxisomal targeting signal receptor, Pex5. J. Biol. Chem. 288, 27220–27231 (2013).
Hara-Kuge, S. & Fujiki, Y. The peroxin Pex14p is involved in LC3-dependent degradation of mammalian peroxisomes. Exp. Cell Res. 314, 3531–3541 (2008).
Deosaran, E. et al. NBR1 acts as an autophagy receptor for peroxisomes. J. Cell Sci. 126, 939–952 (2013).
Gonzalez, F. J. & Shah, Y. M. PPARα: mechanism of species differences and hepatocarcinogenesis of peroxisome proliferators. Toxicology 246, 2–8 (2008).
Tripathi, D. N. et al. Reactive nitrogen species regulate autophagy through ATM-AMPK-TSC2-mediated suppression of mTORC1. Proc. Natl Acad. Sci. USA 110, E2950-2957 (2013).
Ditch, S. & Paull, T. T. The ATM protein kinase and cellular redox signaling: beyond the DNA damage response. Trends Biochem. Sci. 37, 15–22 (2012).
Barlow, C. et al. ATM is a cytoplasmic protein in mouse brain required to prevent lysosomal accumulation. Proc. Natl Acad. Sci. USA 97, 871–876 (2000).
Valentin-Vega, Y. A. et al. Mitochondrial dysfunction in ataxia-telangiectasia. Blood 119, 1490–1500 (2012).
Zhou, B. B. & Bartek, J. Targeting the checkpoint kinases: chemosensitization versus chemoprotection. Nat. Rev. Cancer 4, 216–225 (2004).
Cai, S. L. et al. Activity of TSC2 is inhibited by AKT-mediated phosphorylation and membrane partitioning. J. Cell. Biol. 173, 279–289 (2006).
Pandita, R. K. et al. Mammalian Rad9 plays a role in telomere stability, S- and G2-phase-specific cell survival, and homologous recombinational repair. Mol. Cell. Biol. 26, 1850–1864 (2006).
Hunt, C. R. et al. Hyperthermia activates a subset of ataxia-telangiectasia mutated effectors independent of DNA strand breaks and heat shock protein 70 status. Cancer Res. 67, 3010–3017 (2007).
Gupta, A. et al. MOF phosphorylation by ATM regulates 53BP1-mediated double-strand break repair pathway choice. Cell Rep. 8, 177–189 (2014).
Dhar, S., Squire, J. A., Hande, M. P., Wellinger, R. J. & Pandita, T. K. Inactivation of 14-3-3sigma influences telomere behavior and ionizing radiation-induced chromosomal instability. Mol. Cell. Biol. 20, 7764–7772 (2000).
Acknowledgements
We thank S. Subramani (University of California, San Diego, California) for critical advice and providing the mRFP–EGFP–SKL plasmids, and M. Mancini (Baylor College of Medicine, Houston, Texas for providing the DsRed–SKL plasmid. We are also grateful for the assistance of K. Dunner (U.T.M.D. Cancer Center, Houston, Texas) and D. Townley (Baylor College of Medicine, Houston, Texas) in electron microscopy image acquisition. This work was supported by National Institutes of Health (NIH) Grant R01 CA143811 to C.L.W., a Robert A. Welch Endowed Chair in Chemistry (BE-0023) to C.L.W., and NIH grants CA129537, CA154320 and GM109768 to T.K.P. J.J. was a recipient of the China Scholarship Council (CSC).
Author information
Authors and Affiliations
Contributions
J.Z., D.N.T., J.J. and C.L.W. designed research; J.Z., D.N.T., J.J., J.K., A.A., R.T.P., R.D., J.T.-M., J.-H.L., R.K.P. and V.K.C. performed research; J.Z., D.N.T., J.J., R.D., T.T.P., T.K.P., M.B.K. and C.L.W. analysed data; J.Z., D.N.T. and C.L.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 2 Response of ATM RQ mutant to oxidative and DNA damage.
