Abstract
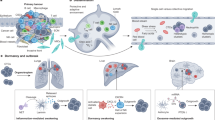
The mechanisms that allow colon cancer cells to form liver and lung metastases, and whether KRAS mutation influences where and when metastasis occurs, are unknown. We provide clinical and molecular evidence showing that different MAPK signalling pathways are implicated in this process. Whereas ERK2 activation provides colon cancer cells with the ability to seed and colonize the liver, reduced p38 MAPK signalling endows cancer cells with the ability to form lung metastasis from previously established liver lesions. Downregulation of p38 MAPK signalling results in increased expression of the cytokine PTHLH, which contributes to colon cancer cell extravasation to the lung by inducing caspase-independent death in endothelial cells of the lung microvasculature. The concerted acquisition of metastatic traits in the colon cancer cells together with the sequential colonization of liver and lung highlights the importance of metastatic lesions as a platform for further dissemination.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Edge, S. B., Byrd, D. R., Compton, C. C. & Fritz, A. G. AJCC Cancer Staging Manual (Springer, 2009).
Gallagher, D. J. & Kemeny, N. Metastatic colorectal cancer: from improved survival to potential cure. Oncology 78, 237–248 (2010).
Wanebo, H. J. et al. Meeting the biologic challenge of colorectal metastases. Clin. Exp. Metastasis 29, 821–839 (2012).
Sadahiro, S. et al. Recurrence patterns after curative resection of colorectal cancer in patients followed for a minimum of ten years. Hepatogastroenterology 50, 1362–1366 (2003).
Kievit, J. Follow-up of patients with colorectal cancer: numbers needed to test and treat. Eur. J. Cancer 38, 986–999 (2002).
Nguyen, D. X., Bos, P. D. & Massagué, J. Metastasis: from dissemination to organ-specific colonization. Nat. Rev. Cancer 9, 274–284 (2009).
Tie, J. et al. KRAS mutation is associated with lung metastasis in patients with curatively resected colorectal cancer. Clin. Cancer Res. 17, 1122–1130 (2011).
Schlüter, K. et al. Organ-specific metastatic tumor cell adhesion and extravasation of colon carcinoma cells with different metastatic potential. Am. J. Pathol. 169, 1064–1073 (2006).
Ding, L. et al. Genome remodelling in a basal-like breast cancer metastasis and xenograft. Nature 464, 999–1005 (2010).
Fidler, I. J. The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nat. Rev. Cancer 3, 453–458 (2003).
Kaiser, S. et al. Transcriptional recapitulation and subversion of embryonic colon development by mouse colon tumor models and human colon cancer. Genome Biol. 8, R131 (2007).
Smith, J. J. et al. Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology 138, 958–968 (2010).
Merlos-Suárez, A. et al. The intestinal stem cell signature identifies colorectal cancer stem cells and predicts disease relapse. Cell Stem Cell 8, 511–524 (2011).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Wagner, E. F. & Nebreda, A. R. Signal integration by JNK and p38 MAPK pathways in cancer development. Nat. Rev. Cancer 9, 537–549 (2009).
Cuadrado, A. & Nebreda, A. R. Mechanisms and functions of p38 MAPK signalling. Biochem. J. 429, 403–417 (2010).
Shin, S., Dimitri, C. A., Yoon, S-O., Dowdle, W. & Blenis, J. ERK2 but not ERK1 induces epithelial-to-mesenchymal transformation via DEF motif-dependent signaling events. Mol. Cell 38, 114–127 (2010).
Xing, L. et al. Discovery and characterization of atropisomer PH-797804, a p38 MAP kinase inhibitor, as a clinical drug candidate. Chem. Med. Chem. 7, 273–280 (2012).
Céspedes, M. V. et al. Orthotopic microinjection of human colon cancer cells in nude mice induces tumor foci in all clinically relevant metastatic sites. Am. J. Pathol. 170, 1077–1085 (2007).
Adams, S. L., Cohen, A. J. & Lassová, L. Integration of signaling pathways regulating chondrocyte differentiation during endochondral bone formation. J. Cell. Physiol. 213, 635–641 (2007).
Hens, J. R. et al. BMP4 and PTHrP interact to stimulate ductal outgrowth during embryonic mammary development and to inhibit hair follicle induction. Development 134, 1221–1230 (2007).
Norberg, E. et al. An increase in intracellular Ca2+ is required for the activation of mitochondrial calpain to release AIF during cell death. Cell Death Differ. 15, 1857–1864 (2008).
