Abstract
Selective macroautophagy is an important protective mechanism against diverse cellular stresses. In contrast to the well-characterized starvation-induced autophagy, the regulation of selective autophagy is largely unknown. Here, we demonstrate that Huntingtin, the Huntington disease gene product, functions as a scaffold protein for selective macroautophagy but it is dispensable for non-selective macroautophagy. In Drosophila, Huntingtin genetically interacts with autophagy pathway components. In mammalian cells, Huntingtin physically interacts with the autophagy cargo receptor p62 to facilitate its association with the integral autophagosome component LC3 and with Lys-63-linked ubiquitin-modified substrates. Maximal activation of selective autophagy during stress is attained by the ability of Huntingtin to bind ULK1, a kinase that initiates autophagy, which releases ULK1 from negative regulation by mTOR. Our data uncover an important physiological function of Huntingtin and provide a missing link in the activation of selective macroautophagy in metazoans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
He, C. & Klionsky, D. J. Regulation mechanisms and signaling pathways of autophagy. Annu. Rev. Genet. 43, 67–93 (2009).
Bjorkoy, G., Lamark, T. & Johansen, T. p62/SQSTM1: a missing link between protein aggregates and the autophagy machinery. Autophagy 2, 138–139 (2006).
Johansen, T. & Lamark, T. Selective autophagy mediated by autophagic adapter proteins. Autophagy 7, 279–296 (2011).
Lamark, T. & Johansen, T. Aggrephagy: selective disposal of protein aggregates by macroautophagy. Int. J. Cell Biol. 2012, 736905 (2012).
Lynch-Day, M. A. & Klionsky, D. J. The Cvt pathway as a model for selective autophagy. FEBS Lett. 584, 1359–1366 (2010).
Jin, M., Liu, X. & Klionsky, D. J. SnapShot: Selective autophagy. Cell 152, 368–368.e2 (2013).
The Huntington’s Disease Collaborative Research Group, A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 72, 971–983 (1993).
Cattaneo, E., Zuccato, C. & Tartari, M. Normal huntingtin function: an alternative approach to Huntington’s disease. Nat. Rev. Neurosci. 6, 919–930 (2005).
Caviston, J. P. & Holzbaur, E. L. Huntingtin as an essential integrator of intracellular vesicular trafficking. Trends Cell Biol. 19, 147–155 (2009).
Harjes, P. & Wanker, E. E. The hunt for huntingtin function: interaction partners tell many different stories. Trends Biochem. Sci. 28, 425–433 (2003).
Martin, D. D., Ladha, S., Ehrnhoefer, D. E. & Hayden, M. R. Autophagy in Huntington disease and huntingtin in autophagy. Trends Neurosci. 38, 26–35 (2015).
Martinez-Vicente, M. et al. Cargo recognition failure is responsible for inefficient autophagy in Huntington’s disease. Nat. Neurosci. 13, 567–576 (2010).
Zhang, S., Feany, M. B., Saraswati, S., Littleton, J. T. & Perrimon, N. Inactivation of Drosophila Huntingtin affects long-term adult functioning and the pathogenesis of a Huntington’s disease model. Dis. Model Mech. 2, 247–266 (2009).
Pircs, K. et al. Advantages and limitations of different p62-based assays for estimating autophagic activity in Drosophila. PloS ONE 7, e44214 (2012).
Bartlett, B. J. et al. p62, Ref(2)P and ubiquitinated proteins are conserved markers of neuronal aging, aggregate formation and progressive autophagic defects. Autophagy 7, 572–583 (2011).
Nezis, I. P. et al. Autophagic degradation of dBruce controls DNA fragmentation in nurse cells during late Drosophila melanogaster oogenesis. J. Cell Biol. 190, 523–531 (2010).
Rubinsztein, D. C. The roles of intracellular protein-degradation pathways in neurodegeneration. Nature 443, 780–786 (2006).
Dolan, P. J. & Johnson, G. V. A caspase cleaved form of tau is preferentially degraded through the autophagy pathway. J. Biol. Chem. 285, 21978–21987 (2010).
Mortimore, G. E. & Poso, A. R. Intracellular protein catabolism and its control during nutrient deprivation and supply. Annu. Rev. Nutr. 7, 539–564 (1987).
Mauvezin, C., Ayala, C., Braden, C. R., Kim, J. & Neufeld, T. P. Assays to monitor autophagy in Drosophila. Methods 68, 134–139 (2014).
Dennis, P. B. & Mercer, C. A. The GST-BHMT assay and related assays for autophagy. Methods Enzymol. 452, 97–118 (2009).
Singh, R. et al. Autophagy regulates lipid metabolism. Nature 458, 1131–1135 (2009).
Koga, H., Kaushik, S. & Cuervo, A. M. Altered lipid content inhibits autophagic vesicular fusion. FASEB J. 24, 3052–3065 (2010).
Zhang, H. et al. Elucidating a normal function of huntingtin by functional and microarray analysis of huntingtin-null mouse embryonic fibroblasts. BMC Neurosci. 9, 38 Special section p1-15 (2008).
Ganley, I. G. et al. ULK1.ATG13.FIP200 complex mediates mTOR signaling and is essential for autophagy. J. Biol. Chem. 284, 12297–12305 (2009).
Bjorkoy, G. et al. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J. Cell Biol. 171, 603–614 (2005).
