Abstract
Matrix stiffness potently regulates cellular behaviour in various biological contexts. In breast tumours, the presence of dense clusters of collagen fibrils indicates increased matrix stiffness and correlates with poor survival. It is unclear how mechanical inputs are transduced into transcriptional outputs to drive tumour progression. Here we report that TWIST1 is an essential mechanomediator that promotes epithelial–mesenchymal transition (EMT) in response to increasing matrix stiffness. High matrix stiffness promotes nuclear translocation of TWIST1 by releasing TWIST1 from its cytoplasmic binding partner G3BP2. Loss of G3BP2 leads to constitutive TWIST1 nuclear localization and synergizes with increasing matrix stiffness to induce EMT and promote tumour invasion and metastasis. In human breast tumours, collagen fibre alignment, a marker of increasing matrix stiffness, and reduced expression of G3BP2 together predict poor survival. Our findings reveal a TWIST1–G3BP2 mechanotransduction pathway that responds to biomechanical signals from the tumour microenvironment to drive EMT, invasion and metastasis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Paszek, M. J. et al. Tensional homeostasis and the malignant phenotype. Cancer Cell 8, 241–254 (2005).
Levental, K. R. et al. Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139, 891–906 (2009).
Jaalouk, D. E. & Lammerding, J. Mechanotransduction gone awry. Nat. Rev. Mol. Cell Biol. 10, 63–73 (2009).
Calvo, F. et al. Mechanotransduction and YAP-dependent matrix remodelling is required for the generation and maintenance of cancer-associated fibroblasts. Nat. Cell Biol. 15, 637–646 (2013).
Butcher, D. T., Alliston, T. & Weaver, V. M. A tense situation: forcing tumour progression. Nat. Rev. Cancer 9, 108–122 (2009).
Colpaert, C., Vermeulen, P., Van Marck, E. & Dirix, L. The presence of a fibrotic focus is an independent predictor of early metastasis in lymph node-negative breast cancer patients. Am. J. Surg. Pathol. 25, 1557–1558 (2001).
Hasebe, T. et al. Prognostic significance of fibrotic focus in invasive ductal carcinoma of the breast: a prospective observational study. Mod. Pathol. 15, 502–516 (2002).
Conklin, M. W. et al. Aligned collagen is a prognostic signature for survival in human breast carcinoma. Am. J. Pathol. 178, 1221–1232 (2011).
Engler, A. J., Humbert, P. O., Wehrle-Haller, B. & Weaver, V. M. Multiscale modeling of form and function. Science 324, 208–212 (2009).
DuFort, C. C., Paszek, M. J. & Weaver, V. M. Balancing forces: architectural control of mechanotransduction. Nat. Rev. Mol. Cell Biol. 12, 308–319 (2011).
Hoffman, B. D., Grashoff, C. & Schwartz, M. A. Dynamic molecular processes mediate cellular mechanotransduction. Nature 475, 316–323 (2011).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Yang, J. & Weinberg, R. A. Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev. Cell 14, 818–829 (2008).
Thiery, J. P., Acloque, H., Huang, R. Y. & Nieto, M. A. Epithelial-mesenchymal transitions in development and disease. Cell 139, 871–890 (2009).
Yang, J. et al. Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell 117, 927–939 (2004).
Fang, X. et al. Twist2 contributes to breast cancer progression by promoting an epithelial–mesenchymal transition and cancer stem-like cell self-renewal. Oncogene 30, 4707–4720 (2011).
Batlle, E. et al. The transcription factor snail is a repressor of E-cadherin gene expression in epithelial tumour cells. Nat. Cell Biol. 2, 84–89 (2000).
Cano, A. et al. The transcription factor snail controls epithelial–mesenchymal transitions by repressing E-cadherin expression. Nat. Cell Biol. 2, 76–83 (2000).
Hajra, K. M., Chen, D. Y. & Fearon, E. R. The SLUG zinc-finger protein represses E-cadherin in breast cancer. Cancer Res. 62, 1613–1618 (2002).
Comijn, J. et al. The two-handed E box binding zinc finger protein SIP1 downregulates E-cadherin and induces invasion. Mol. Cell 7, 1267–1278 (2001).
Eger, A. et al. ΔEF1 is a transcriptional repressor of E-cadherin and regulates epithelial plasticity in breast cancer cells. Oncogene 24, 2375–2385 (2005).
Eckert, M. A. et al. Twist1-induced invadopodia formation promotes tumor metastasis. Cancer Cell 19, 372–386 (2011).
Desprat, N., Supatto, W., Pouille, P. A., Beaurepaire, E. & Farge, E. Tissue deformation modulates twist expression to determine anterior midgut differentiation in Drosophila embryos. Dev. Cell 15, 470–477 (2008).
