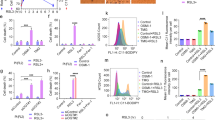
Abstract
Cells rely on autophagy to clear misfolded proteins and damaged organelles to maintain cellular homeostasis. In this study we use the new autophagy inhibitor PIK-III to screen for autophagy substrates. PIK-III is a selective inhibitor of VPS34 that binds a unique hydrophobic pocket not present in related kinases such as PI(3)Kα. PIK-III acutely inhibits autophagy and de novo lipidation of LC3, and leads to the stabilization of autophagy substrates. By performing ubiquitin-affinity proteomics on PIK-III-treated cells we identified substrates including NCOA4, which accumulates in ATG7-deficient cells and co-localizes with autolysosomes. NCOA4 directly binds ferritin heavy chain-1 (FTH1) to target the iron-binding ferritin complex with a relative molecular mass of 450,000 to autolysosomes following starvation or iron depletion. Interestingly, Ncoa4−/− mice exhibit a profound accumulation of iron in splenic macrophages, which are critical for the reutilization of iron from engulfed red blood cells. Taken together, the results of this study provide a new mechanism for selective autophagy of ferritin and reveal a previously unappreciated role for autophagy and NCOA4 in the control of iron homeostasis in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Begun, J. & Xavier, R. J. Autophagy at the crossroads of metabolism and cellular defense. Curr. Opin. Gastroenterol. 29, 588–596 (2013).
Huang, J. & Brumell, J. H. Bacteria–autophagy interplay: a battle for survival. Nat. Rev. Microbiol. 12, 101–114 (2014).
Mizushima, N., Levine, B., Cuervo, A. M. & Klionsky, D. J. Autophagy fights disease through cellular self-digestion. Nature 451, 1069–1075 (2008).
The Wellcome Trust Case Control Consortium, Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Hampe, J. et al. A genome-wide association scan of nonsynonymous SNPs identifies a susceptibility variant for Crohn disease in ATG16L1. Nat. Genet. 39, 207–211 (2007).
Rioux, J. D. et al. Genome-wide association study identifies new susceptibility loci for Crohn disease and implicates autophagy in disease pathogenesis. Nat. Genet. 39, 596–604 (2007).
Metzger, S. et al. Age at onset in Huntington’s disease is modified by the autophagy pathway: implication of the V471A polymorphism in Atg7. Hum. Genet. 128, 453–459 (2010).
Cadwell, K. et al. A key role for autophagy and the autophagy gene Atg16l1 in mouse and human intestinal Paneth cells. Nature 456, 259–263 (2008).
Komatsu, M. et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441, 880–884 (2006).
Rubinsztein, D. C., Codogno, P. & Levine, B. Autophagy modulation as a potential therapeutic target for diverse diseases. Nat. Rev. Drug Discov. 11, 709–730 (2012).
Klionsky, D. J. & Schulman, B. A. Dynamic regulation of macroautophagy by distinctive ubiquitin-like proteins. Nat. Struct. Mol. Biol. 21, 336–345 (2014).
Kihara, A., Noda, T., Ishihara, N. & Ohsumi, Y. Two distinct Vps34 phosphatidylinositol 3-kinase complexes function in autophagy and carboxypeptidase Y sorting in Saccharomyces cerevisiae. J. Cell Biol. 152, 519–530 (2001).
Obara, K., Noda, T., Niimi, K. & Ohsumi, Y. Transport of phosphatidylinositol 3-phosphate into the vacuole via autophagic membranes in Saccharomyces cerevisiae. Genes Cells 13, 537–547 (2008).
Axe, E. L. et al. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J. Cell Biol. 182, 685–701 (2008).
Matsunaga, K. et al. Autophagy requires endoplasmic reticulum targeting of the PI3-kinase complex via Atg14L. J. Cell Biol. 190, 511–521 (2010).
Polson, H. E. et al. Mammalian Atg18 (WIPI2) localizes to omegasome-anchored phagophores and positively regulates LC3 lipidation. Autophagy 6, 506–522 (2010).