Itakura, E. & Mizushima, N. p62 targeting to the autophagosome formation site requires self-oligomerization but not LC3 binding. J. Cell Biol. 192, 17–27 (2011).
Li, Z., Karlovich, C. A., Fish, M. P., Scott, M. P. & Myers, R. M. A putative Drosophila homolog of the Huntington’s disease gene. Hum. Mol. Genet. 8, 1807–1815 (1999).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 282, 24131–24145 (2007).
Newton, K. et al. Ubiquitin chain editing revealed by polyubiquitin linkage-specific antibodies. Cell 134, 668–678 (2008).
Kirkin, V., McEwan, D. G., Novak, I. & Dikic, I. A role for ubiquitin in selective autophagy. Mol. Cell 34, 259–269 (2009).
Egan, D. F. et al. Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 331, 456–461 (2011).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, 132–141 (2011).
Shang, L. et al. Nutrient starvation elicits an acute autophagic response mediated by Ulk1 dephosphorylation and its subsequent dissociation from AMPK. Proc. Natl Acad. Sci. USA 108, 4788–4793 (2011).
Alers, S., Loffler, A. S., Wesselborg, S. & Stork, B. Role of AMPK-mTOR-Ulk1/2 in the regulation of autophagy: cross talk, shortcuts, and feedbacks. Mol. Cell Biol. 32, 2–11 (2012).
Laplante, M. & Sabatini, D. M. mTOR signaling in growth control and disease. Cell 149, 274–293 (2012).
Kundu, M. et al. Ulk1 plays a critical role in the autophagic clearance of mitochondria and ribosomes during reticulocyte maturation. Blood 112, 1493–1502 (2008).
Seibenhener, M. L. et al. Sequestosome 1/p62 is a polyubiquitin chain binding protein involved in ubiquitin proteasome degradation. Mol. Cell Biol. 24, 8055–8068 (2004).
Andrade, M. A. & Bork, P. HEAT repeats in the Huntington’s disease protein. Nat. Genet. 11, 115–116 (1995).
Ehrnhoefer, D. E., Sutton, L. & Hayden, M. R. Small changes, big impact: posttranslational modifications and function of huntingtin in Huntington disease. The Neuroscientist: A Rev. J. Neurobiol. Neurol. Psychiatry 17, 475–492 (2011).
Jeong, H. et al. Acetylation targets mutant huntingtin to autophagosomes for degradation. Cell 137, 60–72 (2009).
Martin, D. D. et al. Identification of a post-translationally myristoylated autophagy-inducing domain released by caspase cleavage of huntingtin. Hum. Mol. Genet. 23, 3166–3179 (2014).
Schilling, B. et al. Huntingtin phosphorylation sites mapped by mass spectrometry. Modulation of cleavage and toxicity. J. Biol. Chem. 281, 23686–23697 (2006).
Steffan, J. S. et al. SUMO modification of Huntingtin and Huntington’s disease pathology. Science 304, 100–104 (2004).
Wellington, C. L. & Hayden, M. R. Caspases and neurodegeneration: on the cutting edge of new therapeutic approaches. Clin. Genet. 57, 1–10 (2000).
Wellington, C. L. et al. Inhibiting caspase cleavage of huntingtin reduces toxicity and aggregate formation in neuronal and nonneuronal cells. J. Biol. Chem. 275, 19831–19838 (2000).
Yanai, A. et al. Palmitoylation of huntingtin by HIP14 is essential for its trafficking and function. Nat. Neurosci. 9, 824–831 (2006).
Duyao, M. P. et al. Inactivation of the mouse Huntington’s disease gene homolog Hdh. Science 269, 407–410 (1995).
Nasir, J. et al. Targeted disruption of the Huntington’s disease gene results in embryonic lethality and behavioral and morphological changes in heterozygotes. Cell 81, 811–823 (1995).
Zeitlin, S., Liu, J. P., Chapman, D. L., Papaioannou, V. E. & Efstratiadis, A. Increased apoptosis and early embryonic lethality in mice nullizygous for the Huntington’s disease gene homologue. Nat. Genet. 11, 155–163 (1995).
Duran, A. et al. The atypical PKC-interacting protein p62 is an important mediator of RANK-activated osteoclastogenesis. Dev. Cell 6, 303–309 (2004).
Rodriguez, A. et al. Mature-onset obesity and insulin resistance in mice deficient in the signaling adapter p62. Cell Metab. 3, 211–222 (2006).
Cheong, H., Lindsten, T., Wu, J., Lu, C. & Thompson, C. B. Ammonia-induced autophagy is independent of ULK1/ULK2 kinases. Proc. Natl Acad. Sci. USA 108, 11121–11126 (2011).
Lee, E. J. & Tournier, C. The requirement of uncoordinated 51-like kinase 1 (ULK1) and ULK2 in the regulation of autophagy. Autophagy 7, 689–695 (2011).
Reiner, A., Dragatsis, I., Zeitlin, S. & Goldowitz, D. Wild-type huntingtin plays a role in brain development and neuronal survival. Mol. Neurobiol. 28, 259–276 (2003).
Dragatsis, I., Efstratiadis, A. & Zeitlin, S. Mouse mutant embryos lacking huntingtin are rescued from lethality by wild-type extraembryonic tissues. Development 125, 1529–1539 (1998).
Cheong, H. et al. Analysis of a lung defect in autophagy-deficient mouse strains. Autophagy 10, 45–56 (2014).