(a) Subcellular fractionation of Zellweger (GM13267) fibroblasts demonstrating the localization of phospho-ATM (S1981) and ATM in response to H2O2 (0.4 mM, 1 h) in the indicated subcellular compartments. LDH, Lamin A/C and β-integrin were used as markers for cytosol (C), nuclear (N) and membrane (M) fractions, respectively. WCE, whole cell extract. (b) AT (GM05849) fibroblasts were transfected with either wild type (WT) or peroxisome localization-deficient R3047Q (RQ) mutant Flag-tagged ATM and treated with H2O2 (0.4 mM) for 1 h. ATM was immunoprecipitated using anti-Flag antibody and activation determined using phosphoserine S1981 specific antibody. (c and d) ATM kinase assay with MR complex, Nbs1, and DNA were performed with 0.2 nM dimeric ATM, 50 nM GST-p53 substrate, 96 nM MR, 195 nM Nbs1, and 140 nM linear double stranded DNA. Phosphorylation of ATM was detected by SDS-PAGE and western blotting with an antibody to p53 phospho-Ser15. (e) Frequency of γ-H2AX foci in AT cells (AT5, GM05849) and HepG2 cells + siATM at different time points post irradiation of 2 Gy. 100 cells were analyzed for each time point in each experiment (Mean ± s.d., n = 3 independent experiments, student’s t test). The mean number of foci is plotted against time. ∗P < 0.05, ∗∗P < 0.01. (f) Representative figures of γ-H2AX foci. A: Untreated cells; B: AT5 (GM05849) + ATM cells 30 min post irradiation; C: AT5 cells 30 min post irradiation; D: AT5 cells 180 min post irradiation; E: AT5 + ATM cells 180 min post irradiation; F: AT5 + ATM-R3047Q cells 180 min post irradiation. Scale bar, 10 μm. (g) Representative images of chromosome aberrations at metaphase. A: AT5 + ATM control metaphase; B: AT5 + ATM showing dicentric (big arrow) and breaks (small arrow); C: AT5 showing radials (big arrow) and breaks (small arrow); D: AT5 + ATM-R3047Q showing radials (big arrow) and breaks (small arrow). (h) Chromosome aberrations at metaphase post irradiation of 3 Gy in AT cells (AT5, GM05849) and HepG2 cells + siATM. For each group 50 metaphases were scored in each experiment (mean ± s.d., n = 3 independent experiments, student’s t test). ∗∗P < 0.01. Uncropped images of western blots are shown in Supplementary Fig. 9. Statistical source data for Supplementary Fig. 1e, h can be found in Supplementary Table 1.
Supplementary Figure 3 Peroxisomal activation of ATM signaling to AMPK-TSC2 to repress mTORC1 and induce autophagy.
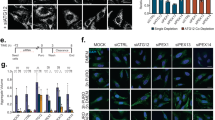
(a) FAO cells treated with clofibrate (0.25 mM) for 48 h activated PPARα to increase the mRNA level of EHHADH by RT–PCR analysis. The data shown is representative of two independent experiments. (b) FAO cells treated with clofibrate (0.25 mM) for 48 h activated PPARα to increase the ACAA1 mRNA levels as demonstrated by RT–PCR analysis. (mean ± s.d., n = 3 independent experiments, Student’s t test). ∗P < 0.05, ∗∗P < 0.01. (c) Western analysis of FAO cells treated with clofibrate at 0.25 mM for indicated times with ACAA1 and EHHADH antibodies. (d) MCF7 cells stably transfected with GFP–LC3 were treated with clofibrate (0.25 mM) for indicated times, and activation of ATM-AMPK-TSC2 signaling and induction of autophagy monitored by western analysis using pATM (S1981), ATM, pAMPK (T172), AMPK, pACC (S79), ACC, pS6K (T389), S6K, pS6 (S235/236), S6, p62 and LC3. (e) Representative images of GFP–LC3 puncta from MCF7 cells stably expressing GFP–LC3 and treated with clofibrate (0.25 mM) for 6 h. Scale bar, 10 μm. (f) HepG2 cells treated with 1 mM clofibrate for indicated times, were monitored for ATM-AMPK-TSC2 signaling, suppression of mTORC1, and induction of autophagy by western analysis using pATM (S1981), ATM, pAMPK (T172), AMPK, pACC (S79), ACC, pS6K (T389), S6K, pS6 (S235/236), S6, p4E-BP1 (T37/46), 4E-BP1, pULK1 (S757), pULK1 (S317), ULK1, p62 and LC3. (g) TSC2+/+ and TSC2−/− MEF cells were treated with clofibrate for 6 h at the indicated doses, and suppression of mTORC1 monitored by western analysis using pS6 (235/236), and S6. (h) Western analysis of FAO cells pre-incubated in the presence or absence of Bafilomycin A1 (Baf A1, 200 nM) for 1 h before treatment with 1 mM clofibrate for 3 h using anti-p62 and LC3-II antibodies. Uncropped images of western blots are shown in Supplementary Fig. 9, statistic source data for Supplementary Fig. 2a, b can be found in Supplementary Table 1.
