Abstract
Asymmetric division of neural precursor cells contributes to the generation of a variety of neuronal types. Asymmetric division is mediated by the asymmetric inheritance of fate determinants by the two daughter cells1,2. In vertebrates, asymmetric fate determinants, such as Par3 and Mib, are only now starting to be identified3,4. Here we show that, during mitosis of neural precursors in zebrafish, directional trafficking of Sara endosomes to one of the daughters can function as such a determinant. In asymmetric lineages, where one daughter cell becomes a neuron (n cell) whereas the other divides again to give rise to two neurons (p cell), we found that the daughter that inherits most of the Sara endosomes acquires the p fate. Sara endosomes carry an endocytosed pool of the Notch ligand DeltaD, which is thereby itself distributed asymmetrically. Sara and Notch are both essential for cell fate assignation within asymmetric lineages. Therefore, the Sara endosome system determines the fate decision between neuronal differentiation and mitosis in asymmetric lineages and thereby contributes to controlling the number of neural precursors and differentiated neurons during neurogenesis in a vertebrate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Neumuller, R. A. & Knoblich, J. A. Dividing cellular asymmetry: asymmetric cell division and its implications for stem cells and cancer. Genes Dev. 23, 2675–2699 (2009).
Gonczy, P. Mechanisms of asymmetric cell division: flies and worms pave the way. Nat. Rev. Mol. Cell Biol. 9, 355–366 (2008).
Alexandre, P., Reugels, A. M., Barker, D., Blanc, E. & Clarke, J. D. Neurons derive from the more apical daughter in asymmetric divisions in the zebrafish neural tube. Nat. Neurosci. 13, 673–679 (2010).
Dong, Z., Yang, N., Yeo, S. Y., Chitnis, A. & Guo, S. Intralineage directional Notch signaling regulates self-renewal and differentiation of asymmetrically dividing radial glia. Neuron 74, 65–78 (2012).
Ando, R., Hama, H., Yamamoto-Hino, M., Mizuno, H. & Miyawaki, A. An optical marker based on the UV-induced green-to-red photoconversion of a fluorescent protein. Proc. Natl Acad. Sci. USA 99, 12651–12656 (2002).
Papan, C. & Campos-Ortega, J. A. Region-specific cell clones in the developing spinal cord of the zebrafish. Dev. Genes Evol. 209, 135–144 (1999).
Coumailleau, F., Furthauer, M., Knoblich, J. A. & Gonzalez-Gaitan, M. Directional delta and Notch trafficking in Sara endosomes during asymmetric cell division. Nature 458, 1051–1055 (2009).
Montagne, C. & Gonzalez-Gaitan, M. Sara endosomes and the asymmetric division of intestinal stem cells. Development 141, 2014–2023 (2014).
Kutateladze, T. G. Phosphatidylinositol 3-phosphate recognition and membrane docking by the FYVE domain. Biochim. Biophys. Acta 1761, 868–877 (2006).
Blatner, N. R. et al. The molecular basis of the differential subcellular localization of FYVE domains. J. Biol. Chem. 279, 53818–53827 (2004).
Hayakawa, A. et al. Structural basis for endosomal targeting by FYVE domains. J. Biol. Chem. 279, 5958–5966 (2004).
Mao, Y. et al. Crystal structure of the VHS and FYVE tandem domains of Hrs, a protein involved in membrane trafficking and signal transduction. Cell 100, 447–456 (2000).
Itoh, F. et al. The FYVE domain in Smad anchor for receptor activation (SARA) is sufficient for localization of SARA in early endosomes and regulates TGF-β/Smad signalling. Genes Cells 7, 321–331 (2002).
Ulrich, F. et al. Wnt11 functions in gastrulation by controlling cell cohesion through Rab5c and E-cadherin. Dev. Cell 9, 555–564 (2005).
Kok, F. O. et al. Reverse genetic screening reveals poor correlation between Morpholino-Induced and mutant phenotypes in Zebrafish. Dev. Cell 32, 97–108 (2014).
Kawaguchi, D., Furutachi, S., Kawai, H., Hozumi, K. & Gotoh, Y. Dll1 maintains quiescence of adult neural stem cells and segregates asymmetrically during mitosis. Nat. Commun. 4, 1880 (2013).