Mizushima, N., Yoshimori, T. & Ohsumi, Y. Role of the Apg12 conjugation system in mammalian autophagy. Int. J. Biochem. Cell Biol. 35, 553–561 (2003).
Johansen, T. & Lamark, T. Selective autophagy mediated by autophagic adapter proteins. Autophagy 7, 279–296 (2011).
Rogov, V., Dotsch, V., Johansen, T. & Kirkin, V. Interactions between autophagy receptors and ubiquitin-like proteins form the molecular basis for selective autophagy. Mol. Cell 53, 167–178 (2014).
Shaid, S., Brandts, C. H., Serve, H. & Dikic, I. Ubiquitination and selective autophagy. Cell Death Differ. 20, 21–30 (2013).
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889 (2006).
Komatsu, M. et al. Impairment of starvation-induced and constitutive autophagy in Atg7-deficient mice. J. Cell Biol. 169, 425–434 (2005).
Gillooly, D. J. et al. Localization of phosphatidylinositol 3-phosphate in yeast and mammalian cells. EMBO J. 19, 4577–4588 (2000).
Folkes, A. J. et al. The identification of 2-(1H-indazol-4-yl)-6-(4-methanesulfonyl-piperazin-1-ylmethyl)-4-morpholin-4-yl-thieno[3,2-d]pyrimidine (GDC-0941) as a potent, selective, orally bioavailable inhibitor of class I PI3 kinase for the treatment of cancer. J. Med. Chem. 51, 5522–5532 (2008).
Maira, S. M. et al. Identification and characterization of NVP-BKM120, an orally available pan-class I PI3-kinase inhibitor. Mol. Cancer Ther. 11, 317–328 (2012).
Miller, S. et al. Shaping development of autophagy inhibitors with the structure of the lipid kinase Vps34. Science 327, 1638–1642 (2010).
Nyfeler, B., Bergman, P., Wilson, C. J. & Murphy, L. O. Quantitative visualization of autophagy induction by mTOR inhibitors. Methods Mol. Biol. 821, 239–250 (2012).
Fujita, N. et al. An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure. Mol. Biol. Cell 19, 4651–4659 (2008).
Kirkin, V., McEwan, D. G., Novak, I. & Dikic, I. A role for ubiquitin in selective autophagy. Mol. Cell 34, 259–269 (2009).
Bjorkoy, G. et al. p62/SQSTM1 forms protein aggregates degraded by autophagy and has a protective effect on huntingtin-induced cell death. J. Cell Biol. 171, 603–614 (2005).
Kirkin, V. et al. A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol. Cell 33, 505–516 (2009).
Thurston, T. L., Ryzhakov, G., Bloor, S., von Muhlinen, N. & Randow, F. The TBK1 adaptor and autophagy receptor NDP52 restricts the proliferation of ubiquitin-coated bacteria. Nat. Immu. 10, 1215–1221 (2009).
Newman, A. C. et al. TBK1 kinase addiction in lung cancer cells is mediated via autophagy of Tax1bp1/Ndp52 and non-canonical NF-κB signalling. PLoS ONE 7, e50672 (2012).
Boada-Romero, E. et al. TMEM59 defines a novel ATG16L1-binding motif that promotes local activation of LC3. EMBO J. 32, 566–582 (2013).
Ohzono, C. et al. Nedd4-interacting protein 2, a short half-life membrane protein degraded in lysosomes, negatively controls down-regulation of connexin43. Biol. Pharm. Bull. 33, 951–957 (2010).
Yeh, S. & Chang, C. Cloning and characterization of a specific coactivator, ARA70, for the androgen receptor in human prostate cells. Proc. Natl Acad. Sci. USA 93, 5517–5521 (1996).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Asano, T. et al. Distinct mechanisms of ferritin delivery to lysosomes in iron-depleted and iron-replete cells. Mol. Cell. Biol. 31, 2040–2052 (2011).
Bridges, K. R. Ascorbic acid inhibits lysosomal autophagy of ferritin. J. Biol. Chem. 262, 14773–14778 (1987).
Ericsson, J. L. Studies on induced cellular autophagy. I. Electron microscopy of cells with in vivo labelled lysosomes. Exp. Cell Res. 55, 95–106 (1969).