Ramesh Babu, J. et al. Genetic inactivation of p62 leads to accumulation of hyperphosphorylated tau and neurodegeneration. J. Neurochem. 106, 107–120 (2008).
Babu, J. R., Geetha, T. & Wooten, M. W. Sequestosome 1/p62 shuttles polyubiquitinated tau for proteasomal degradation. J. Neurochem. 94, 192–203 (2005).
Dragatsis, I., Levine, M. S. & Zeitlin, S. Inactivation of Hdh in the brain and testis results in progressive neurodegeneration and sterility in mice. Nat. Genet. 26, 300–306 (2000).
White, J. K. et al. Huntingtin is required for neurogenesis and is not impaired by the Huntington’s disease CAG expansion. Nat. Genet. 17, 404–410 (1997).
Kegel, K. B. et al. Huntingtin expression stimulates endosomal-lysosomal activity, endosome tubulation, and autophagy. J. Neurosci. 20, 7268–7278 (2000).
Steffan, J. S. Does Huntingtin play a role in selective macroautophagy? Cell Cycle 9, 3401–3413 (2010).
Wong, Y. C. & Holzbaur, E. L. The regulation of autophagosome dynamics by huntingtin and HAP1 is disrupted by expression of mutant huntingtin, leading to defective cargo degradation. J. Neurosci. 34, 1293–1305 (2014).
Zheng, S. et al. Deletion of the huntingtin polyglutamine stretch enhances neuronal autophagy and longevity in mice. PLoS Genet. 6, e1000838 (2010).
Scott, R. C., Juhasz, G. & Neufeld, T. P. Direct induction of autophagy by Atg1 inhibits cell growth and induces apoptotic cell death. Curr. Biol. 17, 1–11 (2007).
Pandey, U. B. et al. HDAC6 rescues neurodegeneration and provides an essential link between autophagy and the UPS. Nature 447, 859–863 (2007).
Venken, K. J. et al. MiMIC: a highly versatile transposon insertion resource for engineering Drosophila melanogaster genes. Nat. Methods 8, 737–743 (2011).
Carre-Mlouka, A. et al. Control of sigma virus multiplication by the ref(2)P gene of Drosophila melanogaster: An in vivo study of the PB1 domain of Ref(2)P. Genetics 176, 409–419 (2007).
Sullivan, W., Ashburner, M. & Hawley, R. S. Drosophila Protocols 240–241 (Cold Spring Harbor Laboratory Press, 2000) Ch. 13, Protocol 13.5
Benzer, S. Behavioral mutants of Drosophila isolated by countercurrent distribution. Proc. Natl Acad. Sci. USA 58, 1112–1119 (1967).
Kamikouchi, A. et al. The neural basis of Drosophila gravity-sensing and hearing. Nature 458, 165–171 (2009).
Juhasz, G. & Neufeld, T. P. Experimental control and characterization of autophagy in Drosophila. Methods Mol. Biol. 445, 125–133 (2008).
Wilk, S. & Orlowski, M. Evidence that pituitary cation-sensitive neutral endopeptidase is a multicatalytic protease complex. J. Neurochem. 40, 842–849 (1983).
Cumming, R. C., Simonsen, A. & Finley, K. D. Quantitative analysis of autophagic activity in Drosophila neural tissues by measuring the turnover rates of pathway substrates. Methods Enzymol. 451, 639–651 (2008).
Kimura, S., Fujita, N., Noda, T. & Yoshimori, T. Monitoring autophagy in mammalian cultured cells through the dynamics of LC3. Methods Enzymol. 452, 1–12 (2009).
Bjorkoy, G. et al. Monitoring autophagic degradation of p62/SQSTM1. Methods Enzymol. 452, 181–197 (2009).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 8, 445–544 (2012).
Pampliega, O. et al. Functional interaction between autophagy and ciliogenesis. Nature 502, 194–200 (2013).
Godin, J. D. et al. Huntingtin is required for mitotic spindle orientation and mammalian neurogenesis. Neuron 67, 392–406 (2010).
Massey, A. C., Follenzi, A., Kiffin, R., Zhang, C. & Cuervo, A. M. Early cellular changes after blockage of chaperone-mediated autophagy. Autophagy 4, 442–456 (2008).
Ikenoue, T., Hong, S. & Inoki, K. Monitoring mammalian target of rapamycin (mTOR) activity. Methods Enzymol. 452, 165–180 (2009).
Bejarano, E. et al. Connexins modulate autophagosome biogenesis. Nat. Cell Biol. 16, 401–414 (2014).
Acknowledgements
We are grateful to T. Neufeld and U. Pandey for their fly lines; P. B. Dennis for the pRK5-GST-BHMT plasmid; D. Contamine for anti-Ref(2)P antibody; I. Bezprozvanny for MEF-Htt-WT and MEF-Htt-KO lines; M. Kundu for MEF-ULK1-KO and MEF-ULK1-WT lines; J. Botas for UAS–hHtt DNA and fly lines. We also thank Z. Mao for technical assistance in confocal microscopy, and Z. Sun and N. Perrimon for critically reading the manuscript. H.J.B. is an investigator of the HHMI and supported by the R&R Belfer and Hufftington foundations. G.D. was supported by a T32 developmental biology training grant from the NICHD. This work was supported by NIH grants R01-NS069880 (to S.Z.) and P01-AG031782 and AG038072 (to A.M.C.) and the generous support of R&R Belfer (to A.M.C.).