Geling, A., Steiner, H., Willem, M., Bally-Cuif, L. & Haass, C. A γ-secretase inhibitor blocks Notch signaling in vivo and causes a severe neurogenic phenotype in zebrafish. EMBO Rep. 3, 688–694 (2002).
Dovey, H. F. et al. Functional γ-secretase inhibitors reduce β-amyloid peptide levels in brain. J. Neurochem. 76, 173–181 (2001).
Appel, B., Givan, L. A. & Eisen, J. S. Delta-Notch signaling and lateral inhibition in zebrafish spinal cord development. BMC Dev. Biol. 1, 13 (2001).
Haddon, C. et al. Multiple delta genes and lateral inhibition in zebrafish primary neurogenesis. Development 125, 359–370 (1998).
Lewis, J. Neurogenic genes and vertebrate neurogenesis. Curr. Opin. Neurobiol. 6, 3–10 (1996).
Itoh, M. et al. Mind bomb is a ubiquitin ligase that is essential for efficient activation of Notch signaling by Delta. Dev. Cell 4, 67–82 (2003).
Scheer, N. & Campos-Ortega, J. A. Use of the Gal4-UAS technique for targeted gene expression in the zebrafish. Mech. Dev. 80, 153–158 (1999).
Scheer, N., Groth, A., Hans, S. & Campos-Ortega, J. A. An instructive function for Notch in promoting gliogenesis in the zebrafish retina. Development 128, 1099–1107 (2001).
Scheer, N., Riedl, I., Warren, J. T., Kuwada, J. Y. & Campos-Ortega, J. A. A quantitative analysis of the kinetics of Gal4 activator and effector gene expression in the zebrafish. Mech. Dev. 112, 9–14 (2002).
Matsuda, M. & Chitnis, A. B. Interaction with Notch determines endocytosis of specific Delta ligands in zebrafish neural tissue. Development 136, 197–206 (2009).
Westerfield, M. The Zebrafish Book: A Guide for The Laboratory Use of Zebrafish (Danio Rerio) (Univ. Oregon Press, 2000).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic development of the zebrafish. Dev. Dyn. 203, 253–310 (1995).
Montero, J. A. et al. Shield formation at the onset of zebrafish gastrulation. Development 132, 1187–1198 (2005).
Acknowledgements
We are very grateful to O. Schaad, C. Seum, C. Alliod, E. Derivery and F. Schütz for technical help, A. Reugels and M. Brand for the Par3–GFP and Palmitoylated–RFP constructs, respectively, and A. Oates, C. Gonzalez and J. Bertrand for critical reading of the manuscript. We also thank R. Finazzi, S. Borgers and O. Seum for fish care. C.C. was supported by Fundação para a Ciência e Tecnologia (SFRH/BD/15210/2004) and ONCASYM and the PGDB PhD programme. M.F. was supported by HFSP, CNRS and INSERM. This work was supported by the Département de l’Instruction Publique of the Canton of Geneva, SNSF, the SystemsX epiPhysX grant, ERC advanced grants (Sara and Morphogen), the NCCR Frontiers in Genetics and Chemical Biology programs and the Polish–Swiss research program to M.G-G.
Author information
Authors and Affiliations
Contributions
S.K., C.C. and I.C. designed and carried out the experiments, generated and interpreted the data and prepared the manuscript. M.F. designed and carried out the iDeltaD antibody uptake, contributed to the Par3 analysis, interpreted the data and contributed to the supervision of the project. M.G-G. supervised the project, interpreted the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Neural progenitor lineages at different developmental stages.