Supplementary Figure 4 Pexophagy in FAO cells with clofibrate.
Representative electron microscopy images of FAO cells treated with vehicle or clofibrate. (a) vehicle 6 h, (b) clofibrate 6 h (1 mM) (c) vehicle 24 h, (d) clofibrate 24 h (0.25 mM), (e) clofibrate 48 h (0.25 mM), and (f) clofibrate 72 h (0.25 mM). Scale bar, 0.5 μm. Autophagosomes containing peroxisomes are represented by yellow arrows and shown enlarged in the boxed area (Scale bar, 0.2 μm).
Supplementary Figure 5 PEX5 contains ATM S*/T*Q phosphorylation motif.
(a) Prediction of PEX5 phosphorylation site and ubiquination site by ScanSite and PhosphoSite. (b) Conserved S141 (ATM phosphorylation site) sequence in PEX5. (c) WT fibroblasts (GM08399) and AT fibroblasts (GM05849) were transfected with Myc-PEX5-WT and treated with H2O2 for 1 h. The complex was immunoprecipitated with anti-Myc and immunoblotted with phospho-(S/T) ATM substrate antibody. Inputs were immunoblotted using the indicated antibodies. Uncropped images of western blots are shown in Supplementary Fig. 9.
Supplementary Figure 6 ROS-induced PEX5 ubiquitination is ATM-dependent.
(a) HEK293 cells expressing Myc-PEX5-WT or PEX5 mutants (K28R, K52R, K170R, K204R, K209R, K292R) and HA-Ub constructs were co-immunoprecipitated using anti-HA and blotting with an anti-Myc antibody. Input lysates were immunoblotted using the indicated antibodies. (b) WT (GM08399) and AT (GM05849) fibroblasts were treated with H2O2 for 6 h and immunoprecipitated with anti-PEX5 and immunoblotted with anti-Ub antibodies. Input lysates were immunoblotted using the indicated antibodies. (c) HEK293 cells transfected with Myc-PEX5-WT and HA-Ub for 24 h following a prior siRNA knockdown of ATM for 48 h, were treated with H2O2 for 6 h and immunoprecipitated with anti-HA and immunoblotted with anti-Myc antibodies. Input lysates were immunoblotted using the indicated antibodies. (d) HEK 293 cells were transfected with SiRNA for PEX2, PEX10 and PEX12, and treated with H2O2 for 6 h. Immunoprecipitation was performed with an anti-PEX5 antibody followed by immunoblotting for endogenous ubiqintin. RT- PCR was performed to confirm siRNA knockdown of PEX5, PEX10 and PEX12. (e) HEK293 cells transfected with Myc-PEX5-WT/Myc-PEX5-K209R and HA-Ub constructs for 24 h following a prior siRNA knockdown of PEX2/10/12 for 48 h were immunoprecipitated with anti-HA and immunoblotted with anti-Myc antibodies. Input lysates were immunoblotted using the indicated antibodies. Uncropped images of western blots are shown in Supplementary Fig. 9.
Supplementary Figure 7 K209 is responsible for PEX5 interaction with p62 in response to ROS.
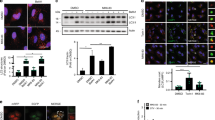
(a) Representative image of HepG2 cells treated with or without H2O2 for 3 h and immunostained for FIP200 or ATG16L (green) and peroxisomal marker PMP70 (red). Scale bar, 10 μm. High-magnification images of boxed areas showed co-localization between FIP200 or ATG16L and PMP70 (Scale bar, 3 μm). (b) HEK293 cells were transfected with Myc-PEX5-WT and HA-p62, and treated with 0.4 mM of H2O2 for indicated times. The complex of HA-p62 and Myc-PEX5 was immunoprecipitated with anti-HA and immunoblotted with anti-Myc. Inputs were immunoblotted using the indicated antibodies. (c) HEK293 cells were transfected with HA-p62 and Myc-PEX5-WT or Myc-PEX5-K209R. The complex of HA-p62 and Myc-PEX5 was immunoprecipitated with anti-HA and immunoblotted with anti-Myc antibodies. Input lysates were immunoblotted using the indicated antibodies. Uncropped images of western blots are shown in Supplementary Fig. 9.
Supplementary Figure 8 Regulation of pexophagy by ROS is ATM-dependent.