Johnson, K. R., Leight, J. L. & Weaver, V. M. Demystifying the effects of a three-dimensional microenvironment in tissue morphogenesis. Methods Cell Biol. 83, 547–583 (2007).
Bissell, M. J., Radisky, D. C., Rizki, A., Weaver, V. M. & Petersen, O. W. The organizing principle: microenvironmental influences in the normal and malignant breast. Differentiation 70, 537–546 (2002).
Lee, G. Y., Kenny, P. A., Lee, E. H. & Bissell, M. J. Three-dimensional culture models of normal and malignant breast epithelial cells. Nat. Methods 4, 359–365 (2007).
Debnath, J., Muthuswamy, S. K. & Brugge, J. S. Morphogenesis and oncogenesis of MCF-10A mammary epithelial acini grown in three-dimensional basement membrane cultures. Methods 30, 256–268 (2003).
Xu, Y. et al. Inducible knockout of Twist1 in young and adult mice prolongs hair growth cycle and has mild effects on general health, supporting Twist1 as a preferential cancer target. Am. J. Pathol. 183, 1281–1292 (2013).
Blick, T. et al. Epithelial mesenchymal transition traits in human breast cancer cell lines. Clin. Exp. Metastasis 25, 629–642 (2008).
Tran, D. D., Corsa, C. A., Biswas, H., Aft, R. L. & Longmore, G. D. Temporal and spatial cooperation of Snail1 and Twist1 during epithelial–mesenchymal transition predicts for human breast cancer recurrence. Mol. Cancer Res. 9, 1644–1657 (2011).
Provenzano, P. P. et al. Collagen reorganization at the tumor-stromal interface facilitates local invasion. BMC Med. 4, 38 (2006).
Xu, J., Lamouille, S. & Derynck, R. TGF-β-induced epithelial to mesenchymal transition. Cell Res. 19, 156–172 (2009).
Leight, J. L., Wozniak, M. A., Chen, S., Lynch, M. L. & Chen, C. S. Matrix rigidity regulates a switch between TGF-β1-induced apoptosis and epithelial-mesenchymal transition. Mol. Biol. Cell 23, 781–791 (2012).
Friedland, J. C., Lee, M. H. & Boettiger, D. Mechanically activated integrin switch controls α5β1 function. Science 323, 642–644 (2009).
Zhao, B. et al. Inactivation of YAP oncoprotein by the Hippo pathway is involved in cell contact inhibition and tissue growth control. Genes Dev. 21, 2747–2761 (2007).
Kudo, N. et al. Leptomycin B inactivates CRM1/exportin 1 by covalent modification at a cysteine residue in the central conserved region. Proc. Natl Acad. Sci. USA 96, 9112–9117 (1999).
Singh, S. & Gramolini, A. O. Characterization of sequences in human TWIST required for nuclear localization. BMC Cell Biol. 10, 47 (2009).
Kim, M. M., Wiederschain, D., Kennedy, D., Hansen, E. & Yuan, Z. M. Modulation of p53 and MDM2 activity by novel interaction with Ras-GAP binding proteins (G3BP). Oncogene 26, 4209–4215 (2007).
Prigent, M., Barlat, I., Langen, H. & Dargemont, C. IκBα and IκBα /NF-κ B complexes are retained in the cytoplasm through interaction with a novel partner, RasGAP SH3-binding protein 2. J. Biol. Chem. 275, 36441–36449 (2000).
Wu, H. Y. et al. Combining alkaline phosphatase treatment and hybrid linear ion trap/Orbitrap high mass accuracy liquid chromatography-mass spectrometry data for the efficient and confident identification of protein phosphorylation. Anal. Chem. 81, 7778–7787 (2009).
Casas, E. et al. Snail2 is an essential mediator of Twist1-induced epithelial mesenchymal transition and metastasis. Cancer Res. 71, 245–254 (2011).
Miller, F. R., Santner, S. J., Tait, L. & Dawson, P. J. MCF10DCIS.com xenograft model of human comedo ductal carcinoma in situ. J. Natl Cancer Inst. 92, 1185–1186 (2000).
Kakkad, S. M. et al. Collagen I fiber density increases in lymph node positive breast cancers: pilot study. J. Biomed. Opt. 17, 116017 (2012).
Provenzano, P. P. et al. Collagen density promotes mammary tumor initiation and progression. BMC Med. 6, 11 (2008).
Alexander, N. R. et al. N-cadherin gene expression in prostate carcinoma is modulated by integrin-dependent nuclear translocation of Twist1. Cancer Res. 66, 3365–3369 (2006).