Sanges, D., Comitato, A., Tammaro, R. & Marigo, V. Apoptosis in retinal degeneration involves cross-talk between apoptosis-inducing factor (AIF) and caspase-12 and is blocked by calpain inhibitors. Proc. Natl Acad. Sci. USA 103, 17366–17371 (2006).
Vilardaga, J-P., Romero, G., Friedman, P. A. & Gardella, T. J. Molecular basis of parathyroid hormone receptor signaling and trafficking: a family B GPCR paradigm. Cell. Mol. Life Sci. 68, 1–13 (2011).
Susin, S. A. et al. Two distinct pathways leading to nuclear apoptosis. J. Exp. Med. 192, 571–580 (2000).
Joza, N. et al. Essential role of the mitochondrial apoptosis-inducing factor in programmed cell death. Nature 410, 549–554 (2001).
Gallwitz, W. E., Guise, T. A. & Mundy, G. R. Guanosine nucleotides inhibit different syndromes of PTHrP excess caused by human cancers in vivo. J. Clin. Invest. 110, 1559–1572 (2002).
Takayama, Y., Mori, T., Nomura, T., Shibahara, T. & Sakamoto, M. Parathyroid-related protein plays a critical role in bone invasion by oral squamous cell carcinoma. Int. J. Oncol. 36, 1387–1394 (2010).
Guise, T. A. Molecular mechanisms of osteolytic bone metastases. Cancer 88, 2892–2898 (2000).
Malakouti, S., Asadi, F. K., Kukreja, S. C., Abcarian, H. A. & Cintron, J. R. Parathyroid hormone-related protein expression in the human colon: immunohistochemical evaluation. Am. Surg. 62, 540–544 (1996).
Ponomarev, V. et al. A novel triple-modality reporter gene for whole-body fluorescent, bioluminescent, and nuclear noninvasive imaging. Eur. J. Nucl. Med. Mol. Imaging 31, 740–751 (2004).
Giavazzi, R., Campbell, D. E., Jessup, J. M., Cleary, K. & Fidler, I. J. Metastatic behavior of tumor cells isolated from primary and metastatic human colorectal carcinomas implanted into different sites in nude mice. Cancer Res. 46, 1928–1933 (1986).
Céspedes, M. V. et al. Orthotopic microinjection of human colon cancer cells in nude mice induces tumor foci in all clinically relevant metastatic sites. Am. J. Pathol. 170, 1077–1085 (2007).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Rossell, D., Guerra, R. & Scott, C. Semi-parametric differential, expression analysis via partial mixture estimation. Stat. Appl. Genet. Mol. Biol. 7, Article15 (2008).
Mallick, B. K., Gold, D. & Baladandayuthapani, V. Bayesan Analysis of Gene Expression Data (Wiley, 2009).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy. Stat. Soc. Ser. B 57, 289–300 (1995).
Alonso, G., Ambrosino, C., Jones, M. & Nebreda, A. R. Differential activation of p38 mitogen-activated protein kinase isoforms depending on signal strength. J. Biol. Chem. 275, 40641–40648 (2000).
Garcia-Albeniz, X. et al. Serum matrilysin correlates with poor survival independently of KRAS and BRAF status in refractory advanced colorectal cancer patients treated with irinotecan plus cetuximab. Tumour Biol. 32, 417–424 (2011).
Martínez-Fernandez, A. et al. Serum matrilysin levels predict outcome in curatively resected colorectal cancer patients. Ann. Surg. Oncol. 16, 1412–1420 (2009).
Smith, J. J. et al. Experimentally derived metastasis gene expression profile predicts recurrence and death in patients with colon cancer. Gastroenterology 138, 958–968 (2010).
Jorissen, R. N. et al. Metastasis-associated gene expression changes predict poor outcomes in patients with dukes stage B and C colorectal cancer. Clin. Cancer Res. 15, 7642–7651 (2009).
Acknowledgements
We would like to thank J. Maurel for providing the data on patients with colon and rectal cancers depicted in Supplementary Table 3. We also thank H. Auer, J. Colombelli and M. Gay from the Functional genomics, Microscopy and Mass spectrometry core facilities of IRB Barcelona, respectively. J.U. was supported by Fundación BBVA, a Juan de la Cierva grant from Spanish Ministerio de Ciencia e Innovación (MICINN) and a grant from Fundación Olga Torres. X.G-A. was supported by Hospital Clinic-IDIBAPS (Ajut a la Recerca Josep Font) and an ASISA-Harvard Fellowship for Excellence in Clinical Research. S.R. was supported by a S. Borrel contract from Instituto de Salud Carlos III. S.G. and M.P. were supported by La Caixa Fellowships and A.B. by FPI from MICINN. R.R.G., E.B. and A.R.N. are supported by the Institució Catalana de Recerca i Estudis Avançats (ICREA). Research support was provided by MICINN (BFU2010-17850), the European Commission FP7 (ERC 294665) and the Fundación BBVA to A.R.N, by the Generalitat de Catalunya (2009 SGR 1437), MINECO P12/01861 and CIBER-BBN NanoCoMets to R.M., and by the Fundación BBVA, AECC, Generalitat de Catalunya (2009 SGR 1429) and MICINN (SAF2010-21171) to R.R.G.