Author information
Authors and Affiliations
Contributions
Y-N.R. and Z.X. designed and performed most Drosophila studies and most of the studies on starvation- and MG132-induced autophagy response and Tau-ΔC degradation in mammalian cells; B.P. designed and performed part of the studies on autophagic flux in mammalian cells, and most of the studies of lipophagy and mitophagy, some of the co-immunoprecipitation studies and all the electron microscopy studies and morphometric analysis; Z.C., D.C., A.T., E.F.S. and Y.S. contributed to part of these studies; G.D. performed larva starvation, LysoTracker staining and LC3 reporter assay; G.D. and H.J.B. designed experiments, analysed data and contributed to part of the writing and revision of the manuscript; A.M.C. coordinated the study, designed experiments, analysed data and contributed to the writing and revision of the manuscript. S.Z. coordinated the study, designed experiments, analysed data and contributed to the main writing and revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Ectopic Tau expression reveals genetic interaction between dhtt and autophagy pathway.
(a) Tau expression induced the collapsed thorax phenotype in homozygous dhttko−/− mutants (arrow in a3) that can be rescued by the presence of a dhttRescue transgene (a4). Bright field images of adult thorax, dorsal view (pooled from n = 3 independent experiments). (b,c) Tau expression induced more severe mobility (b) and viability phenotypes (c) in dhttko mutants that can be rescued by the presence of a dhttRescue transgene (pooled from n = 3 independent experiments). (d,e) dhtt genetically interacts with multiple autophagy pathway components. Bright field images of adult thorax, dorsal view. Tau expression induced the collapsed thorax phenotype both in homozygous dhttko−/− (arrow in a3) and in atg8ad4−/− (arrow in d7) mutant flies. Similar phenotype was also observed in Tau-expressing flies carrying double heterozygous mutations for both dhttko and one of the following mutant alleles: atg8ad4 (d5), ref(2)pc003993 (e3) or atg1Δ3D (e5), but not in all other controls. The penetrance of the phenotype was quantified as bar graphs to the right (d) or below (e). Genotypes are as indicated (pooled from n = 3 independent experiments). All values are mean + s.e.m. and differences are significant for ∗P < 0.05 using analysis of variance + Bonferroni test. Scale bars: 100 μm.
Supplementary Figure 2 dhtt does not affect proteasome activity and starvation response in Drosophila.
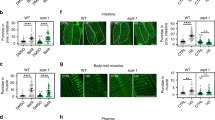
(a) Increased accumulation of total ubiquitinated proteins in aged (5-week-old) dhttko−/− mutants compared to aged-matched controls (wildtype (WT) and dhttko−/− carrying a wildtype dhtt rescue transgene dhttRescue). Young flies (1-week-old) showed similar low levels of total ubiquitinated proteins in all the three genotypes. Bar graph below shows the quantification of the levels of total ubiquitinated proteins detected with anti-ubiquitin antibody by western blotting (n = 3 independent experiments). (b and c) Abnormal eye phenotypes (b) and increase ubiquitination (c) associated with proteasome dysfunction. (b) Bright field images of adult fly eyes. Note the rough eye phenotype due to proteasome inactivation by DTS5 or DTS7, two temperature-sensitive, dominant-negative alleles of proteasome subunits at the non-permissive 28 °C, as compared to the normal eye morphology in flies of the same genotypes raised at permissive temperature of 22 °C (compare a2 and a3 with a5 and a6). GMR-Gal4 driver was used to direct eye-specific expression of UAS-DTS5 and DTS7. LacZ was used as negative control (a1 and a4). Scale bar: 50 μm. Anterior up and ventral to the right in all panels. n > 15 fly eyes from 3 independent experiments. (c) Total proteins from flies of the indicated genotypes were extracted by lysis buffer containing 1% Triton X-100 and separated by 10% SDS-PAGE gel. The level of ubiquitination was revealed by anti-Ub antibody (FK2). Actin served as loading control. Note the significantly increased levels of ubiquitinated proteins in flies expressing the dominant-negative DTS7 mutant driven by arm-Gal4 under restrictive temperature of 28 °C but not at 22 °C or in controls (n = 3 independent experiments). (d) Normal thorax morphology (top panels) and internal muscle structures (bottom panels) in flies expressing Tau together with the dominant-negative DTS5 or DTS7 alleles of proteasome subunits at either permissive 22 °C or non-permissive 28 °C temperatures, as indicated. Top panels: bright field images of external thorax morphology; bottom panels: cross section views of underlying muscle structures, as revealed by