Abstract
Dynamics of epithelial tissues determine key processes in development, tissue healing and cancer invasion. These processes are critically influenced by cell–cell adhesion forces. However, the identity of the proteins that resist and transmit forces at cell–cell junctions remains unclear, and how these proteins control tissue dynamics is largely unknown. Here we provide a systematic study of the interplay between cell–cell adhesion proteins, intercellular forces and epithelial tissue dynamics. We show that collective cellular responses to selective perturbations of the intercellular adhesome conform to three mechanical phenotypes. These phenotypes are controlled by different molecular modules and characterized by distinct relationships between cellular kinematics and intercellular forces. We show that these forces and their rates can be predicted by the concentrations of cadherins and catenins. Unexpectedly, we identified different mechanical roles for P-cadherin and E-cadherin; whereas P-cadherin predicts levels of intercellular force, E-cadherin predicts the rate at which intercellular force builds up.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Friedl, P. & Gilmour, D. Collective cell migration in morphogenesis, regeneration and cancer. Nat. Rev. Mol. Cell Biol. 10, 445–457 (2009).
Weber, G. F., Bjerke, M. A. & DeSimone, D. W. A mechanoresponsive cadherin-keratin complex directs polarized protrusive behavior and collective cell migration. Dev. Cell 22, 104–115 (2012).
Heisenberg, C. P. & Bellaiche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962 (2013).
Leckband, D. E., le Duc, Q., Wang, N. & de Rooij, J. Mechanotransduction at cadherin-mediated adhesions. Curr. Opin. Cell Biol. 23, 523–530 (2011).
Budnar, S. & Yap, A. S. A mechanobiological perspective on cadherins and the actin-myosin cytoskeleton. F1000Prime Rep. 5, 35 (2013).
Tambe, D. T. et al. Collective cell guidance by cooperative intercellular forces. Nat. Mater. 10, 469–475 (2011).
Theveneau, E. et al. Chase-and-run between adjacent cell populations promotes directional collective migration. Nat. Cell Biol. 15, 763–772 (2013).
Campinho, P. et al. Tension-oriented cell divisions limit anisotropic tissue tension in epithelial spreading during zebrafish epiboly. Nat. Cell Biol. 15, 1405–1414 (2013).
Bosveld, F. et al. Mechanical control of morphogenesis by Fat/Dachsous/Four-jointed planar cell polarity pathway. Science 336, 724–727 (2012).
Legoff, L., Rouault, H. & Lecuit, T. A global pattern of mechanical stress polarizes cell divisions and cell shape in the growing Drosophila wing disc. Dev. Suppl. 140, 4051–4059 (2013).
Brugués, A. et al. Forces driving epithelial wound healing. Nat. Phys. 10, 683–690 (2014).
Rauzi, M., Lenne, P. F. & Lecuit, T. Planar polarized actomyosin contractile flows control epithelial junction remodelling. Nature 468, 1110–1114 (2010).
Serra-Picamal, X. et al. Mechanical waves during tissue expansion. Nat. Phys. 8, 628–634 (2012).
Foty, R. A. & Steinberg, M. S. The differential adhesion hypothesis: a direct evaluation. Dev. Biol. 278, 255–263 (2005).
Borghi, N. et al. E-cadherin is under constitutive actomyosin-generated tension that is increased at cell–cell contacts upon externally applied stretch. Proc. Natl Acad. Sci. USA 109, 12568–12573 (2012).
Maitre, J. L. & Heisenberg, C. P. Three functions of cadherins in cell adhesion. Curr. Biol. 23, R626–R633 (2013).
Zaidel-Bar, R. Cadherin adhesome at a glance. J. Cell Sci. 126, 373–378 (2013).
Nieman, M. T., Prudoff, R. S., Johnson, K. R. & Wheelock, M. J. N-cadherin promotes motility in human breast cancer cells regardless of their E-cadherin expression. J. Cell Biol. 147, 631–644 (1999).
Ribeiro, A. S. et al. Extracellular cleavage and shedding of P-cadherin: a mechanism underlying the invasive behaviour of breast cancer cells. Oncogene 29, 392–402 (2010).
Batlle, E. et al. The transcription factor snail is a repressor of E-cadherin gene expression in epithelial tumour cells. Nat. Cell Biol. 2, 84–89 (2000).