Mortensen, M. et al. Loss of autophagy in erythroid cells leads to defective removal of mitochondria and severe anemia in vivo. Proc. Natl Acad. Sci. USA 107, 832–837 (2010).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 8, 445–544 (2012).
Seglen, P. O. & Gordon, P. B. 3-Methyladenine: specific inhibitor of autophagic/lysosomal protein degradation in isolated rat hepatocytes. Proc. Natl Acad. Sci. USA 79, 1889–1892 (1982).
Wu, Y. T. et al. Dual role of 3-methyladenine in modulation of autophagy via different temporal patterns of inhibition on class I and III phosphoinositide 3-kinase. J. Biol. Chem. 285, 10850–10861 (2010).
Simonsen, A. & Tooze, S. A. Coordination of membrane events during autophagy by multiple class III PI3-kinase complexes. J. Cell Biol. 186, 773–782 (2009).
Jaber, N. et al. Class III PI3K Vps34 plays an essential role in autophagy and in heart and liver function. Proc. Natl Acad. Sci. USA 109, 2003–2008 (2012).
Morishita, H. et al. Deletion of autophagy-related 5 (Atg5) and Pik3c3 genes in the lens causes cataract independent of programmed organelle degradation. J. Biol. Chem. 288, 11436–11447 (2013).
Parekh, V. V. et al. Impaired autophagy, defective T cell homeostasis, and a wasting syndrome in mice with a T cell-specific deletion of Vps34. J. Immu. 190, 5086–5101 (2013).
Willinger, T. & Flavell, R. A. Canonical autophagy dependent on the class III phosphoinositide-3 kinase Vps34 is required for naive T-cell homeostasis. Proc. Natl Acad. Sci. USA 109, 8670–8675 (2012).
Zhou, X. et al. Deletion of PIK3C3/Vps34 in sensory neurons causes rapid neurodegeneration by disrupting the endosomal but not the autophagic pathway. Proc. Natl Acad. Sci. USA 107, 9424–9429 (2010).
Devereaux, K. et al. Regulation of mammalian autophagy by class II and III PI 3-kinases through PI3P synthesis. PLoS ONE 8, e76405 (2013).
Crichton, R. R. & Declercq, J. P. X-ray structures of ferritins and related proteins. Biochim. Biophys. Acta 1800, 706–718 (2010).
Coffey, J. W. & DeDuve, C. Digestive activity of lysosomes. I. The digestion of proteins by extracts of rat liver lysosomes. J. Biol. Chem. 243, 3255–3263 (1968).
Andrews, S. C., Treffry, A. & Harrison, P. M. Siderosomal ferritin. The missing link between ferritin and haemosiderin? Biochem. J. 245, 439–446 (1987).
Ganz, T. & Nemeth, E. Hepcidin and disorders of iron metabolism. Annu. Rev. Med. 62, 347–360 (2011).
Donovan, A., Roy, C. N. & Andrews, N. C. The ins and outs of iron homeostasis. Physiology (Bethesda) 21, 115–123 (2006).
Kidane, T. Z., Sauble, E. & Linder, M. C. Release of iron from ferritin requires lysosomal activity. Am. J. Physiol. Cell Physiol. 291, C445-455 (2006).
Mancias, J. D., Wang, X., Gygi, S. P., Harper, J. W. & Kimmelman, A. C. Quantitative proteomics identifies NCOA4 as the cargo receptor mediating ferritinophagy. Nature 509, 105–109 (2014).
Gregory, A., Polster, B. J. & Hayflick, S. J. Clinical and genetic delineation of neurodegeneration with brain iron accumulation. J. Med. Genet. 46, 73–80 (2009).
Levi, S., Cozzi, A. & Arosio, P. Neuroferritinopathy: a neurodegenerative disorder associated with L-ferritin mutation. Best Pract. Res. Clin. Haematol. 18, 265–276 (2005).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. 66, 213–221 (2010).
Proenca, C. C. et al. Atg4b-dependent autophagic flux alleviates Huntington’s disease progression. PLoS ONE 8, e68357 (2013).