confocal imaging of F-Actin patterning from phalloidin staining (bottom). Note the similar smooth thorax surface and intact internal muscle structures in all panels. Simultaneous expression of Tau and DTS5 or DTS7 were driven by A307-Gal4 line, as indicated. Scale bar: 100 μm. n > 10 fly bodies from 3 independent experiments. (e) Normal UPS activity in dhttko−/− mutants. No obvious accumulation of UPS reporter GFP-CL1 signal in dhttko−/− mutants in either the eye imaginal discs of third instar larvae (c2 and c4) or 3-day-old adult brains (c6 and c8) at either 22 °C or 28 °C. In contrast, in the positive controls that expressed a temperature-sensitive dominant-negative UPS mutant (DTS7) at a non-permissive temperature of 28 °C (c3 and c7), there is a strong accumulation of the UPS reporter GFP-CL1. Scale bar: 100 μm. n > 8 fly samples from 3 independent experiments. (f) Normal catalytic activities of proteasome subunits in dhttko−/− mutants. Caspase-like, trypsin-like and chymotrypsin-like peptidase activities were measured using whole-animal extracts from young (5-day-old) or old (40-day-old) wild-type and dhttko−/− flies, and their relative activities are presented with results from young wildtype flies as standard value of 1. n. s. : no statistical significance. n = 18 fly extracts for each genotype, pooled from 3 independent experiments. (g,h) dhtt is not required for the starvation-induced autophagy. Autophagosomes and autolysosome in fat bodies of 80-hour larvae were labeled with mCherry-Atg8a (red). Starvation induced similar strong increase of red puncta (autophagosomes and autolysosome) in both the wildtype and dhttko−/− mutants, as quantified in the bar graphs to the right. UAS-mCherry-Atg8a reporter was driven by the fat body-specific Cg-Gal4 driver. Tissues were double-labeled with DAPI (blue) for cell nuclei. (h) Bar graph shows the quantification of the average number of mCherry-Atg8a-positive punctae. n = 60 cells for each genotype, pooled from 3 independent experiments. (i) Starvation-resistance assay. Newly eclosed or 5-day-old adult flies were subjected to starvation. dhttko−/− mutants showed similar survival curve as wildtype control (CS, Canton S). n = 155 flies (newly eclosed CS), n = 160 (newly eclosed dhttko−/−), n = 203 (5 days CS), n = 180 (5 days dhttko−/−), pooled from 3 independent experiments. All values are mean + s.e.m. and differences are significant for ∗P < 0.05 using Student’s t-test or analysis of variance + Bonferroni test.
Supplementary Figure 3 Human Htt rescues dhttko mutant-associated phenotypes.
(a) Schematics of the C-terminus truncated Tau (Tau-ΔC, missing last 49 amino acids) used in the study, as compared to the full-length (FL) Tau. Below: western blotting analysis to confirm the expression of Tau deletion used in the study. Note the smaller size of the C-terminus truncated Tau (Tau-ΔC), as compared to that of the full-length Tau. Both Tau transgenes are tagged with GFP epitope at their C-termini. In (a-d), the ubiquitous arm arm-Gal4 driver was used to drive the ectopic Tau expression, protein expression was examined by anti-GFP antibody, with Actin served as loading control. + / + : samples from wildtype non-transgenic flies. (b) Confirmation of the proper expression of endogenous dHtt as well as the human full-length Huntingtin (hHtt) and GFP proteins from the corresponding transgenes by Western blotting analyses. The ubiquitous arm-Gal4 driver was used to drive the hHtt and GFP transgene expression. n = 3 independent experiments. (c and d) Ectopic expression of full-length human Huntingtin (hHtt) significantly rescued several dhttko associated phenotypes, including (c) the reduced mobility and (d) shortened lifespan of mutant adults. N values are indicated in the figure and were pooled from 4 independent experiments. (e and f) Ectopic expression of hHtt significantly rescued Tau-induced phenotypes in dhttko mutant background, including the exacerbated (e) mobility decline and (f) reduced lifespan. A307-Gal4 driver was used to drive the Tau-ΔC-GFP and hHtt expression. N values are indicated in the figure and were pooled from n = 4 independent experiments. (g–i) hHtt partially suppressed the increased accumulations of (g–h) total ubiquitinated proteins and (f) Tau-ΔC in dhttko−/− mutant flies. Ubiquitous arm-Gal4 driver was used to drive the hHtt expression; Tau-ΔC level was examined by anti-GFP antibody, with Actin serving as loading control. n = 3 independent experiments. All values are mean + s.e.m. and differences are significant for ∗P < 0.05 using analysis of variance + Bonferroni test.
Supplementary Figure 4 Htt knockdown does not affect proteasomal activity or lysosomal function.