Kumper, S. & Ridley, A. J. p120ctn and P-cadherin but not E-cadherin regulate cell motility and invasion of DU145 prostate cancer cells. PLoS ONE 5, e11801 (2010).
Ribeiro, A. S. et al. P-cadherin functional role is dependent on E-cadherin cellular context: a proof of concept using the breast cancer model. J. Pathol. 229, 705–718 (2013).
Tabdili, H. et al. Cadherin-dependent mechanotransduction depends on ligand identity but not affinity. J. Cell Sci. 125, 4362–4371 (2012).
Buckley, C. D. et al. Cell adhesion. The minimal cadherin-catenin complex binds to actin filaments under force. Science 346, 1254211 (2014).
Abe, K. & Takeichi, M. EPLIN mediates linkage of the cadherin catenin complex to F-actin and stabilizes the circumferential actin belt. Proc. Natl Acad. Sci. USA 105, 13–19 (2008).
Yamada, S., Pokutta, S., Drees, F., Weis, W. I. & Nelson, W. J. Deconstructing the cadherin-catenin-actin complex. Cell 123, 889–901 (2005).
Runkle, E. A. & Mu, D. Tight junction proteins: from barrier to tumorigenesis. Cancer Lett. 337, 41–48 (2013).
Harris, A. R. et al. Characterizing the mechanics of cultured cell monolayers. Proc. Natl Acad. Sci. USA 109, 16449–16454 (2012).
Herrmann, H., Bar, H., Kreplak, L., Strelkov, S. V. & Aebi, U. Intermediate filaments: from cell architecture to nanomechanics. Nat. Rev. Mol. Cell Biol. 8, 562–573 (2007).
Wang, N. & Stamenovic, D. Contribution of intermediate filaments to cell stiffness, stiffening, and growth. Am. J. Physiol. 279, C188–C194 (2000).
Bao, L., Sachs, F. & Dahl, G. Connexins are mechanosensitive. Am. J. Physiol. 287, C1389–C1395 (2004).
Poujade, M. et al. Collective migration of an epithelial monolayer in response to a model wound. Proc. Natl Acad. Sci. USA 104, 15988–15993 (2007).
Hur, S. S. et al. Roles of cell confluency and fluid shear in 3-dimensional intracellular forces in endothelial cells. Proc. Natl Acad. Sci. USA 109, 11110–11115 (2012).
Tambe, D. T. et al. Monolayer stress microscopy: limitations, artifacts, and accuracy of recovered intercellular stresses. PLoS ONE 8, e55172 (2013).
Trepat, X. & Fredberg, J. J. Plithotaxis and emergent dynamics in collective cellular migration. Trends Cell Biol. 21, 638–646 (2012).
Trepat, X. et al. Physical forces during collective cell migration. Nat. Phys. 5, 426–430 (2009).
Liu, Z. et al. Mechanical tugging force regulates the size of cell–cell junctions. Proc. Natl Acad. Sci. USA 107, 9944–9949 (2010).
Maruthamuthu, V., Sabass, B., Schwarz, U. S. & Gardel, M. L. Cell–ECM traction force modulates endogenous tension at cell–cell contacts. Proc. Natl Acad. Sci. USA 108, 4708–4713 (2011).
Ng, M. R., Besser, A., Brugge, J. S. & Danuser, G. Mapping the dynamics of force transduction at cell–cell junctions of epithelial clusters. eLife 4, e03282 (2014).
Simpson, K. J. et al. Identification of genes that regulate epithelial cell migration using an siRNA screening approach. Nat. Cell Biol. 10, 1027–1038 (2008).
Sales-Pardo, M., Guimera, R., Moreira, A. A. & Amaral, L. A. Extracting the hierarchical organization of complex systems. Proc. Natl Acad. Sci. USA 104, 15224–15229 (2007).
Fanning, A. S., Van Itallie, C. M. & Anderson, J. M. Zonula occludens-1 and -2 regulate apical cell structure and the zonula adherens cytoskeleton in polarized epithelia. Mol. Biol. Cell 23, 577–590 (2012).
Tuomi, S. et al. PKCepsilon regulation of an α5 integrin-ZO-1 complex controls lamellae formation in migrating cancer cells. Sci. Signal. 2, ra32 (2009).