Wee, S. et al. PTEN-deficient cancers depend on PIK3CB. Proc. Natl Acad. Sci. USA 105, 13057–13062 (2008).
Maira, S. M. et al. Identification and characterization of NVP-BEZ235, a new orally available dual phosphatidylinositol 3-kinase/mammalian target of rapamycin inhibitor with potent in vivo antitumor activity. Mol. Cancer Ther. 7, 1851–1863 (2008).
Nyfeler, B. et al. Relieving autophagy and 4EBP1 from rapamycin resistance. Mol. Cell. Biol. 31, 2867–2876 (2011).
Rush, J. et al. Immunoaffinity profiling of tyrosine phosphorylation in cancer cells. Nat. Biotechnol. 23, 94–101 (2005).
Udeshi, N. D., Mertins, P., Svinkina, T. & Carr, S. A. Large-scale identification of ubiquitination sites by mass spectrometry. Nat. Protoc. 8, 1950–1960 (2013).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Acknowledgements
We would like to thank E. George, A. Donovan, K. Mansfield, L. Klickstein, D. Glass, R. Xavier and M. Fishman for their input and advice.
Author information
Authors and Affiliations
Contributions
W.E.D., B.N., E.T., S.L., Z.W., S.D’A., H.W., D.W., J.K., P.B., S.U. and L.O.M. performed cell biology experiments. J.N., J.T., D.S., and M.S. performed proteomic experiments. R.A.E., J.C., C.L., D.E.B. and M.S.K. performed protein crystallography studies and analysed the co-structural data. P.F., M.E.D. and F.H., performed high-throughput screens, I.C-T., E.H., A.H. and E.P.K. performed medicinal chemistry experiments. W.E.D., H.L., Q.F. and T.N. generated mice. W.E.D., B.N., R.A.V. and L.O.M. analysed mouse data and provided interpretation. B.N., I.C-T., S.B.H., J.B., J.T., C.J.W., V.E.M., J.A.P., D.B., P.M.F., M.A.L., X.M., L.G.H., B.D.M., T.N., M.S., K.M.K., E.P.K. and L.O.M. provided supervision. W.E.D., B.N. and L.O.M. wrote the manuscript with input from other co-authors. L.O.M. devised the concept and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 PIK-III is a selective inhibitor of VPS34 in vitro and in mammalian cells.
(a) U2OS cells expressing GFP-FYVE were incubated with the indicated compounds (2 h), fixed, stained with Hoechst 33342 and imaged by automated epifluorescence microscopy. Representative images of GFP-FYVE are shown and scale bars are equivalent to 10 μm. GFP-positive puncta/cell were determined using an image analysis algorithm and these data were then used to determine IC50 values for each compound as shown in Fig. 1c. (b) U2OS cells expressing GFP-FOXO3A were incubated with the indicated compounds (2 h), fixed, stained with Hoechst 33342 and imaged by automated epifluorescence microscopy. Representative images of GFP-FOXO3A (green) and Hoechst 33342 nuclear staining (blue) are shown and scale bars are equivalent to 10 μm. The ratio of nuclear to cellular GFP fluorescence intensity was determined using an image analysis algorithm and these data were then used to determine IC50 values for each compound as shown in Fig. 1d.
Supplementary Figure 2 Validation of ATG7 conditional knockout DLD1 cells.
ATG7−/− DLD1 cells harbour a conditional knockout of ATG7, with exon 15 of the first allele being disrupted and exon 15 of the second allele being flanked with two loxP sites (see Experimental Procedures). ATG7+/+, ATG7−/+ or two ATG7flox/− DLD1 cell clones were infected with adenoviral-Cre recombinase (+ Adeno-Cre), lysed, and the levels of ATG7 and ATG5 evaluated using western blotting. Note the complete loss of ATG7 in the ATG7−/− clones treated with Adeno-Cre which results in full inhibition of ATG5-ATG12 conjugation. Cells treated with Adeno-Cre were used to generate data shown in Fig. 4d.
Supplementary Figure 3 The effect of autophagy disruption on NCOA4 mRNA.