Chaudhuri, T., Rehfeldt, F., Sweeney, H. L. & Discher, D. E. Preparation of collagen-coated gels that maximize in vitro myogenesis of stem cells by matching the lateral elasticity of in vivo muscle. Methods Mol. Biol. 621, 185–202 (2010).
Klenova, E., Chernukhin, I., Inoue, T., Shamsuddin, S. & Norton, J. Immunoprecipitation techniques for the analysis of transcription factor complexes. Methods 26, 254–259 (2002).
Guo, Y., Ma, S. F., Grigoryev, D., Van Eyk, J. & Garcia, J. G. 1-DE MS and 2-D LC-MS analysis of the mouse bronchoalveolar lavage proteome. Proteomics 5, 4608–4624 (2005).
Artimo, P. et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 40, W597–W603 (2012).
Acknowledgements
We thank members of the Yang laboratory, especially M. Eckert, for helpful discussions. We thank the UCSD Shared Microscope Facility (P30NS047101), the UCSD Cancer Center Support Grant P30CA23100, and the NCI Cancer Diagnosis Program (CDP) for providing breast tumour tissue microarrays. The shRFP control pLKO.1 plasmid was a kind gift from S. Stewart (Washington University in St Louis, USA). This work was supported by grants from NIH (DP2OD002420-01, 1RO1CA168689), DOD Breast Cancer Program W81XWH-13-1-0132, and ACS (RSG-09-282-01-CSM) to J.Y., from DOD W81XWH-13-1-0133 to A.J.E., from NIH (DK54441) and HHMI to S.S.T., and from NIH (P01AG007996) to R.L.S. S.C.W. was supported by a NIH Cancer Cell Biology Training grant (2T32CA067754), NIH Molecular Pathology of Cancer Training grant (5T32CA077109), and was an ARCS Foundation Scholar. L.F. was supported by a postdoctoral fellowship from Fondation pour la Recherche Médicale (SPE20130326547).
Author information
Authors and Affiliations
Contributions
S.C.W. and J.Y. conceived the project and wrote the manuscript. S.C.W. and L.F. performed most of the experiments and prepared the figures. J.H.T., Y.G., V.H.P., H.E.M. and A.C.C. contributed to the experimental work. R.L.S., S.S.T. and A.J.E. advised on experimental design. L.F., J.H.T. and A.J.E. revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 2 TWIST1 is required for matrix stiffness-induced EMT.
(A) Confocal microscopy of MCF10A cells grown in 3D culture for 5 days on varying matrix rigidities stained for Laminin V (green) and DAPI (blue) (scale bar, 50 μm). (B–C) Brightfield images of MCF10A (B) and Eph4Ras (C) cells expressing control and shTwist1 shRNAs (scale bar, 75 μm). (D) Lysates of control and shTwist expressing MCF10A cells analyzed by SDS-PAGE and probed for TWIST1 and β-Actin. (E) Quantification of invasive acini of MCF10A shTwist1 cells in 3D culture (∗∗∗, P < 0.001, unpaired two-tailed T-test with Welch’s correction, n = 50 acini/experiment, 3 independent experiments, error bars represent s.d.). (F) qPCR analysis of E-cadherin (CDH1) mRNA expression in MCF10A cells in 3D culture on PA hydrogels treated or not with 5 ng/ml TGF-β for 5 days (P < 0.05, unpaired two-tailed T-test with Welch’s correction, n = 4 independent experiments, statistics source data can be found in Supplementary Table 1, error bars represent s.d.). (G) qPCR analysis of the mRNA expression of mesenchymal markers, Fibronectin (FN1) and Vimentin (VIM), in MCF10A cells in 3D culture on PA hydrogels treated or not with 5 ng/ml TGF-β for 5 days (P < 0.05, unpaired two-tailed T-test with Welch’s correction, n = 4 independent experiments, statistics source data can be found in Supplementary Table 1, error bars represent s.d.).
Supplementary Figure 3 Mechanoregulation of Twist1 nuclear localization in Eph4Ras cells.
Brightfield images (A) and confocal images (scale bar, 50 μm) (B) of Eph4Ras cells cultured on micropatterned glass coverslips for 6 h stained for Twist1 (green) and DAPI (blue) (scale bar, 20 μm). (C) Quantification of nuclear localized Twist1 in percentage of the total cell number (#, not significant, unpaired two-tailed T-test with Welch’s correction, n = 25 cells/experiment, 3 independent experiments, error bars represent s.d.).
Supplementary Figure 4 G3BP2 is a TWIST1 binding protein that localizes in the cytoplasm.