Author information
Authors and Affiliations
Contributions
J.U. designed and performed experiments, analysed data and participated in text writing. X.G-A designed and established the model system, performed microarrays and participated in text writing and statistical analyses of microarrays and patient data. E.P. performed statistical analyses of patient data. S.R., I.D. and F.M.B. performed experiments and analysed data on p38 MAPK. M.V.C. and R.M carried out intracaecum injections and analysed the data. A.B. and S.G. contributed to western blots and immunohistochemical analysis. M.G. and M.P. carried out intrasplenic, tail vein and intracaecum injections. E.F. carried out immunohistochemical analysis. C.N. provided and analysed patient data. N.K, E.B. and A.R.N. participated in data analyses and manuscript writing. R.R.G conceived the project, analysed data, supervised the overall project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Characterization of liver and lung metastatic potential of SW620 parental and LiM2 cells.
(a) Invasion capacity of SW620 parental and LiM2 cells was measured using matrigel-coated Boyden chambers. Results represent values of three independent experiments where each cell line was seeded in six Boyden chambers and five fields per chamber were analysed and average for each chamber obtained (n = 18 chambers per group). Statistical significance was calculated using two-tailed Mann Whitney test; the box extends from 25 to 75 percentile where black line within the box represents median, whiskers extend from 10 to 90 percentile. (b) In vivo proliferation of parental and LiM2 cells (0.5 × 105) injected into cecum of nude mice. Bioluminescent images were taken at day 54. n = 8 (Parental) and n = 9 (LiM2) number of mice used for each cell line. Statistical analysis was done using two-tailed Mann Whitney test and no statistically significant differences were found. (c) Percentage of lymph node and liver metastasis as well as lung metastasis in mice that developed liver metastasis on intracecum injections of parental and LiM2 cells determined post-mortem. Number of mice (n) used is defined in (b). Statistical significance was calculated using one-sided Fisher’s exact test. (d) Representative bioluminescent images of intestine, liver and lung from mice injected intracecum with SW620 parental and LiM2 cells. Data is presented as average plus s.d.
Supplementary Figure 2 Gene set enrichment analyses (GSEA) of the CCM downregulated gene set and phosphorylated p38 MAPK levels in liver metastatic lesions formed by LiM2 cells.
(a) GSEA analysis of the CCM downregulated gene set in human colon cancer dataset (pooled GSE17537 and GSE14333 expression set). NES-normalized enrichment score; FDR-false discovery rate. (b) SW620 parental and LiM2 cells were injected intraliver and 40 days later mice were sacrificed. Size matched liver lesions were selected for immunohistochemistry staining with anti- phospho-p38 (P-p38) antibodies. Scale bar, 500 μm; inset scale bar, 50 μm; H-healthy tissue; T-tumour.
Supplementary Figure 3 ERK1 and ERK2 influence on primary tumours and metastasis.
(a) ERK1 and ERK2 downregulation was confirmed by Western blotting. (b) Percentage of mice intrasplenically injected with LiM2 cells expressing the indicated shRNAs that developed liver metastases (up) or lung metastasis (bottom). n = 17 (shControl), n = 10 (sh1 ERK1) and n = 18 (sh2 ERK1) number of mice used per group pooled from two independent experiments. Statistical analysis was done using two-sided Fisher’s exact test. n.s.-not significant. (c) LiM2 cells expressing the indicated shRNAs were injected orthotopically into mice cecum and growth rates by photon flux quantification were determined. n = 9 (shControl), n = 9 (sh1 ERK1), n = 8 (sh2 ERK1), n = 10 (sh1 ERK2), n = 8 (sh2 ERK2) number of mice used per group. Data is presented as average plus s.d. Statistical analysis was done using one-way ANOVA and no statistical differences were found. (d, e) Size matched lesions from (c) were selected for immunohistochemistry Ki-67 (d) and cleaved caspase-3 staining (e). n = 20 total fields per group analysed where five different fields from size matched lesions per mouse were quantified. Number of mice used in each group was 4. Statistical analysis was done using two-tailed Mann Whitney test. All values are presented in the graphs together with average ±s.d. (f) Kaplan–Meier curves representing association of proportion of recurrence-free patients with relative FRA1 gene expression levels in human primary colon cancer dataset (pooled GSE17537 and GSE14333; n = 267). HR-hazard ratio. Statistical analysis was done using Cox proportional hazards model.