Bausch, A. R., Moller, W. & Sackmann, E. Measurement of local viscoelasticity and forces in living cells by magnetic tweezers. Biophys. J. 76, 573–579 (1999).
Le Duc, Q. et al. Vinculin potentiates E-cadherin mechanosensing and is recruited to actin-anchored sites within adherens junctions in a myosin II-dependent manner. J. Cell Biol. 189, 1107–1115 (2010).
Riveline, D. et al. Focal contacts as mechanosensors: externally applied local mechanical force induces growth of focal contacts by an mDia1-dependent and ROCK-independent mechanism. J. Cell Biol. 153, 1175–1186 (2001).
Roca-Cusachs, P., Gauthier, N. C., Del Rio, A. & Sheetz, M. P. Clustering of α(5)β(1) integrins determines adhesion strength whereas α(v)β(3) and talin enable mechanotransduction. Proc. Natl Acad. Sci. USA 106, 16245–16250 (2009).
Yonemura, S., Wada, Y., Watanabe, T., Nagafuchi, A. & Shibata, M. α-Catenin as a tension transducer that induces adherens junction development. Nat. Cell Biol. 12, 533–542 (2010).
Yao, M. et al. Force-dependent conformational switch of α-catenin controls vinculin binding. Nat. Commun. 5, 4525 (2014).
Barry, A. K. et al. α-catenin cytomechanics–role in cadherin-dependent adhesion and mechanotransduction. J. Cell Sci. 127, 1779–1791 (2014).
Cano, A. et al. The transcription factor snail controls epithelial–mesenchymal transitions by repressing E-cadherin expression. Nat. Cell Biol. 2, 76–83 (2000).
Thiery, J. P., Acloque, H., Huang, R. Y. & Nieto, M. A. Epithelial-mesenchymal transitions in development and disease. Cell 139, 871–890 (2009).
Stark, C. et al. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 34, D535–D539 (2006).
Albergaria, A. et al. P-cadherin role in normal breast development and cancer. Int. J. Dev. Biol. 55, 811–822 (2011).
Van Roy, F. Beyond E-cadherin: roles of other cadherin superfamily members in cancer. Nat. Rev. Cancer 14, 121–134 (2014).
Vleminckx, K., Vakaet, L. Jr, Mareel, M., Fiers, W. & van Roy, F. Genetic manipulation of E-cadherin expression by epithelial tumor cells reveals an invasion suppressor role. Cell 66, 107–119 (1991).
Wong, A. S. & Gumbiner, B. M. Adhesion-independent mechanism for suppression of tumor cell invasion by E-cadherin. J. Cell Biol. 161, 1191–1203 (2003).
Van Marck, V. et al. P-cadherin promotes cell–cell adhesion and counteracts invasion in human melanoma. Cancer Res. 65, 8774–8783 (2005).
Jacobs, K. et al. P-cadherin counteracts myosin II-B function: implications in melanoma progression. Mol. Cancer 9, 255 (2010).
Bechhoefer, J. Feedback for physicists: a tutorial essay on control. Rev. Mod. Phys. 77, 783–836 (2005).
Cloutier, M. & Wellstead, P. The control systems structures of energy metabolism. J. R. Soc. Interface 7, 651–665 (2010).
Kandow, C. E., Georges, P. C., Janmey, P. A. & Beningo, K. A. Polyacrylamide hydrogels for cell mechanics: steps toward optimization and alternative uses. Methods Cell Biol. 83, 29–46 (2007).
Yeung, T. et al. Effects of substrate stiffness on cell morphology, cytoskeletal structure, and adhesion. Cell. Motil. Cytoskeleton 60, 24–34 (2005).
Ostuni, E., Kane, R. S., Chen, C. S., Ingber, D. E. & Whitesides, G. M. Patterning mammalian cells using elastomeric membranes. Langmuir 16, 7811–7819 (2000).
Blanchard, G. B. et al. Tissue tectonics: morphogenetic strain rates, cell shape change and intercalation. Nat. Methods 6, 458–464 (2009).
Roca-Cusachs, P. et al. Integrin-dependent force transmission to the extracellular matrix by α-actinin triggers adhesion maturation. Proc. Natl Acad. Sci. USA 110, E1361–E1370 (2013).
Fabry, B. et al. Time scale and other invariants of integrative mechanical behavior in living cells. Phys. Rev. 68, 041914 (2003).