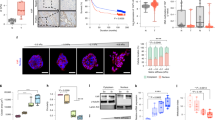
(A) Immunoprecipitation of endogenous TWIST1 from MCF10A cell lysates resolved by SDS-PAGE and silver stained. Unique bands were identified, excised, and analyzed by mass spectrometry. (B) Confocal images of MCF10A and Bt-549 cells grown in 3D culture stained for endogenously expressed G3BP2 (red) and DAPI (blue) (scale bar, 50 μm). (C) Exogenously expressed Twist1 from 293T cell lysates was immunoprecipitated and analyzed by SDS-PAGE, and probed for G3BP2 and Twist1.
Supplementary Figure 5 G3BP2 mediates mechanoregulation of TWIST1 and EMT.
(A) Brightfield images of Eph4Ras (left panel) and MCF10A (right panel) cells expressing control and G3BP2 shRNAs (scale bar, 75 μm). (B) Confocal images of MCF10A cells expressing shRNAs against G3BP2 grown in 3D culture for 5 days on varying matrix rigidities and stained for endogenously expressed TWIST1 (green) and DAPI (blue) (scale bar, 25 μm). (C) qPCR analysis of G3bp2, Twist1 and Snai2 in Eph4Ras cells expressing control (shCtrl#1) or G3bp2 shRNAs, together with control (shCtrl#2) or Twist1 shRNA (shTwist1#5), 3D cultured under indicated matrix rigidities for 5 days (∗, P < 0.05;∗∗, P < 0.01;∗∗∗, P < 0.001, unpaired two-tailed T-test with Welch’s correction, n = 3 independent experiments, statistics source data can be found in Supplementary Table 1; double knockdown compared to the respective single knockdown, error bars represent s.d.). (D) Confocal images of MCF10A cells expressing shRNAs against G3BP2 grown in 3D culture for 5 days on varying matrix rigidities and stained for YAP1 (red) and DAPI (blue) (scale bar, 50 μm).
Supplementary Figure 6 G3BP2 is required for mechanosensing in MCF10DCIS cells.
(A) Brightfield images of MCF10DCIS cells expressing control and G3BP2 shRNAs (scale bar, 25 μm). (B) Brightfield images of MCF10DCIS cells expressing control and G3BP2 shRNAs cultured in 3D at indicated matrix rigidities for 5 days (scale bar, 150 μm). (C) Confocal images of MCF10DCIS cells expressing shRNAs against G3BP2 grown in 3D culture for 5 days on varying matrix rigidities and stained for endogenously expressed TWIST1 (green) and DAPI (blue) (scale bar, 25 μm).
Supplementary Figure 7 G3BP2 expression profile in normal and cancer human tissues.
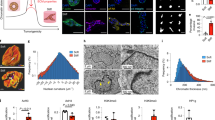
(A) Kaplan–Meier survival curve of patients stratified by G3BP2 expression in the TCGA breast cancer dataset (TCGA_BRCA_G4502A_07_3) (P = 0.2435, Log-Rank). (B) Statistics of overall survival of patients stratified by G3BP2 expression in the TCGA breast cancer dataset (TCGA_BRCA_G4502A_07_3). (C) Kaplan–Meier curve of recurrence free survival in stage 3 breast cancer patients based on SHG imaging (∗∗, P = 0.0047, Log-Rank, n = 197 breast tumours). (D) Confocal microscopy of normal human colon luminal epithelial cells stained for G3BP2 (red) and nuclei (blue) (scale bar, 50 μm). (E,F) Correlation between G3BP2 expression and ER positivity (E) or tumour grade (F) in stage 3 breast cancer patient samples analyzed in (C) (#, not significant, Fisher’s Exact, n = 197 breast tumours, error bars represent s.d.).
Supplementary information
Supplementary Information
Supplementary Information (PDF 1009 kb)
Supplementary Table 1
Supplementary Information (XLSX 75 kb)
Rights and permissions
About this article
Cite this article
Wei, S., Fattet, L., Tsai, J. et al. Matrix stiffness drives epithelial–mesenchymal transition and tumour metastasis through a TWIST1–G3BP2 mechanotransduction pathway. Nat Cell Biol 17, 678–688 (2015). https://doi.org/10.1038/ncb3157
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3157
This article is cited by
-
Linking cell mechanical memory and cancer metastasis
Nature Reviews Cancer (2024)
-
The impact of tumor microenvironment: unraveling the role of physical cues in breast cancer progression
Cancer and Metastasis Reviews (2024)
-
Extracellular matrix remodeling in tumor progression and immune escape: from mechanisms to treatments
Molecular Cancer (2023)
-
Roles of Twist1 in lipid and glucose metabolism
Cell Communication and Signaling (2023)
-
Tumor matrix stiffness provides fertile soil for cancer stem cells
Cancer Cell International (2023)