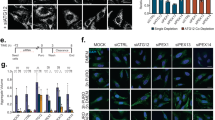
(a–c) Htt knockdown does not affect the proteasome activity in mammalian cells. Proteasome activity in HEK293T cells was evaluated by examining the level of (a) a UPS reporter CL1-GFP and (b) ubiquitinated β-catenin, a bona fide substrate of proteasome. Htt knockdown by two independent siRNA did not cause an abnormal accumulation of (a) GFP-CL1 (lanes 3 and 4) or (b) ubiquitinated β-catenin, resembling that from samples treated with the control siRNA (Ctrl, lane 2) or autophagy inhibitor BafA1 (lane 6). In contrast, treatment with proteasome inhibitor MG132 induced a higher level of both (a) GFP-CL1 (lanes 5) and (b) ubiquitinated β-catenin. BafA1 was applied at 100 nM for two hours and MG132 was applied at 10 μM for 6 hours before the assay. n = 3 independent experiments. (c) Normal catalytic activities of proteasome subunits in Htt knock down NIH3T3 fibroblasts. Caspase-like, trypsin-like and chymotrypsin-like peptidase activities were measured using whole cell extracts from HEK293T cells treated with si-Ctrl, si-Htt or MG132. Their relative activities are presented with results from si-Ctrl cells as the standard of 1. n. s. : no statistical significance. n = 3 independent experiments. (d–h) Htt knockdown does not affect lysosomal function. (d) Staining of acid compartments with LysoTracker does not reveal problems of acidification in lysosomes although in agreement with the electromicroscopy studies shows some level of expansion of these acid compartments. (e–f) Internalization of fluorescent transferrin is comparable in both cell types supporting the absence of major alterations in the endocytic system. (e) Representative images of Transferrin loaded cells. Inserts: higher magnification images of the transferring channel and (f) quantification of the number of transferrin positive puncta per cell. n = 3 independent experiments where 80 cells were examined in total from 3 independent fields in each condition. (g,h) Levels of LAMPs are not reduced in Htt(-) cells, in fact a slight increase in levels of LAMP1 as measured by immunoblot (g) and of LAMP-2 positive puncta (h) was noticeable in Htt(-) cells either in the presence or absence of serum. Scale bar: 10 μm. (i–k) Time-course analyses of the levels of Tau-ΔC-GFP protein expressed from stably transfected Dox-inducible HeLa cell line. Degradation of Tau-ΔC-GFP was significantly delayed by (i) autophagy inhibitor 3-MA or (j) by siRNA-mediated knockdown of ATG7, whereas (k) proteasome inhibitor MG132 showed little effect. Control cells were treated with only water (i), scramble siRNA (j) or DMSO (k). Quantification of multiple repeat experiments is shown at the bottom n = 9 dishes for each condition pooled from 3 independent experiments. The difference was most evident around 48 hours after induction of Tau-ΔC-GFP expression, which was detected by anti-GFP antibody. Actin served as loading control. In i and j, additional panels at the bottom show reduced LC3 lipidation upon addition of 3-MA or Atg7 siRNA supporting the efficacy of these treatments in bloking autophagy (note that due to the low basal steady-state levels of LC3-II in these cells, Baf1 was added to detect changes in LC3 lipidation). All values are mean + s.e.m. and differences are significant for ∗P < 0.05 using analysis of variance + Bonferroni test (c) or student’s t-test (I,j,k,f).
Supplementary Figure 5 Effect of Htt depletion on autophagic flux in mammalian cells.
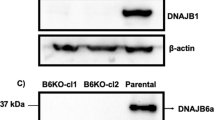
(a,b) Htt does not affect the basal level of autophagic activity in dividing mammalian HEK293T cells. (a) LC3 lipidation assay. Compared to controls, Htt knockdown did not have obvious effect on the level of lipidated LC3 (LC3-II) either in the absence or the presence of lysosomal inhibitor BafA1, which blocks autophagy flux and increases cellular LC3-II levels. (b) GST-BHMT assay. Htt knockdown in the same cells did not show obvious effect on the level of the cleaved GST-BHMT-FRAG product. MG132-treated samples were included as positive controls (lanes 5 and 10 in (a) and lane 5 in (b)), which showed robust induction of LC3-II and GST-BHMT fragmentation. Actin served as loading control. n = 3 independent experiments. (c) Restored autophagosome formation by GFP-Htt overexpression in Htt-knockdown HeLa cells. Immunofluorescent staining for endogenous LC3 (red) in HeLa wildtype (Ctrl), or si-Htt interfered cells (Htt-) untreated or co-transfected with GFP-Htt cDNA (GFP-Htt). Cells were cultured in the presence or absence of MG132 for 6 hours and/or BafA1 for 2 hours before harvesting. Overexpression of GFP-Htt, confirmed by anti-GFP antibody staining (green), rescued the reduced number of LC3-positive puncta in MG132-treated Htt (-) cells, as shown by quantification at the bottom. n = 3 independent experiments where a total of 120 cells were analyzed per condition. Scale Bar: 5 μm. (d) Restoration of Htt in MEFs from Htt knockout (Htt-KO) mice rescues defective autophagosome formation during induction of selective autophagy. Htt-KO MEFs were transiently transfected with a plasmid carrying GFP-Htt and subjected to the indicated treatments. Images show staining for LC3 with a Cy5-conjugated secondary antibody. Representative regions containing a transfected and untransfected cell are shown as merge channels (left) and red channel only (bottom). Nuclei are highlighted with DAPI. Scale Bar: 5 μm. Bottom shows quantification of the average number of LC3 puncta per cell in both cell populations. n = 3 independent experiments where a total of 120 cells were analyzed per condition. (e) Htt affects MG132-induced autophagosome biogenesis but not its fusion with lysosomes. HeLa cells were double labeled for LC3 (green) and lysosome marker LAMP1 (red). BafA1 treatment led to a significant lysosomal accumulation of LC3. Knockdown of Htt expression by two independent siRNA both significantly suppressed the MG132-induced formation of LC3-positive puncta, but did not affect the accumulation of remaining LC3 in the lysosomes. The levels of co-localization between LC3 and LAMP1 are quantified in bar graphs (bottom). Scale Bar: 2 μm. n = 3 independent experiments where a total of 45 (in none), 90 (in MG132) and 100 (in Serum-) cells were analyzed per condition. (f) Htt does not affect starvation-induced autophagy in dividing mammalian HEK293T cells. Compared to controls, Htt knockdown did not have obvious effect on the level of lipidated LC3 (LC3-II) either in the absence or the presence of lysosomal inhibitor BafA1. HeLa cells cultured in nutrient rich medium (RM) or EBSS medium were used for the studies. Untreated (RM) and control siRNA (si-Ctrl)-treated samples were included as negative controls. n = 3 independent experiments. (g–k) Depletion of Htt in different types of mammalian cells reduces selective autophagic flux but not starvation-induced autophagic flux. Control and Htt knock-down (Htt(-)) NIH3T3 (g), N2a (i) or mouse striatal derived (j,k) cells and MEFs from wild-type or Htt knock-out mice (Htt KO) (h) were transduced with a vector encoding mCherry-GPF-LC3 and subjected to the indicated stimuli to activate nutrient-induced autophagy (serum- and EBSS) or selective autophagy in response to proteostasis (Lactacystin, MG132), lipotoxicity (oleic) or mitochondria depolarization (FCCP) as labeled. Cells were imaged using high content microscopy and autophagic flux was determined as conversion of autophagosomes (yellow puncta) into autolysosomes (red only puncta). n = 3 independent experiments where a total of 2250 (g–i) and 3000 (j) cells were analyzed per condition. Representative images of mouse striatal-derived cells upon the indicated treatments. Merged channels are shown. Nuclei are highlighted with DAPI. Scale bar: 5 μm. All values are mean + s.e.m. and differences are significant for ∗P < 0.05 using analysis of variance + Bonferroni test (e) or student’s t-test (c,d,g–j).