Supplementary Figure 4 Lung metastasis in colon cancer associates with low levels of phopsho-p38 MAPK in primary tumours.
(a) Association of lung metastasis with levels of phosphorylated p38 (P-p38) protein in samples of 7 primary CRC tumour samples that developed metastasis. P-p38 levels were normalized to p38 total amount of protein and P-p38 expressed in the respective healthy mucosa samples. n = 3 tumours with lung mets and n = 4 tumours with other mets. Statistical analysis was done using one-tailed Mann Whitney test. (b) HCT116 cells expressing p38α shRNAs or constitutively active MKK6EE were injected orthotopically into mice cecum and size matched lesions were selected for Ki-67 and cleaved caspase-3 staining. Five different fields per lesion and mouse for Ki67 staining and ten different fields /lesion/mouse for cleaved caspase-3 staining were scored. n = 40 (Mock), n = 30 (sh p38α) and n = 40 (MKK6EE) total number of fields per group analysed for Ki67 staining. n = 80 (Mock), n = 60 (sh p38α) and n = 80 (MKK6EE) total number of fields per group analysed for cleaved caspase-3 staining. (c) HCTT116 cells expressing p38αshRNAs or constitutively active MKK6EE were injected orthotopically into mice cecum and the percentage of mice that developed liver metastasis, the area of liver and lung metastasis were determined. #, † indicate the groups which statistically significant. #P = 0.001; †P = 0.017. Values for area represent mean ± s.e.m. n = 41 (Mock), n = 14 (sh p38α) and n = 61 (MKK6EE) number of lung lesions analysed. (d) Representative H&E staining of lung and liver histological sections from (c). Dashed lines delineate metastasis. Scale bar, 200 μm. (e) Emerging lung metastatic lesions from HCTT116 cells expressing p38αshRNAs or constitutively active MKK6EE injected orthotopically into mice cecum were selected for cleaved caspase-3 staining. n = 9 (Mock), n = 11 (sh p38α) and n = 5 (MKK6EE) total number of lesions analysed per group. (f) Emerging lung metastatic lesions from SW620 parental and LiM2 cells were selected for cleaved caspase-3 staining. n = 6 (Parental) and n = 9 (LiM2) total number of lesions analysed per group. Scale bar, 50 μm. For (b),(c), (e) and (f) two-tailed Mann Whitney test was used for statistical analysis; n.s.-not significant. In panels (a), (b), and (f) all values are presented in the graphs together with average ±s.d. In panel (e) all values are presented in the graph together with average plus s.d.
Supplementary Figure 5 PTHLH gene expression is associated with poor clinical outcome in patients with colon cancer.
(a) Kaplan–Meier curves representing association of proportion of recurrence-free patients with relative PTHLH gene expression levels in human primary colon cancer dataset (pooled GSE17537 and GSE14333; n = 267). HR-hazard ratio. Statistical analysis was done using Cox proportional hazard’s model. (b, c) ELISA analysis of PTHLH levels in HCTT116 cells (n = 3 values from independent experiments) expressing constitutively active MKK6EE (b) or immunofluorescence analysis of PTHLH levels in SW620 cells expressing PTHLH (c). Statistical analysis was done using one-tailed Student’s t test in (b) and graphs represent average plus s.d. Scale bar, in (c) 50 μM. (d) Detection of PTHLH by mass-spec analysis in conditioned medium from SW620 parental, LiM2 shControl and LiM2 shPTHLH cell lines.
Supplementary Figure 6 PTHLH does not affect the growth of LiM2 cells in the lung.
(a) Photon flux of dorsal upper back area of mice injected intrapulmonary with LiM2 cells expressing two different shRNA against PTHLH. Normalization was done to values of day 0. n = 9 (sh Control), n = 9 (sh1 PTHLH) and n = 9 (sh2 PTHLH) number of mice used per group. Inset represents qRT–PCR analysis of PTHLH expression levels in LiM2 cells infected with lentivirus expressing shRNA against PTHLH or the control shRNA in three independent experiments (n = 3). Statistical analysis for main graph was done using two-tailed Mann Whitney test and no statistically significant differences were found. Statistics for the graph presented in inset was done using two-tailed Student’s t test. **P < 0.001. Values represented in the main graph as well as in inset represent average plus s.d. (b) Representative bioluminescent images of mice injected intrapulmonary with LiM2 cells expressing two different shRNAs against PTHLH. (c) Lung metastasis-free survival of mice (n = 14) on tail vein injection of LiM2 cells. Representative bioluminescent images are shown.