Birmingham, A. et al. Statistical methods for analysis of high-throughput RNA interference screens. Nat. Methods 6, 569–575 (2009).
Simon, R. in Fundamentals of Data Mining in Genomics and Proteomics (eds Dubitzky, W., Granzow, M. & Berrar, D. P.) 177–178 (Springer Science+Business Media, 2007).
Acknowledgements
We thank F. Supek, B. Lehner, A. Brugués and R. Vincent for discussions, R. Zaidel-Bar for sharing unpublished data, and E. Sahai for contributing reagents. This research was supported by the European Research Council (StG-242993 and CoG - 616480 to X.T.), the 7th European Community Framework Programme (PCIG10-GA-2011-303848 to P.R-C., PIRG-GA-2010-277166 to R.G., PIRG-GA-2010-268342 to M.S-P., FET Grant 317532 to M.S-P. and R.G.), the Spanish Ministerio de Economía y Comptetitividad (BFU2012-38146 to X.T., DPI2013-43727-R to J.J.M., FIS2010-18639 and FIS2013-47532-C3 to M.S-P. and R.G., BFU2011-23111 to P.R-C., Juan de la Cierva Fellowship JCI-2012-15123 to V.C.), the National Institutes of Health (R01HL107561 to X.T.), Fundació La Caixa, Fundació la Marató de TV3 (20133330 to P.R-C.), and the James S. McDonnell Foundation (R.G. and M.S-P.).
Author information
Authors and Affiliations
Contributions
E.B. and X.T. conceived the study and designed experiments. E.B., M.B-M. and A.E-A. performed experiments. E.B., V.C. and A.E-A. analysed data. X.S-P. and P.R-C. developed data analysis tools. V.C. and J.J.M. developed computational mechanics tools. M.S-P. and R.G. performed unsupervised clustering analysis and LOOCV analysis. E.B., M.S-P., R.G. and X.T. wrote the manuscript. All authors discussed and interpreted results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
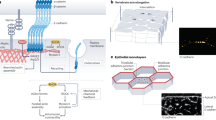
Supplementary Figure 5 mRNA and protein expression levels after siRNA transfections.
(a) Scheme illustrating the localization of all targeted proteins. (b) Levels of mRNA after 5 days of transfection. Data were quantified by RT-PCR. Data are presented as mean ± SEM (normalized to control cells). RT-PCR was run in triplicate. n = 12 samples pooled from 4 independent experiments for CT siRNA, Ncad siRNA, Pcad siRNA, DSC3 siRNA, αcat siRNA, p120 siRNA, LIMA1 siRNA, CLDN8 siRNA, JAM-A siRNA; n = 9 samples pooled from 3 independent experiments for βcat siRNA, DDR1 siRNA, VCL siRNA, CLDN1 siRNA, CLDN7 siRNA, ZO-1 siRNA, ZO-3 siRNA, PKP2 siRNA, JUP siRNA; n = 6 samples pooled from 2 independent experiments for CLDN4 siRNA, CX43 siRNA, OCLN siRNA. (c) Protein levels after 5 days of transfection. Data are presented as mean ± SEM. For each protein, n = 3 samples pooled from 3 independent transfections (normalized to control cells).
Supplementary Figure 6 Representative maps of monolayer mechanics for siRNA perturbations targeting E-, N-, and P-cadherin.
Data show phase contrast images (first column), velocities in the x direction (second column), traction forces in the x direction (third column), and monolayer tension (fourth column) for two time points (0h first row and 6h second row). Scale bar, 100 μm.
Supplementary Figure 7 Representative maps of monolayer mechanics for siRNA perturbations targeting α-catenin, β-catenin, and p120.
Data show phase contrast images (first column), velocities in the x direction (second column), traction forces in the x direction (third column), and monolayer tension (fourth column) for two time points (0h first row and 6h second row). Scale bar, 100 μm.
Supplementary Figure 8 Representative maps of monolayer mechanics for siRNA perturbations targeting DDR1, LIMA1, and VCL.
Data show phase contrast images (first column), velocities in the x direction (second column), traction forces in the x direction (third column), and monolayer tension (fourth column) for two time points (0h first row and 6h second row). Scale bar, 100 μm.