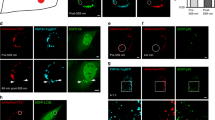
(a) Frame capture from time course imaging of pexophagy in live RPE cells stably expressing GFP–LC3 and DsRed-SKL following H2O2 (0.4 mM) treatment. Scale bar, 10 μm. (b) Western analysis of HepG2 cells pre-incubated in the presence or absence of Bafilomycin A1 (Baf A1, 200 nM) for 1 h before treatment with 0.4 mM H2O2 for 3 h using anti-PEX1, anti-PEX14, anti-p62 and LC3-II antibodies. (c) Quantification of PEX1 and PEX14 intensity normalized to GAPDH from Supplementary Fig. 7b (mean ± s.d., n = 3 independent experiments, Student’s t test). ∗P < 0.05, ∗∗P < 0.01, ∗∗∗P < 0.001, NS = not significant. (d) Western analysis of whole cell extracts (WCE) and peroxisome fractions (P) of HepG2 cells treated with Baf A1 (200 nM) for 4 h immunoblotted using anti-ATM, anti-pPEX5(S141) and PEX5 antibodies. (e) Western analysis of HepG2 cells transfected with or without sip62 for 72 h and treated with 0.4 mM H2O2 for 3 h using anti-PEX1, anti-PEX14, anti-pATM, anti-ATM, anti-p62 and LC3-II antibodies. (f) Quantification of PEX1 and PEX14 intensity normalized to GAPDH from Supplementary Fig. 7e (mean ± s.d., n = 3 independent experiments, Student’s t test). ∗∗P < 0.01, ∗∗∗P < 0.001, NS = not significant. (g) Representative images of wild type (WT, GM08399) and ataxia telengiectasia (AT, GM05849) patient fibroblasts treated with H2O2 for 6 h, and immunostained with PEX14 antibody. Scale bar, 15 μm. (h) Quantification of PEX14 puncta per cell was performed from Supplementary Fig. 7g from n = 4 independent experiments, 100 cells were analyzed in each experiment. All error bars represent s.d., (Student’s t test) ∗∗∗P < 0.001, NS = not significant. (i) Western analysis of AT (GM05849) fibroblasts transfected with Flag-ATM WT or Flag-ATM KD (kinase dead) mutant and treated with 0.4 mM H2O2 for 6 h using anti-PEX1, anti-PEX14, anti-pATM, anti-Flag and GAPDH antibodies. (j) Western analysis of HEK293 cells transfected with Pex2/10/12 siRNA for 72 h and treated with H2O2 (0.4 mM) for 6 h using anti-PEX1, anti-PEX14, anti-pATM, ATM and GAPDH antibodies. Uncropped images of western blots are shown in Supplementary Fig. 9. Statistic source data for Supplementary Fig. 7c, f, h can be found in Supplementary Table 1.
Supplementary Figure 9 Pexophagy in HepG2 cells with H2O2.
Representative electron microscopy images of HepG2 cells treated with vehicle or H2O2. (a) Vehicle 6 h, (b,c and d) H2O2 6 h (0.4 mM). Scale bar, 0.5 μm. Autophagosomes containing peroxisome are represented by yellow arrows, and shown enlarged in the boxed area (Scale bar, 0.25 μm).
Supplementary information
Supplementary Information
Supplementary Information (PDF 2796 kb)
Formation of autophagosomes over peroxisome.
RPE cells stably expressing GFP–LC3 and DsRed-SKL and treated with H2O2 (0.4 mM, from 30 min to 3 h) imaged by time-lapse microcopy. (AVI 188 kb)
Induction of pexophagic flux with H2O2.
MCF7 cells transfected with mRFP-EGFP-SKL and treated with H2O2 (0.4 mM, from 30 min to 3 h) imaged by time-lapse microcopy. (AVI 873 kb)
Rights and permissions
About this article
Cite this article
Zhang, J., Tripathi, D., Jing, J. et al. ATM functions at the peroxisome to induce pexophagy in response to ROS. Nat Cell Biol 17, 1259–1269 (2015). https://doi.org/10.1038/ncb3230
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3230
This article is cited by
-
PEX26 Functions as a Metastasis Suppressor in Colorectal Cancer
Digestive Diseases and Sciences (2024)
-
The peroxisome: an update on mysteries 3.0
Histochemistry and Cell Biology (2024)
-
Extracellular release in the quality control of the mammalian mitochondria
Journal of Biomedical Science (2023)
-
Autophagy and autophagy-related pathways in cancer
Nature Reviews Molecular Cell Biology (2023)
-
Cyclers’ kinases in cell division: from molecules to cancer therapy
Cell Death & Differentiation (2023)