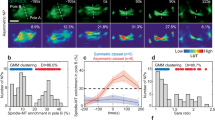
a–b, Schemes of lineage tracing by Kaede photoconversion of either neural precursor mother cells (a) or daughter cells (b). Single neural precursor cells (a) or one of their daughters (b) expressing Kaede in the neural plate (c), neural rod (d) and neural tube (e) were photoconverted from green to red. When we photoconverted mothers or daughters at plate, rod or tube stage, the second division of the daughters in the case of n ⋅ p and p ⋅ p lineages occurred late during tube stage. Therefore, the photoconverted mothers in early tube stage are not the daughters of neural precursors that divided at plate or rod stage. c–e, Neural precursor mother cell photoconversion at plate stage shows that 75% of the lineages are p ⋅ p, while 25% corresponds to the n ⋅ n lineage (c). The frequencies change at rod and tube stages (d,e). At rod stage, 53% corresponds to p ⋅ p lineage, while 35% and 12% correspond to the n ⋅ p and n ⋅ n lineages, respectively (d). At tube stage, the frequencies obtained were 18% for the p ⋅ p lineage, 58% for the n ⋅ p and 24% for the n ⋅ n lineages (e). From the observed lineages through photoconversions of mother cells at plate (c), rod (d) and tube (e) stages, the expected amount of 1-cell and 2- or more-cell clones after photoconversion of the daughter cells can be calculated (“Expected”). The total expected frequencies correlate with the “Observed” frequencies. It is worth noting that only the sum of all lineages was given for the observed frequency since our assay does not allow distinguish between the three different lineages.
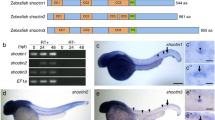
Supplementary Figure 2 sara ubiquitous expression, endosomal localization and lack of correlation between Sara endosome asymmetry and lineage type.
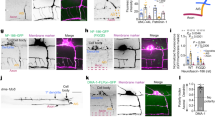
a, sara mRNA expression pattern during embryogenesis. Control: sara sense probe. Scale bar: 250 μm. b, Colocalization between Venus-Sara and CFP-Rab5-positive early endosomes (arrowheads) in the spinal cord. Right panels, magnification of the boxes in the left panels. c,e, Cell frequency distribution of YFPRab5c (c) and YFP-Rab7-positive endosome ratios between the two daughters during neural precursor divisions (e). Both Rab5- and Rab7-positive endosomes segregate symmetrically. d,f, Confocal images of dividing neural precursors showing symmetric distribution of either YFP-Rab5c (d) or YFP-Rab7 endosomes (f). Cell profile counterstained by Gap43-CFP. g, Percentage of cells with an endosomal ratio higher than 1.5 for Venus-Sara-, YFP-Rab5c-, YFP-Rab11a-, and YFP-Rab7-positive endosomes. h, Sara endosome ratio between the daughter cells and the type of lineages generated subsequently (data set from Fig. 2a). Statistical analyses (t-test) comparing the averages of endosome asymmetry in the different types of lineages show that there is not correlation between Sara endosome asymmetry and the type of lineage. Three outliers with abnormally high levels of asymmetry (6, 7 and 12 fold) were excluded from the statistics. i, Correlation between Sara endosome ratio and n vs. p fate in dividing neural precursors expressing Rab5Q81L. While there is no correlation between Sara endosome ratio and n vs. p fate acquisition in wildtype (h), Sara endosome asymmetry is much higher in neural precursors expressing Rab5Q81L and almost all lineages are n ⋅ p, under this condition. Grey area, Sara endosome asymmetry range in wildtype as seen in (h). Dashed lines correspond to the 1.5 fold threshold, which defines “asymmetric dispatch” in this work. j, Confocal image of neural precursors expressing YFP-Rab5Q81L and mRFP-Sara showing Sara endosomes as few large vesicular structures (white arrows). k, Scheme and corresponding confocal images of daughter photoconversion during division of precursor cells expressing YFP-Rab5Q81L/Sara and Kaede. Either the daughter cell that inherits fewer (upper panel) or more YFP-Rab5Q81L/Sara endosomes (lower panel) is photoconverted (middle panels: right panel red channel; left, merge). The fate of the photoconverted daughter is determined 48 h later by the number of cells in the red lineages (red cells; most right panel). l, Percentage of daughters acquiring p vs. n fate among those inheriting Rab5Q81L/Sara or not as in k. Scale bars: 5 μm.
Supplementary Figure 3 Sara endosome segregation correlates with n versus p fate.
a–c, Schemes showing the frequency distribution of lineages and the correlation between Sara endosome segregation and mitotic fate at plate (a), rod (b) and tube (c) stages. Frequencies of the n ⋅ n, n ⋅ p and p ⋅ p lineages when the neural progenitor mother cell was photoconverted are shown as in Supplementary Fig. 1. The frequency of the different lineages was estimated (“expected”) according to the lineages observed when the neural precursor mother cell was photoconverted. The frequencies observed when either the daughter that inherited more Sara (Sara+) or the daughter that inherited less Sara endosomes (Sara−) was photoconverted do correlate with the expected frequencies.