Supplementary Figure 6 Structural-functional analysis of Htt interaction with p62 and ULK1 proteins.
(a–d) Htt physically interacts with p62 and ULK1 in HEK293T cells. Reciprocal co-IP experiments between (a and b) HA- or FLAG-tagged Htt with FLAG- or HA-tagged p62, or (c and d) between HA- or Myc-tagged Htt with FLAG- or HA-tagged ULK1, as indicated. Htt was detected in the same immunoprecipitates both with p62 and with ULK1 in co-IP assays from either direction (lanes 2). Lanes 4 are positive controls for Myc-tagged mHAP1 (a and c), a known Htt-associated protein, Myc-tagged LC3 (b), a known p62 interactor, or FLAG-tagged Atg13 (d), a known ULK1 binding partner. n = 3 independent experiments. (e) Schematics of putative functional and structural motifs in human Htt protein: N17 (grey color): the first 17 amino acids of Htt protein, which is right in front of the polyglutamine tract (polyQ, blue color); PolyP (yellow): the proline-rich region adjacent to polyQ tract; NES: nuclear export signal. The predicted 40 HEAT repeat in the Htt protein was presented as red cylinders and numbered accordingly13. (f) Schematic illustration of the five most conserved regions in human Htt protein (labeled alphabetically from “a” to “e”) that bear the highest sequence similarity with its Drosophila Htt counterpart. (g) Schematics of the nine Htt deletions used in the study (top), designated as NT (N-terminal), MD (middle), CT (C-terminal), and D2 to D7 regions, as illustrated. The deletions were designed based on the distribution of the predicted structural motifs and conserved sequences as illustrated in (e) and (f).
Supplementary Figure 7 Htt facilitates the cargo recognition efficiency by p62 protein.
(a) Time-course analysis of p62 distribution in Triton X-100 soluble (top) and insoluble fractions (middle), as well as total p62 protein (bottom) in HEK293T cells after MG132 treatment, as indicated. Actin served as loading control. Note the gradual decrease of soluble p62 with concomitant increase of insoluble p62, as well as the marked increase of total p62 at hour 12. n = 3 independent experiments. (b and c) MG132-induced turnover of soluble p62 is autophagy-dependent. (b) Beclin 1 knockdown in HEK293T cells suppressed MG132-induced depletion of Triton X-100-soluble p62 (compare lanes 3 with 2). 5 nM siRNA against beclin 1 was effective in reducing the expression of Beclin 1 protein (middle panels). (c) Bar graph representation of the soluble p62 levels, which were quantified and normalized against loading control Actin, as indicated. Scale bars indicate s.e.m. n = 9 dishes for each condition pooled from 3 independent experiments ∗p < 0.05 using analysis of variance + Bonferroni test. (d) Htt does not affect p62 level under basal and starvation conditions in HEK293T cells. Under both conditions, si-Htt treated samples showed similar levels of p62 protein as in controls of mock- or control siRNA-treated samples. Note the overall reduced level of p62 protein under starvation condition (EBSS incubation). n = 3 independent experiments. (e) Htt does not affect p62 self-polymerization. in vitro co-IP assay to assess the interaction between HA- and Myc-tagged p62 in transfected HEK293T cells. Similar levels of Myc-p62 were pulled down by HA-p62 between the siHtt-treated (lanes 3 and 4) and control siRNA-treated (lane 2) samples. n = 3 independent experiments. (f–i) Htt specifically affects the binding affinity between endogenous p62 protein and substrates with Ub-K63, but not Ub-K48, modifications. Co-IP experiments against endogenous ubiquitin using (f) anti-Ub-K63 or (g) anti-Ub-K48 from HEK293T cells, or using anti-HA antibody from HEK293T cells transfected with HA-tagged (h) Ub-K63 (HA-Ub-K63) or (i) Ub-K48 (HA-Ub-K48). In both cells, Htt knockdown significantly reduced the amount of endogenous p62 that co-immunoprecipitated with Ub-K63 modified proteins, but showed no obvious effect on that co-immunoprecipitated with Ub-K48 modified proteins (compare lanes 2 and 3 with lane 1 in both (f) and in (g), and in both (h) and in (i)). n = 3 independent experiments.
Supplementary Figure 8 Htt forms distinctive complex with ULK1.