(a) Total RNA was isolated from cells treated as in Fig. 4C and the relative levels of NCOA4 mRNA determined using quantitative RT-PCR. (b) Total RNA was isolated from cells treated as in Fig 4d and relative levels of NCOA4 and ATG7 mRNA determined using quantitative RT-PCR. Three different PCR probes (11/12, 14/15 and 15/16) were used to amplify ATG7 mRNA with the latter two flanking the targeted exon 15. (c) Total RNA was isolated from cells treated as in Fig. 4e and relative levels of NCOA4 mRNA determined using quantitative RT-PCR. Values are reported are two independent repeats from qPCR performed in quadruplicate.
Supplementary Figure 4 FTH1 and FTL interact with SBP-NCOA4 in DLD1 cells.
Steptavidin agarose beads were incubated with protein extracts from control or SBP-NCOA4-expressing DLD1 cells, bound proteins were eluted and analysed by LC-MS. Proteins enriched in the SBP-NCOA4 over control sample were determined as described in the methods and log10 fold ratios and adjusted P-Values are shown on the y and x axis, respectively. Note the robust enrichment of NCOA4, FTH1, FTL and TMSB10. The complete dataset is shown in Supplementary Table 3. P-values were calculated using a one-way T-test (arbitrarily set to 1 for non-significant single peptide quantitations) and adjusted using the Benjamini-Hochberg False Discovery Rate.
Supplementary Figure 5 Starvation-induced degradation of NCOA4, FTH1 and FTL does not require p62.
DLD1 cells transfected with the indicated siRNAs were left in rich medium (Fed) or nutrient starved (Starve) for the indicated times before cell lysis and western blot analysis.
Supplementary Figure 6 NCOA4 interacts specifically with FTH1.
(a) Endogenous NCOA4 was immunoprecipitated from FTH1−/− cells transfected with empty vector, N-SBP-FTH1 or N-SBP-FTL and the presence of FTH1 and FTL in the immunoprecipitates was determined by western blot. (b) Endogenous NCOA4 was immunoprecipitated from FTH1−/−/FTL−/− double knockout cells transfected with empty vector, N-SBP-FTH1 or N-SBP-FTL and the presence of FTH1 and FTL in the immunoprecipitates was determined by western blot. A specific interaction between NCOA4 and FTH1 only was detected. Asterisk indicates IgG band and arrow highlights location of FTL band.
Supplementary Figure 7 Uncropped scans of key western blots.
Area shown in figures is outlined in red and molecular weight markers are shown on the left.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1154 kb)
Supplementary Table 1
Supplementary Information (PDF 83 kb)
Supplementary Table 2
Supplementary Information (XLSX 5760 kb)
Supplementary Table 3
Supplementary Information (XLSX 186 kb)
Supplementary Table 4
Supplementary Information (PDF 68 kb)
Supplementary Table 5
Supplementary Information (XLSX 8 kb)
Supplementary Table 6
Supplementary Information (PDF 78 kb)
Rights and permissions
About this article
Cite this article
Dowdle, W., Nyfeler, B., Nagel, J. et al. Selective VPS34 inhibitor blocks autophagy and uncovers a role for NCOA4 in ferritin degradation and iron homeostasis in vivo. Nat Cell Biol 16, 1069–1079 (2014). https://doi.org/10.1038/ncb3053
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3053
This article is cited by
-
Mechanisms controlling cellular and systemic iron homeostasis
Nature Reviews Molecular Cell Biology (2024)
-
Lysosomes as coordinators of cellular catabolism, metabolic signalling and organ physiology
Nature Reviews Molecular Cell Biology (2024)
-
Targeting autophagy as a therapeutic strategy in pediatric acute lymphoblastic leukemia
Scientific Reports (2024)
-
Alleviated NCOA4-mediated ferritinophagy protected RA FLSs from ferroptosis in lipopolysaccharide-induced inflammation under hypoxia
Inflammation Research (2024)
-
Iron and copper: critical executioners of ferroptosis, cuproptosis and other forms of cell death
Cell Communication and Signaling (2023)