Supplementary Figure 4 Sara endosomes, Par3 and Mindbomb.
a, Confocal images showing the normal apical localization of Sara endosomes in par3 morphants (cf. Fig. 1k–m). Scale bar: 20 μm. b,c Normal Sara endosome asymmetric segregation during neural precursor division in par3 morphants. Confocal image (b; dotted white line marks cell outline) and frequencies of cells with Sara endosome ratio above 1.5 in wildtype and par3 morphant embryos (c; t-test; p > 0.7). d–e, Apical localization of Par3-GFP in both wildtype (d) and sara morphant (e). Scale bar: 20 μm. f,g, Asymmetric segregation of Par3-GFP into the two daughter cells during the division of a wild type (f) and sara morphant neural progenitor cell (g). Dotted white line marks cell outline of dividing cell. In a,b and f, cell profile is counterstained by palmitoylated-RFP. h, Cell frequency distribution of apical Par3-GFP ratio between the two daughters during neural precursor divisions (e). Fisher’s Exact Test; p > 0.1. i–l, Specificity of the Delta antibody uptake assay. i,j, Confocal images showing the spinal cord of embryos injected with either a DeltaD antibody labeled with Zenon 488 (j) or with Zenon 488 alone (i). k,l, Confocal images showing DeltaD staining after the iDeltaD internalization assay in wildtype (k) and mib morphant embryos (l). Note the plasma membrane staining in mib morphants. Scale bar: 20 μm. m, Confocal images showing colocalization between Mib-GFP intracellular vesicular structures and Sara-positive endosomes (white arrowheads) in the spinal cord. Scale bar: 20 μm. n, Confocal images showing that mRFP-Sara endosomes and Mib-GFP segregate preferentially into the same daughter cell (arrowhead). Dotted white line marks cell outline of dividing cell. o, Confocal images showing the asymmetric segregation of Venus-Sara endosomes during neural progenitor in notch1a/notch3 double morphants, and deltad and mib morphants. Dotted white line marks cell outline of dividing cell. p, Venus-Sara endosome ratio between the two daughter cells after neural progenitor division in wildtype and different Notch pathway blockage conditions. Sara endosomes asymmetric segregation (Sara endosome ratio > 1.5) is unaffected in notch1a/notch3 double morphants, and deltad and mib single morphants compared with wild type. Scale bar: 5 μm, unless stated otherwise.
Supplementary information
Supplementary Information
Supplementary Information (PDF 3420 kb)
Segregation of Sara endosomes in a dividing neural progenitor cell.
Sara endosomes (marked by Venus-Sara, depicted in green) are asymmetrically segregated into the two daughter cells. GAP43-CFP (depicted in red) outlines the membrane of the dividing neural precursor cell. (MOV 1753 kb)
Dynamics of Sara endosomes and their DeltaD cargo in dividing neural progenitor cells.
Sara endosomes (marked by mRFP-Sara, depicted in green) transport endogenous internalized DeltaD ligands (iDeltaD, red). GAP43-GFP (depicted in magenta) outlines the membrane of both the dividing neural precursor and neighboring neuro-epithelial cells. (AVI 174 kb)
Rights and permissions
About this article
Cite this article
Kressmann, S., Campos, C., Castanon, I. et al. Directional Notch trafficking in Sara endosomes during asymmetric cell division in the spinal cord. Nat Cell Biol 17, 333–339 (2015). https://doi.org/10.1038/ncb3119
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3119
This article is cited by
-
A cell fate decision map reveals abundant direct neurogenesis bypassing intermediate progenitors in the human developing neocortex
Nature Cell Biology (2024)
-
Elongator stabilizes microtubules to control central spindle asymmetry and polarized trafficking of cell fate determinants
Nature Cell Biology (2022)
-
WDR81 regulates adult hippocampal neurogenesis through endosomal SARA-TGFβ signaling
Molecular Psychiatry (2021)
-
Notch-mediated inhibition of neurogenesis is required for zebrafish spinal cord morphogenesis
Scientific Reports (2019)
-
Sara phosphorylation state controls the dispatch of endosomes from the central spindle during asymmetric division
Nature Communications (2017)