Supplementary Figure 7 Effect of PTHLH on endothelial cells of the lung.
(a) Migration of LiM2 cells mock infected or infected with a retrovirus that expresses PTHLH was scored in Boyden chambers covered with fibronectin or with fibronectin and HPMEC monolayer. Each cell line was seeded in triplicate and 5 fields per chamber were counted. Results represent three independent experiments for migration through fibronectin and four independent experiments for migration through fibronectin and HPMEC monolayer. n = 45 total number of fields analysed per group for migration through fibronectin and n = 60 total number of fields analysed per group for migration through fibronectin and HPMEC monolayer. Statistical analysis was done using two-tailed Mann Whitney test (b) Western blot analysis of PTH1R expression levels in HPMEC cells. Tubulin was used as a loading control. (c) HPMEC cells were grown until reaching tight confluence and then were treated for 24 h with conditioned medium from SW620 parental, LiM2, LiM2 shPTHLH and LiM2 treated cells with PTHLH-Antagonist Peptide (AP). Phalloidin staining was performed and total gap area per field was determined. Results represent values of three independent experiments where each sample was seeded in duplicate and for each coverslip at least 5 fields were analysed. n = 40, 40, 40 and 30 total number of fields analysed per group respectively. Statistical analysis was done using two-tailed Student’s t test; scale bar, 100 μm. (d) Average number of TUNEL-positive cells in ovary or liver 4 h post tail vein injection of recombinant PTHLH (1–34). Three 30 μm distant sections per animal were counted. PBS, 1 nM, 10 nM, and 100 nM rPTHLH (5 mice per group). n = 15 total number of sections analysed per group. Statistical analysis was done using two-tailed Mann Whitney test; n.s.-not significant. All values are presented in the graphs together with average ±s.d. (e) Western blot analysis of total and cleaved PARP, Caspases-3 and Caspases-7 in HPMEC cells treated for 20 h with PTHLH or Staurosporine. Tubulin was used as a loading control. (f) Western blot analysis of pro-apoptotic and anti-apoptotic protein levels in HPMEC cells on 4 h treatment with the indicated doses of recombinant PTHLH. Tubulin was used as a loading control. (g) HPMEC cells were treated for 4 h with the indicated doses of recombinant PTHLH and the cytosolic and mitochondrial fractions were isolated and subjected to western blot analysis for AIFM1 protein expression levels. Tom 20 was used as a loading control of mitochondria and tubulin as a loading control of cytosol. In panels (a) and (c) the box extends from 25 to 75 percentile where black line within the box represents median, the whiskers extend from 10 to 90 percentile.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1489 kb)
Supplementary Table 1
Supplementary Information (XLSX 58 kb)
Supplementary Table 2
Supplementary Information (XLSX 41 kb)
Supplementary Table 3
Supplementary Information (XLSX 35 kb)
Supplementary Table 4
Supplementary Information (XLSX 33 kb)
PTHLH treatment increases intracellular Ca2+ levels.
Cells were treated for 15 min with Fluo4-AM calcium indicator and then imaged for 9 min. First two minutes of imaging were used for determining the basal Ca2+ levels. At 2 min, control media was added to cells and at 5 min media with 1.25 μM recombinant PTHLH. (AVI 2664 kb)
Rights and permissions
About this article
Cite this article
Urosevic, J., Garcia-Albéniz, X., Planet, E. et al. Colon cancer cells colonize the lung from established liver metastases through p38 MAPK signalling and PTHLH. Nat Cell Biol 16, 685–694 (2014). https://doi.org/10.1038/ncb2977
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2977
This article is cited by
-
Litchi procyanidins inhibit colon cancer proliferation and metastasis by triggering gut-lung axis immunotherapy
Cell Death & Disease (2023)
-
Homogeneous and heterogeneous risk and prognostic factors for lung metastasis in colorectal cancer patients
BMC Gastroenterology (2022)
-
Intracellular and extracellular factors of colorectal cancer liver metastasis: a pivotal perplex to be fully elucidated
Cancer Cell International (2022)
-
Seven key hub genes identified by gene co-expression network in cutaneous squamous cell carcinoma
BMC Cancer (2021)
-
Diversity and versatility of p38 kinase signalling in health and disease
Nature Reviews Molecular Cell Biology (2021)