(a and b) Htt does not affect the binding and phosphorylation of ULK1 by AMPK. (a) Co-IP experiment using anti-FLAG antibody to examine the physical interactions between ULK1 and AMPK, as indicated. HEK293T cells were transfected with FLAG-tagged ULK1 together with HA-tagged AMPKα, AMPKβ, AMPKγ subunits. Htt knockdown showed no effect on the levels of AMPK subunits co-immunoprecipitated with ULK1. WCE: whole cell extract. (b) Htt knockdown in HEK293T cells did not affect the phenformin-stimulated phosphorylation of ULK1 at the targeted Ser555 site (ULK1-S555) by AMPK (compare lanes 3 and 4 with lane 2). Samples were treated in 1 μM phenformin for 2 hours before the assay. A constitutive active AMPK (AMPK-CA) was included as the positive control for the kinase assay (lane 5). n = 3 independent experiments. (c) Htt knockdown does not affect the stability of the ULK1-FIP200-Atg13-Atg101 complex. Co-IP experiment in HEK293T cells to assess the effect of Htt knockdown on the physical associations among the ULK1 complex components. HA-tagged ULK1 was co-transfected with other ULK1 complex components FIP200, Atg13 and Atg101, followed by co-IP using anti-HA antibody, as indicated. Htt knockdown by two independent siRNA showed no obvious effect on the binding affinity among these ULK1 complex components, resembling that observed from the control siRNA treated sample (compare lanes 3 and 4 with lane 2). n = 3 independent experiments. (d–f) Htt does not affect mTOR kinase activity. (d and e) in vivo mTOR kinase assay. Neither (d) Htt knock-down nor (e) Htt overexpression in HEK293T cells showed any obvious effect on the mTOR kinase activity in vivo. (e) Htt overexpression also did not affect basal mTOR kinase activity (compare lane 3 and 4 with lane 1) or Rheb-induced mTOR activation (compare lane 5 with lane 2). The phosphorylation level of S6K at Thr389 (p-S6K (Thr389)) was used as the readout of in vivo mTOR kinase activity. Upstream mTOR activator Rheb was included as the positive control for the kinase assay. (f) in vitro mTOR kinase assay. Htt knockdown did not affect mTOR kinase activity in vitro (compare lanes 3 and 4 with lane 2). 4E-BP1 was used as the substrate and its phosphorylation level at position Ser65 (p-4E-BP1 (S65)) was used as the readout of in vitro mTOR kinase activity. A dominant-negative kinase dead (KD) mutant of mTOR (Myc-mTOR-KD) was included as negative control for the assay. n = 3 independent experiments. (g–i) Endogenous co-IP experiments in HEK293T cells using (g) anti-ULK1, (h) anti-Htt or (i) anti-mTOR antibodies. (g) ULK1 co-immunoprecipitated with both endogenous Htt and mTOR (lane 4). (h) Htt co-immunoprecipitated with endogenous ULK1 but not mTOR (lane 4). (i) mTOR co-immunoprecipitated with endogenous ULK1 but not Htt (lane 4). Cells treated with siRNA against ULK1 (g), Htt (h) or mTOR (i) were included as negative controls. n = 3 independent experiments. (j and k) Increased ULK1-Htt complex and concomitant loss of ULK1-mTOR complex after lipotoxic or mitotoxic challenges. HEK293T cells co-transfected with Myc-mTOR and Flag-ULK1 were incubated with (j) 200uM Oleic acid (OA) for 12 hours or (k) with 10uM FCCP for 1 hour before harvesting for co-IP experiments using anti-FLAG antibody. Under either basal or OA and FCCP challenges, FLAG-tagged ULK1 could immunoprecipitate both Myc-tagged mTOR and endogenous Htt. However, both (j) lipid and (k) mitochondrial stresses led to a significant higher levels of Htt and the simultaneous loss of mTOR that co-immunoprecipitated with ULK1 (compare lanes 2 and 6 in (j) and (k)), while Htt knockdown largely restored the ULK1 association with mTOR (compare lanes 7 and 8 with lane 6 in (j) and (k)). n = 3 independent experiments. (l,m) Elevated Htt protein levels in response to induction of selective autophagy. Western analyses of Htt protein levels in NIH 3T3 (l) and N2a neuroblastoma (m) cells that were mock treated (None), serum deprived (serum-) or stressed with lactacystin (Lacta), MG132, FCCP or oleic acid, as indicated. Graph in l shows quantification of changes in Htt levels in NIH3T3 cells (n = 7 different cellular extracts for condition from 5 independent experiments. error bars indicate s.e.m., and ∗p < 0.05 compared to None using student’s t-test). Actin served as loading controls.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2699 kb)
Rights and permissions
About this article
Cite this article
Rui, YN., Xu, Z., Patel, B. et al. Huntingtin functions as a scaffold for selective macroautophagy. Nat Cell Biol 17, 262–275 (2015). https://doi.org/10.1038/ncb3101
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3101
This article is cited by
-
Cell-type-specific CAG repeat expansions and toxicity of mutant Huntingtin in human striatum and cerebellum
Nature Genetics (2024)
-
Upregulated pexophagy limits the capacity of selective autophagy
Nature Communications (2024)
-
Autophagy markers, cognitive deficits and depressive symptoms in Parkinson’s disease
Journal of Neural Transmission (2024)
-
Emerging role of lipophagy in liver disorders
Molecular and Cellular Biochemistry (2024)
-
The regulatory role of lipophagy in central nervous system diseases
Cell Death Discovery (2023)