Supplementary Figure 9 Representative maps of monolayer mechanics for siRNA perturbations targeting JAM-A, OCLN, ZO-1, and ZO-3.
Data show phase contrast images (first column), velocities in the x direction (second column), traction forces in the x direction (third column), and monolayer tension (fourth column) for two time points (0h first row and 6h second row). Scale bar, 100 μm.
Supplementary Figure 10 Representative maps of monolayer mechanics for siRNA perturbations targeting claudins 1, 4, 7, and 8.
Data show phase contrast images (first column), velocities in the x direction (second column), traction forces in the x direction (third column), and monolayer tension (fourth column) for two time points (0h first row and 6h second row). Scale bar, 100 μm.
Supplementary Figure 11 Representative maps of monolayer mechanics for siRNA perturbations targeting desmocollin 3, plakophilin 2, plakoglobin, and connexin 43.
Data show phase contrast images (first column), velocities in the x direction (second column), traction forces in the x direction (third column), and monolayer tension (fourth column) for two time points (0h first row and 6h second row). Scale bar, 100 μm.
Supplementary Figure 12 Time evolution of monolayer mechanics for each siRNA perturbation.
Data are presented as mean ± SEM. n = 13 independent cell monolayers (CT siRNA), n = 3 independent cell monolayers (Ecad siRNA, βcat siRNA, JAM-A siRNA, ZO-3 siRNA, Ncad siRNA, LIMA1 siRNA, DDR1 siRNA, PKP2 siRNA), n = 4 independent cell monolayers (Pcad siRNA, αcat siRNA, p120 siRNA, CX43 siRNA, VCL siRNA, JUP siRNA), n = 5 independent cell monolayers (DSC3 siRNA, CLDN1 siRNA, CLDN8 siRNA), n = 6 independent monolayers (OCLN siRNA, CLDN4 siRNA), n = 7 independent cell monolayers (ZO-1 siRNA), n = 8 independent cell monolayers (CLDN7 siRNA); monolayers were assessed from 10 experiments (CT siRNA), 4 experiments (CLDN7 siRNA, CLDN4 siRNA), 3 experiments (αcat siRNA, CX43 siRNA, DSC3 siRNA), 2 experiments (Ecad siRNA, βcat siRNA, JAM-A siRNA, ZO-3 siRNA, Ncad siRNA, LIMA1 siRNA, DDR1 siRNA, PKP2 siRNA, Pcad siRNA, p120 siRNA, VCL siRNA, JUP siRNA, CLDN1 siRNA, CLDN8 siRNA, OCLN siRNA, ZO-1 siRNA).
Supplementary information
Supplementary Information
Supplementary Information (PDF 5154 kb)
Expansion of a micropatterned monolayer of MCF10A cells.
Scale bar, 100 μm. (AVI 3111 kb)
Dynamics of an expanding cell monolayer of MCF10A cells.
Top: velocity field Vx overlaid on phase contrast images. Middle: traction force field Tx overlaid on phase contrast images. Bottom: monolayer tension σxx overlaid on phase contrast images. Scale bar, 100 μm. (AVI 3523 kb)
Time evolution of monolayer tension in response to siRNAs targeting cell-cell junctions.
Composition of representative experiments for the 21 siRNA pools and 3 controls. Each panel shows monolayer tension σxx overlaid on phase contrast images. Scale bar, 100 μm. (AVI 3648 kb)
Rights and permissions
About this article
Cite this article
Bazellières, E., Conte, V., Elosegui-Artola, A. et al. Control of cell–cell forces and collective cell dynamics by the intercellular adhesome. Nat Cell Biol 17, 409–420 (2015). https://doi.org/10.1038/ncb3135
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3135
This article is cited by
-
Programmable integrin and N-cadherin adhesive interactions modulate mechanosensing of mesenchymal stem cells by cofilin phosphorylation
Nature Communications (2022)
-
Two Rac1 pools integrate the direction and coordination of collective cell migration
Nature Communications (2022)
-
Intrinsic cell rheology drives junction maturation
Nature Communications (2022)
-
Endothelial cell-specific molecule 1 drives cervical cancer progression
Cell Death & Disease (2022)
-
Machine learning phenomics (MLP) combining deep learning with time-lapse-microscopy for monitoring colorectal adenocarcinoma cells gene expression and drug-response
Scientific Reports (2022)